

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145221.10 - phase: 0
(68 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
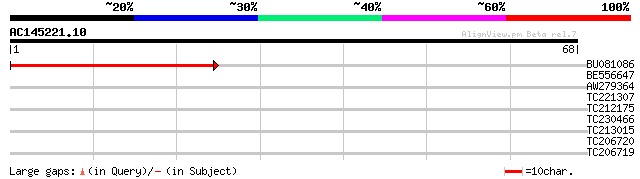
Score E
Sequences producing significant alignments: (bits) Value
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 44 2e-05
BE556647 similar to GP|1763230|gb|A 3-hydroxy-3-methylglutaryl c... 28 0.91
AW279364 similar to GP|29420787|db L-asparaginase {Glycine max},... 27 2.0
TC221307 similar to UP|Q94A51 (Q94A51) AT3g49060/T2J13_100, part... 26 4.5
TC212175 weakly similar to UP|Q9M3Y6 (Q9M3Y6) 68 kDa protein, pa... 25 5.9
TC230466 similar to UP|Q8RY60 (Q8RY60) At1g47330/T3F24_2, partia... 25 5.9
TC213015 UP|O07902 (O07902) Cytochrome c oxidase subunit III (Fr... 25 5.9
TC206720 25 7.7
TC206719 similar to UP|2ACA_HUMAN (Q06190) Serine/threonine prot... 25 7.7
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 43.5 bits (101), Expect = 2e-05
Identities = 20/25 (80%), Positives = 21/25 (84%)
Frame = +2
Query: 1 MTINKSQGQSLKHDGVYLPTPVFSH 25
MTINKSQGQ L G+YLPTPVFSH
Sbjct: 350 MTINKSQGQLLASVGLYLPTPVFSH 424
>BE556647 similar to GP|1763230|gb|A 3-hydroxy-3-methylglutaryl coenzyme a
reductase {Camptotheca acuminata}, partial (16%)
Length = 286
Score = 28.1 bits (61), Expect = 0.91
Identities = 16/39 (41%), Positives = 24/39 (61%)
Frame = +3
Query: 8 GQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREWLKILII 46
G SLK + +P +FS GQ+++ SKV S WL ++I
Sbjct: 39 GASLKIL*IVIPCHLFSTGQVYLQGSKVFSVLWLGRMLI 155
>AW279364 similar to GP|29420787|db L-asparaginase {Glycine max}, partial
(37%)
Length = 370
Score = 26.9 bits (58), Expect = 2.0
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = +1
Query: 8 GQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREW 40
G+ KH+ V LP P + ++ +VT+ EW
Sbjct: 73 GKGSKHNPV*LPNPNIGVRDVRILCGEVTANEW 171
>TC221307 similar to UP|Q94A51 (Q94A51) AT3g49060/T2J13_100, partial (6%)
Length = 629
Score = 25.8 bits (55), Expect = 4.5
Identities = 9/23 (39%), Positives = 16/23 (69%)
Frame = -2
Query: 17 YLPTPVFSHGQLHVVVSKVTSRE 39
YLP+P+F G +V S ++++E
Sbjct: 376 YLPSPIFQDGSKSLVASFISTKE 308
>TC212175 weakly similar to UP|Q9M3Y6 (Q9M3Y6) 68 kDa protein, partial (33%)
Length = 990
Score = 25.4 bits (54), Expect = 5.9
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = -1
Query: 11 LKHDGVYLPTPVFSHG 26
L H+G+YLP FSHG
Sbjct: 612 LCHNGLYLPIANFSHG 565
>TC230466 similar to UP|Q8RY60 (Q8RY60) At1g47330/T3F24_2, partial (13%)
Length = 704
Score = 25.4 bits (54), Expect = 5.9
Identities = 15/41 (36%), Positives = 24/41 (57%)
Frame = +2
Query: 19 PTPVFSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETSNI 59
P PVF + VVV +T + ++ L+ +E ++TDE NI
Sbjct: 191 PLPVFPPNE--VVVGVITMEDVIEELLQEEILDETDEYVNI 307
>TC213015 UP|O07902 (O07902) Cytochrome c oxidase subunit III (Fragment),
partial (5%)
Length = 465
Score = 25.4 bits (54), Expect = 5.9
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = -2
Query: 11 LKHDGVYLPTPVFSHG 26
L H+G+YLP FSHG
Sbjct: 50 LCHNGLYLPIANFSHG 3
>TC206720
Length = 693
Score = 25.0 bits (53), Expect = 7.7
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +2
Query: 38 REWLKILIIDEDGEDTDETSNIVYEE 63
RE+++ L ++EDGED S V+EE
Sbjct: 218 REYIR-LSMEEDGEDMSNASGDVWEE 292
>TC206719 similar to UP|2ACA_HUMAN (Q06190) Serine/threonine protein
phosphatase 2A, 72/130 kDa regulatory subunit B (PP2A,
subunit B, B''-PR72/PR130) (PP2A, subunit B, B72/B130
isoforms) (PP2A, subunit B, PR72/PR130 isoforms) (PP2A,
subunit B, R3 isoform) , partial (3%)
Length = 1892
Score = 25.0 bits (53), Expect = 7.7
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +2
Query: 38 REWLKILIIDEDGEDTDETSNIVYEE 63
RE+++ L ++EDGED S V+EE
Sbjct: 1589 REYIR-LSMEEDGEDMSNASGDVWEE 1663
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,873,833
Number of Sequences: 63676
Number of extensions: 30869
Number of successful extensions: 123
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 123
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 123
length of query: 68
length of database: 12,639,632
effective HSP length: 44
effective length of query: 24
effective length of database: 9,837,888
effective search space: 236109312
effective search space used: 236109312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC145221.10


