

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
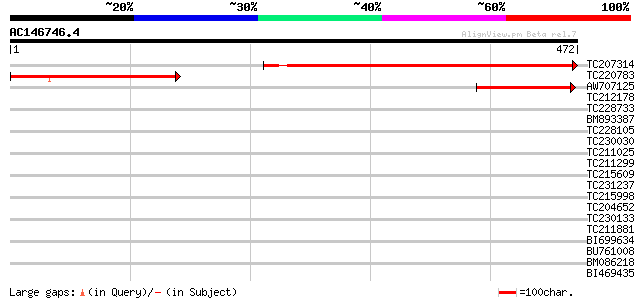
Query= AC146746.4 - phase: 0
(472 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC207314 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associat... 420 e-118
TC220783 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associat... 207 8e-54
AW707125 149 3e-36
TC212178 similar to UP|Q9SKH2 (Q9SKH2) Expressed protein, partia... 37 0.015
TC228733 weakly similar to GB|AAT41789.1|48310278|BT014806 At3g4... 37 0.025
BM893387 weakly similar to PIR|A86321|A86 F6A14.17 protein - Ara... 37 0.025
TC228105 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {A... 36 0.043
TC230030 similar to UP|Q9M9U3 (Q9M9U3) F6A14.17 protein (At1g187... 36 0.043
TC211025 similar to PIR|T05269|T05269 adenylyl cyclase-associate... 35 0.056
TC211299 weakly similar to GB|AAP37814.1|30725584|BT008455 At1g7... 35 0.072
TC215609 weakly similar to UP|Q70DK2 (Q70DK2) Blue copper bindin... 35 0.095
TC231237 weakly similar to GB|AAL69460.1|18491271|AY074644 At2g4... 35 0.095
TC215998 similar to UP|Q9LV44 (Q9LV44) Similarity to signal pept... 34 0.12
TC204652 similar to UP|H1_PEA (P08283) Histone H1, partial (81%) 34 0.12
TC230133 weakly similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%) 34 0.12
TC211881 similar to PIR|T48135|T48135 nucleotide sugar epimerase... 34 0.16
BI699634 33 0.21
BU761008 similar to PIR|T48135|T481 nucleotide sugar epimerase-l... 33 0.21
BM086218 similar to PIR|T04107|T041 calmodulin-binding heat-shoc... 33 0.21
BI469435 33 0.21
>TC207314 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associated protein,
partial (51%)
Length = 1248
Score = 420 bits (1079), Expect = e-118
Identities = 209/261 (80%), Positives = 228/261 (87%)
Frame = +2
Query: 212 QTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVT 271
Q GK+ APSK +A P+APPPPPASLFSSESSQASSSKPK GMSAVFQ+I +GNVT
Sbjct: 23 QPGKIIAPSKASA------PAAPPPPPASLFSSESSQASSSKPKEGMSAVFQQISSGNVT 184
Query: 272 AGLRKVTDDMKTKNRADRSGVVGNSVKESQAAPRAFSKVGPPKLELQMGRKWVVENQIDQ 331
+GL+KVT+DMKTKNR DR+GVVG KE+ A+ R FSK GPPK ELQMGRKWVVENQI++
Sbjct: 185 SGLKKVTNDMKTKNRTDRTGVVGAIEKETPASSRVFSKAGPPKFELQMGRKWVVENQIEK 364
Query: 332 KSLVIEDCDSKQSVYVYGCKNSVLQIQGKVNNITIDNCKKTGVVFKDVVAAFEVVNSNGV 391
K LVIEDCDSKQSVY+YGCKNSVLQI GKVNNITID C K GVVFKDVVAA E+VNSNGV
Sbjct: 365 KDLVIEDCDSKQSVYIYGCKNSVLQILGKVNNITIDKCTKIGVVFKDVVAACEIVNSNGV 544
Query: 392 EVQCQGSAPTISVDNTSGCQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHSLPQ 451
EVQCQG+APTISVDNTSGCQ+YLSKDSLE SISTAKSSEINVLVP + DGDWVEHSLPQ
Sbjct: 545 EVQCQGAAPTISVDNTSGCQLYLSKDSLEASISTAKSSEINVLVPGADPDGDWVEHSLPQ 724
Query: 452 QYIHLFKEGRFETTPASHSGG 472
QYIHLFK FETTPA+HSGG
Sbjct: 725 QYIHLFKNEHFETTPAAHSGG 787
>TC220783 similar to UP|Q9ZWM6 (Q9ZWM6) Adenylyl cyclase associated protein,
partial (26%)
Length = 575
Score = 207 bits (527), Expect = 8e-54
Identities = 109/145 (75%), Positives = 126/145 (86%), Gaps = 3/145 (2%)
Frame = +3
Query: 1 MDETLIKRLESAVTRLEALSTGIHHSSSASD---ASDAASDPSVVAFVDLIDQYVSRLSK 57
MDE LI+RLESAV+RLE+LS G+H S++ S A DAA DPS+VAF DLIDQYV R+S
Sbjct: 141 MDEKLIQRLESAVSRLESLSAGLHPSATQSGGAHAPDAALDPSIVAFADLIDQYVGRVSG 320
Query: 58 AADIIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTE 117
AA++IGGQVLDVT V+EAF+VQKELLIKL+QTQKP+ AGL+EFLKPLNEVI KA+ LTE
Sbjct: 321 AAEVIGGQVLDVTKLVQEAFNVQKELLIKLRQTQKPNLAGLSEFLKPLNEVITKATKLTE 500
Query: 118 GRRSDFFNHLKAAVDSLSALAWIAF 142
GRRSDFFNHLKAA DSLSALAWIA+
Sbjct: 501 GRRSDFFNHLKAAADSLSALAWIAY 575
>AW707125
Length = 413
Score = 149 bits (375), Expect = 3e-36
Identities = 70/83 (84%), Positives = 76/83 (91%)
Frame = +2
Query: 389 NGVEVQCQGSAPTISVDNTSGCQIYLSKDSLETSISTAKSSEINVLVPNVESDGDWVEHS 448
+GVEVQCQG+APTISVDNTSGCQ+YLSKDSLE SISTAKSSEINVLVP + DGDWVEHS
Sbjct: 5 DGVEVQCQGAAPTISVDNTSGCQLYLSKDSLEASISTAKSSEINVLVPGADPDGDWVEHS 184
Query: 449 LPQQYIHLFKEGRFETTPASHSG 471
LPQQYIHLFK F+TTPA+HSG
Sbjct: 185 LPQQYIHLFKNEHFKTTPAAHSG 253
>TC212178 similar to UP|Q9SKH2 (Q9SKH2) Expressed protein, partial (12%)
Length = 728
Score = 37.4 bits (85), Expect = 0.015
Identities = 27/98 (27%), Positives = 45/98 (45%), Gaps = 2/98 (2%)
Frame = -3
Query: 231 PSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKN--RAD 288
P PPPPP S SS ++ AS P +A + G+ + A R+ D + + A
Sbjct: 441 PLTPPPPPCSRRSSVAASASRPAPPHSPAAAARSRGSCGIEAARRRGRDRSRWERLFPAT 262
Query: 289 RSGVVGNSVKESQAAPRAFSKVGPPKLELQMGRKWVVE 326
RSG ++ PR +++ + L R+WV++
Sbjct: 261 RSGA-------GRSGPRWETRIWRSAVSLGPKRRWVLK 169
>TC228733 weakly similar to GB|AAT41789.1|48310278|BT014806 At3g44220
{Arabidopsis thaliana;} , partial (31%)
Length = 797
Score = 36.6 bits (83), Expect = 0.025
Identities = 23/83 (27%), Positives = 38/83 (45%), Gaps = 2/83 (2%)
Frame = +1
Query: 228 PAAPSA--PPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDDMKTKN 285
P+ P PPP P+ L S SS + + +++ E +G + G ++T M T
Sbjct: 34 PSGPPTFTPPPSPSPLSLSRSSSRTEPPEEFWLTSKSAEE*SGRLELGFPEITILMSTVR 213
Query: 286 RADRSGVVGNSVKESQAAPRAFS 308
S V+G + +S P +FS
Sbjct: 214 HTSGSPVIGTTPLDSPVPPSSFS 282
>BM893387 weakly similar to PIR|A86321|A86 F6A14.17 protein - Arabidopsis
thaliana, partial (50%)
Length = 399
Score = 36.6 bits (83), Expect = 0.025
Identities = 21/51 (41%), Positives = 25/51 (48%)
Frame = +1
Query: 210 WSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSA 260
WS TG PS +S +P P PP+S S+ S ASS P G SA
Sbjct: 58 WSSTGASSPPSSTPSSTSPWITRPAPSPPSSPSSAGSLPASSLIPSPGRSA 210
>TC228105 similar to GB|AAP37692.1|30725340|BT008333 At4g29830 {Arabidopsis
thaliana;} , complete
Length = 1113
Score = 35.8 bits (81), Expect = 0.043
Identities = 23/59 (38%), Positives = 28/59 (46%), Gaps = 4/59 (6%)
Frame = +1
Query: 200 VKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAP----PPPPASLFSSESSQASSSKP 254
V S P G T AP+ ASA+P +P P PPPP S SS SS + + P
Sbjct: 103 VPSTRPSGSGAPTTSFWTAPTPATASASPPSPPTPSALSPPPPPSTASSASSMSIPTPP 279
>TC230030 similar to UP|Q9M9U3 (Q9M9U3) F6A14.17 protein
(At1g18720/F6A14_17), partial (17%)
Length = 579
Score = 35.8 bits (81), Expect = 0.043
Identities = 19/50 (38%), Positives = 25/50 (50%)
Frame = +2
Query: 210 WSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMS 259
WS TG PS +S +P P PP+S +S+ S A+S P G S
Sbjct: 359 WSSTGASSPPSSTPSSTSPWITRPAPSPPSSPYSAGSLTATSLIPSPGRS 508
>TC211025 similar to PIR|T05269|T05269 adenylyl cyclase-associated protein
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (4%)
Length = 679
Score = 35.4 bits (80), Expect = 0.056
Identities = 23/64 (35%), Positives = 27/64 (41%), Gaps = 33/64 (51%)
Frame = +1
Query: 347 VYGCKNSVLQIQGK---------------------------------VNNITIDNCKKTG 373
+YGC+NSVL+IQGK VNNITID C K G
Sbjct: 223 LYGCQNSVLKIQGKSFFLSFSYLLSSYLALDEKAKSN*IISKQYAGKVNNITIDKCTKIG 402
Query: 374 VVFK 377
VF+
Sbjct: 403 GVFE 414
>TC211299 weakly similar to GB|AAP37814.1|30725584|BT008455 At1g76920
{Arabidopsis thaliana;} , partial (25%)
Length = 450
Score = 35.0 bits (79), Expect = 0.072
Identities = 17/53 (32%), Positives = 31/53 (58%)
Frame = +3
Query: 202 SFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKP 254
S +GP S + S +A+++P +PS PP PP+S +S S+ ++ ++P
Sbjct: 117 SMSSIGPASSVAATAIS-SSWSAASSPPSPSTPPAPPSSSSASTSTTSAGTRP 272
>TC215609 weakly similar to UP|Q70DK2 (Q70DK2) Blue copper binding protein,
partial (14%)
Length = 843
Score = 34.7 bits (78), Expect = 0.095
Identities = 17/44 (38%), Positives = 27/44 (60%)
Frame = +1
Query: 218 APSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAV 261
+PS V+ +A+PA ++PP PP S SES S++ P + S +
Sbjct: 214 SPSSVSPAASPATGNSPPAPPTS-SPSESPLPSAATPNISPSTI 342
>TC231237 weakly similar to GB|AAL69460.1|18491271|AY074644
At2g41330/F13H10.12 {Arabidopsis thaliana;} , partial
(24%)
Length = 583
Score = 34.7 bits (78), Expect = 0.095
Identities = 16/35 (45%), Positives = 21/35 (59%)
Frame = +2
Query: 226 AAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSA 260
+AP AP APPPPP S +S +S+S P +A
Sbjct: 95 SAPGAPHAPPPPPHSRPASTRVSSSTSPPSASSAA 199
>TC215998 similar to UP|Q9LV44 (Q9LV44) Similarity to signal peptidase,
partial (33%)
Length = 542
Score = 34.3 bits (77), Expect = 0.12
Identities = 17/37 (45%), Positives = 22/37 (58%)
Frame = -1
Query: 219 PSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPK 255
PS +AS +P P +PPPPP SS+ S A S P+
Sbjct: 260 PSSFSASRSPPPPPSPPPPPP--LSSDLSTALKSSPE 156
>TC204652 similar to UP|H1_PEA (P08283) Histone H1, partial (81%)
Length = 1347
Score = 34.3 bits (77), Expect = 0.12
Identities = 21/61 (34%), Positives = 28/61 (45%)
Frame = +3
Query: 201 KSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSA 260
KS P P + K AP+K +A P A +A P PA+ + A+ SKP A
Sbjct: 603 KSKAPAKPKPAAKPKAKAPAKAKPAAKPKASAAAKPKPAAAKPKAAKPAAKSKPAAKTKA 782
Query: 261 V 261
V
Sbjct: 783 V 785
>TC230133 weakly similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%)
Length = 850
Score = 34.3 bits (77), Expect = 0.12
Identities = 16/38 (42%), Positives = 26/38 (68%)
Frame = -1
Query: 223 NASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSA 260
++S++P++ S PPPP AS SSES+ + S P+ S+
Sbjct: 454 SSSSSPSSSSLPPPPLASGSSSESASSPPSSPEKSSSS 341
>TC211881 similar to PIR|T48135|T48135 nucleotide sugar epimerase-like
protein - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (23%)
Length = 529
Score = 33.9 bits (76), Expect = 0.16
Identities = 19/48 (39%), Positives = 27/48 (55%)
Frame = +3
Query: 225 SAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTA 272
SA+P+ P A PPPPASL SS ASS+ S+ + + + T+
Sbjct: 312 SASPS-PLAAPPPPASLSSSPEPPASSAPTSPSPSSAVATVSSASTTS 452
Score = 31.2 bits (69), Expect = 1.0
Identities = 17/44 (38%), Positives = 25/44 (56%)
Frame = +3
Query: 208 PVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASS 251
P + TGK +PS + A PA+ S+ P PPAS + S +S+
Sbjct: 288 PGAAPTGKSASPSPLAAPPPPASLSSSPEPPASSAPTSPSPSSA 419
>BI699634
Length = 421
Score = 33.5 bits (75), Expect = 0.21
Identities = 17/41 (41%), Positives = 26/41 (62%), Gaps = 1/41 (2%)
Frame = +3
Query: 215 KVFAPSKVNASAAPAAPSAP-PPPPASLFSSESSQASSSKP 254
+V+ S+ SA+P A S P PPPP+S ++ SS ++ S P
Sbjct: 282 RVWISSRTTTSASPRALSRPSPPPPSSPVNTSSSTSTPSSP 404
>BU761008 similar to PIR|T48135|T481 nucleotide sugar epimerase-like protein
- Arabidopsis thaliana, partial (11%)
Length = 444
Score = 33.5 bits (75), Expect = 0.21
Identities = 17/40 (42%), Positives = 24/40 (59%)
Frame = +3
Query: 213 TGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSS 252
TG+ PS A PA+PS+ P PPASL + S +S++
Sbjct: 309 TGRSAWPSPPAAPPPPASPSSSPAPPASLAPTSPSPSSAA 428
>BM086218 similar to PIR|T04107|T041 calmodulin-binding heat-shock protein -
common tobacco, partial (8%)
Length = 430
Score = 33.5 bits (75), Expect = 0.21
Identities = 16/27 (59%), Positives = 17/27 (62%)
Frame = +2
Query: 226 AAPAAPSAPPPPPASLFSSESSQASSS 252
AA AP PPPPP S S +S ASSS
Sbjct: 194 AATTAPRGPPPPPTSSTRSHASAASSS 274
>BI469435
Length = 426
Score = 33.5 bits (75), Expect = 0.21
Identities = 18/35 (51%), Positives = 22/35 (62%)
Frame = +3
Query: 225 SAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMS 259
S + AAPSAPPPP ++ S Q SS KPK +S
Sbjct: 99 STSTAAPSAPPPPSPQTTTTTSPQ-SSPKPKSSLS 200
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.128 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,197,364
Number of Sequences: 63676
Number of extensions: 267910
Number of successful extensions: 5701
Number of sequences better than 10.0: 251
Number of HSP's better than 10.0 without gapping: 3575
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4957
length of query: 472
length of database: 12,639,632
effective HSP length: 101
effective length of query: 371
effective length of database: 6,208,356
effective search space: 2303300076
effective search space used: 2303300076
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146746.4


