

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146571.7 + phase: 0 /pseudo
(2018 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
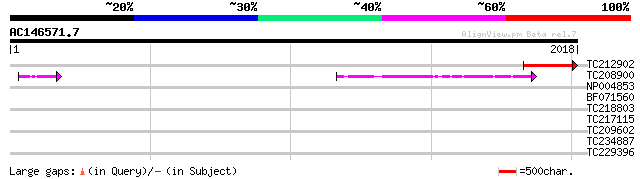
Score E
Sequences producing significant alignments: (bits) Value
TC212902 similar to UP|O64795 (O64795) T1F15.3 protein, partial ... 356 5e-98
TC208900 UP|DPOD_SOYBN (O48901) DNA polymerase delta catalytic s... 293 5e-79
NP004853 nodulin 33 1.3
BF071560 32 2.1
TC218803 similar to UP|Q84P72 (Q84P72) Receptor-like protein kin... 32 2.8
TC217115 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D... 32 3.7
TC209602 similar to UP|Q940G9 (Q940G9) Periaxin-like protein, pa... 31 4.8
TC234887 weakly similar to UP|Q6H7K3 (Q6H7K3) Wall-associated ki... 31 6.2
TC229396 similar to UP|Q9LVG4 (Q9LVG4) Emb|CAB86035.1 (Pseudo-re... 30 8.1
>TC212902 similar to UP|O64795 (O64795) T1F15.3 protein, partial (9%)
Length = 803
Score = 356 bits (914), Expect = 5e-98
Identities = 171/191 (89%), Positives = 178/191 (92%)
Frame = +2
Query: 1828 IHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNQWFAEM 1887
IHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLN WF+EM
Sbjct: 2 IHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQIIPALQRVFGLVGADLNHWFSEM 181
Query: 1888 PRPVREASVKHAFTPNFQRTRIDYYYLSKHCVLCGGLVQASARLCSQCSENEVAAATAVI 1947
PRP REAS KH T NF +TRIDYYYLSKHCVLC LVQASARLC+QCSENEVAAATAVI
Sbjct: 182 PRPTREASAKHTLTTNFHQTRIDYYYLSKHCVLCDRLVQASARLCNQCSENEVAAATAVI 361
Query: 1948 GKTAKLEQEMQHLVSICQHCGGGDRLLESGVKCTSISCLVFYERRKVQKELLAATHVAAD 2007
KT+KLEQEMQHLV++C HCGGGDRLLE+GVKCTSISCLVFYERRKVQKELLAATHVAAD
Sbjct: 362 SKTSKLEQEMQHLVAVCHHCGGGDRLLENGVKCTSISCLVFYERRKVQKELLAATHVAAD 541
Query: 2008 KGFYPRCTVEW 2018
K YPRCTVEW
Sbjct: 542 KDLYPRCTVEW 574
>TC208900 UP|DPOD_SOYBN (O48901) DNA polymerase delta catalytic subunit ,
partial (90%)
Length = 3070
Score = 293 bits (750), Expect = 5e-79
Identities = 216/727 (29%), Positives = 346/727 (46%), Gaps = 15/727 (2%)
Frame = +2
Query: 1162 LLSIEVLAESRGDLLPDPQFDAVNIVALGFQKDCDATVEVLVLLHSKFVPCQRSLDGLSD 1221
+LS ++ R P+P D V +A LV L + P R++ L
Sbjct: 938 ILSFDIECAGRKGHFPEPTHDPVIQIAN------------LVTLQGEDQPFIRNVMTLKS 1081
Query: 1222 CK------VLNFTDEKHLLKEFTKIVSSSDPDILMGWEIQGSSLGFLAERASHLGFGLLN 1275
C V+ F E+ +L + + DPDI++G+ I L +L ERA +L
Sbjct: 1082 CSPIVGVDVMPFETEREVLLAWRDFIREVDPDIIIGYNICKFDLPYLIERALNLKIAEFP 1261
Query: 1276 DLSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGG 1335
L R NS + +D S + ++G + V V G
Sbjct: 1262 ILGRI-RNSRVRVKDTTFSSR-------------------------QYGTRESKEVAVEG 1363
Query: 1336 RIVLNLWRLIRGEVKLNLYSVEAVAEAVLRRKVPSLNHNVLTKWFSRGPGQARYQCIKYI 1395
R+ +L ++++ + KL+ YS+ +V+ L + ++H++++ G + R + Y
Sbjct: 1364 RVTFDLLQVMQRDYKLSSYSLNSVSSHFLSEQKEDVHHSIISD-LQNGNAETRRRLAVYC 1540
Query: 1396 VDRAKLNLEILNQLDMVNRTSELARVFGIDFFSVLSRGSQYRVESMFLRLAHTQNYVAIS 1455
+ A L +L++L + E+ARV G+ +LSRG +V S LR A +N V
Sbjct: 1541 LKDAYLPQRLLDKLMFIYNYVEMARVTGVPISFLLSRGQSIKVLSQLLRRARQKNLVI-- 1714
Query: 1456 PGNQQVASQPAMECLPLVMEPESGFYSDPVVVLDFQSLYPSMIIAYNLCFCTCLGKVVPS 1515
P +Q S+ V+E +GFY P+ LDF SLYPS+++AYNLC+CT V+P
Sbjct: 1715 PNAKQAGSEQGTFEGATVLEARAGFYEKPIATLDFASLYPSIMMAYNLCYCTL---VIPE 1885
Query: 1516 KTNTLGVSPFVPEQHILQDLKDQILLTPNGVMFVPSKIRRGVLPRLLEEILSTRIMVKQA 1575
L + P + + TP+G FV S +++G+LP +LEE+L+ R K+A
Sbjct: 1886 DARKLNIPP------------ESVNRTPSGETFVKSNLQKGILPEILEELLTAR---KRA 2020
Query: 1576 MKKLSPSEQVLQR-IFNARQLALKLISNVTYGYTAAGFSGRMPCAELADSIVQCGRCTLE 1634
L ++ L++ + + RQLALK+ +N YG+T A G++PC E++ S+ GR +E
Sbjct: 2021 KADLKEAKDPLEKAVLDGRQLALKISANSVYGFTGATI-GQLPCLEISSSVTSYGRQMIE 2197
Query: 1635 KAISFV-------NLHEKWNAKVIYGDTDSMFVLLKGRTAKEAFQIGSEIASAITAMNPS 1687
V N +E NA+VIYGDTDS+ V +EA +G E A I+
Sbjct: 2198 HTKKIVEDKFTTLNGYEH-NAEVIYGDTDSVMVQFGVSAVEEAMNLGREAAEHISGTFTK 2374
Query: 1688 PVTLKMEKVYHPCFLITKKRYVGYSYENPNQIEPVFDAKGIETVRRDTCEAVAKIMEQSL 1747
P+ L+ EKVY+P LI+KKRY G + P+ + + D KGIETVRRD C V ++ L
Sbjct: 2375 PIKLEFEKVYYPYLLISKKRYAGLFWTKPDNFDKM-DTKGIETVRRDNCLLVKNLVNDCL 2551
Query: 1748 RLFFEHQSLLEVKTYLQRQWKRILSGRVSLKDFIFAKEVRLGTYSARISSLPPAAIVATK 1807
+ + Y++ +L R+ L + K + T + + A + +
Sbjct: 2552 HKILIDRDIPGAVQYVKNAISDLLMNRMDLSLLVITKGL---TKTGDDYEVKAAHVELAE 2722
Query: 1808 AMRV-DRRAEPRYAERIPYVVIHGEPGARLVDMVVDPLEVLAIDSPFRINDLYYINKQII 1866
MR D P +R+PYV+I GA+ + DP+ VL + P I+ YY+ QI
Sbjct: 2723 RMRKRDAATAPNVGDRVPYVIIKAAKGAKAYERSEDPIYVLENNIP--IDPHYYLENQIS 2896
Query: 1867 PALQRVF 1873
+ R+F
Sbjct: 2897 KPILRIF 2917
Score = 50.8 bits (120), Expect = 6e-06
Identities = 40/158 (25%), Positives = 72/158 (45%), Gaps = 5/158 (3%)
Frame = +2
Query: 32 SHSTFHRGKVNEVPVIRVYGSTPAGQKTCLHIHGALPYLYVSCSDIPLQLDQEGDAYTYM 91
SH V +IR++G T G C ++HG PY Y+ C P + + ++ +
Sbjct: 341 SHGELLPNSSGPVAIIRIFGVTKEGHSVCCNVHGFEPYFYICC---PPGMGPDDISHFH- 508
Query: 92 VASSLEKALKLKGSADSSRQHVHGCSLVRAKK-FYGYCSFEEFFVKIYLRYYPQDVSRAA 150
+LE ++ + + V +V+ + Y S + F+KI + P V+
Sbjct: 509 --QTLEGRMREANRNSNVGKFVRRIEMVQRRSIMYYQQSNSQPFLKIVVA-LPTMVASCR 679
Query: 151 NLLLAGAVLN----KSLQPHESHIPFILQFLVDYNLYG 184
+L G L+ KS +ES++ F L+F++D N+ G
Sbjct: 680 GILDRGIQLDGLGMKSFLTYESNVLFALRFMIDCNIVG 793
>NP004853 nodulin
Length = 1431
Score = 33.1 bits (74), Expect = 1.3
Identities = 17/36 (47%), Positives = 20/36 (55%), Gaps = 6/36 (16%)
Frame = -3
Query: 238 WM-WSLPSESGGSSNDNTHSPKRQSI-----CELEG 267
W+ W LPSES G S NTH RQ + C+ EG
Sbjct: 529 WLGWQLPSESSGPSKRNTHDLARQQVA*RWRCKSEG 422
>BF071560
Length = 397
Score = 32.3 bits (72), Expect = 2.1
Identities = 27/98 (27%), Positives = 39/98 (39%)
Frame = -3
Query: 948 LVDKNLKSHLSLPTFPNNLHLDEDDEMPGNALDVFLPISARNSQKQKKPWNKCVTIETPR 1007
L +K L+ HLS+PT L +D+ LD AR+ PWN C + R
Sbjct: 347 LWEKTLR-HLSIPTCHRRTVLSTEDDSRNPLLD-----QARSKMSPLCPWNSCKGFDDMR 186
Query: 1008 SSGTKGVSTYYQNDGSHLYLLTPNILPPSAGSVQRWLF 1045
++ D + +LT L S G+ LF
Sbjct: 185 KVFGGTITQVASEDLLNFSVLTNFSLSTSRGATHEKLF 72
>TC218803 similar to UP|Q84P72 (Q84P72) Receptor-like protein kinase-like
protein (Fragment), partial (35%)
Length = 1266
Score = 32.0 bits (71), Expect = 2.8
Identities = 16/37 (43%), Positives = 19/37 (51%), Gaps = 3/37 (8%)
Frame = -1
Query: 54 PAGQKTCLHIHGAL---PYLYVSCSDIPLQLDQEGDA 87
P K C HIHG P + SCS +P L +GDA
Sbjct: 816 PFFDKRCQHIHGMTHIWPLAWFSCSAVPC*LSNDGDA 706
>TC217115 similar to GB|CAD41408.1|21741286|OSJN00110 OSJNBb0078D11.16 {Oryza
sativa (japonica cultivar-group);} , partial (8%)
Length = 777
Score = 31.6 bits (70), Expect = 3.7
Identities = 28/115 (24%), Positives = 49/115 (42%)
Frame = -1
Query: 559 SEYKRASENHLLHNTDTSTVNTDKRNRQWGSLPFSMTGKVNNDGEHATSQVAHLFERETE 618
S ++ S +++ N+D T T + + W S +M ++N+ + SQ L
Sbjct: 723 SNIRKCSRHYIHSNSDDQTFQTSEMTKNWQSFSVTMFRYISNELSQSKSQKILLILYNQN 544
Query: 619 DSSLSDYLTRNEIKNNKYIKRNVGEGASDSKEVHSLVNCSLRDLMRRKRSYRVEH 673
S S + +K+N ++ R G SD E S +CSL + R+ H
Sbjct: 543 IFSCSLSM*SA*LKSNFHLIR----GCSDEAEGMS-SSCSLMTIS*RRTKQAQYH 394
>TC209602 similar to UP|Q940G9 (Q940G9) Periaxin-like protein, partial (20%)
Length = 744
Score = 31.2 bits (69), Expect = 4.8
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = +3
Query: 403 LHCVKMPNCLRKGPTQVSSC*KLVKCNQLKRLKSQNLLKWL 443
L+C+K NCL + SC + V C LKS N L+++
Sbjct: 282 LNCLKYLNCLNPSCLKYLSCLRFVNCLSPSFLKSPNYLRFM 404
>TC234887 weakly similar to UP|Q6H7K3 (Q6H7K3) Wall-associated kinase-like,
partial (10%)
Length = 592
Score = 30.8 bits (68), Expect = 6.2
Identities = 18/62 (29%), Positives = 31/62 (49%), Gaps = 14/62 (22%)
Frame = +3
Query: 1702 LITKKRYVGYSYENP--------------NQIEPVFDAKGIETVRRDTCEAVAKIMEQSL 1747
LIT ++ + + YE+ NQ+ +FDA+ ++ R+D AVA + + L
Sbjct: 72 LITGRKPISFLYEDEGQNLVAQFISLMKKNQVSEIFDARVLKDARKDDILAVANLAMRCL 251
Query: 1748 RL 1749
RL
Sbjct: 252 RL 257
>TC229396 similar to UP|Q9LVG4 (Q9LVG4) Emb|CAB86035.1 (Pseudo-response
regulator 3), partial (28%)
Length = 1013
Score = 30.4 bits (67), Expect = 8.1
Identities = 23/120 (19%), Positives = 50/120 (41%), Gaps = 9/120 (7%)
Frame = +1
Query: 470 AATIDKMLEKANMAYESESQQECQDILDSIDDM---------LALDLPKKKPSRSFDHNC 520
A + EK + + +++EC +++D +DD+ ++L++ + P + N
Sbjct: 610 AQVMQTKTEKVSSRWVHATEKECHELIDPLDDVAMGKDLAMGISLNMRLEHPLKELSCN- 786
Query: 521 PIEASSDNMIPQVDGSNDDEFSSPCASLAEISSAVEINSEYKRASENHLLHNTDTSTVNT 580
P+ N + VD + S ++ E N + R EN ++ D + N+
Sbjct: 787 PMMGKGANKMSDVDDMQIIKRQSNACEKGQL----EYNGDKTRTRENQAMNVIDVTDSNS 954
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 91,496,601
Number of Sequences: 63676
Number of extensions: 1339676
Number of successful extensions: 6312
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 6212
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6304
length of query: 2018
length of database: 12,639,632
effective HSP length: 112
effective length of query: 1906
effective length of database: 5,507,920
effective search space: 10498095520
effective search space used: 10498095520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146571.7


