

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
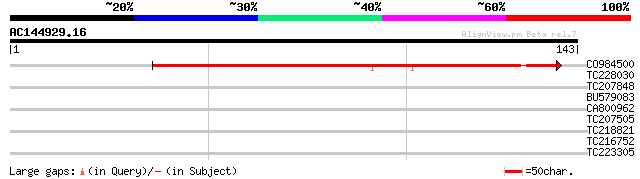
Query= AC144929.16 + phase: 0
(143 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CO984500 96 8e-21
TC228030 similar to UP|Q8LPS4 (Q8LPS4) At3g06670/T8E24.10, parti... 28 1.1
TC207848 homologue to UP|Q84RS0 (Q84RS0) ZR4 protein (Fragment),... 27 3.3
BU579083 27 4.3
CA800962 26 5.7
TC207505 26 5.7
TC218821 weakly similar to UP|Q7PBB4 (Q7PBB4) NADH dehydrogenase... 26 7.4
TC216752 similar to UP|Q8LA49 (Q8LA49) Globulin-like protein, pa... 25 9.7
TC223305 similar to GB|AAQ89640.1|37202050|BT010618 At4g09550 {A... 25 9.7
>CO984500
Length = 758
Score = 95.5 bits (236), Expect = 8e-21
Identities = 62/124 (50%), Positives = 76/124 (61%), Gaps = 21/124 (16%)
Frame = -1
Query: 37 SFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVR------ 90
S+ S +TLS K+ DG + ED NK +CPLQ YLLGSSFE+ E EVR ERAS+
Sbjct: 755 SYVSIITLSEKEIDGPKAEDYRNKAVCPLQGYLLGSSFELPETKEVRKERASLAELLYKT 576
Query: 91 --------------ETQVKQGRRS-ALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEK 135
ETQVK+ +S A+HI++K+ V SSSKSCNT N AD A STN+K
Sbjct: 575 KSTSQDCIDTGIQGETQVKRTPKSAAMHIMRKILKRVHSSSKSCNTSRNDADSA-STNKK 399
Query: 136 LCKV 139
L KV
Sbjct: 398 LQKV 387
>TC228030 similar to UP|Q8LPS4 (Q8LPS4) At3g06670/T8E24.10, partial (12%)
Length = 1344
Score = 28.5 bits (62), Expect = 1.1
Identities = 15/34 (44%), Positives = 19/34 (55%)
Frame = -1
Query: 12 KLISYELEKFVEADEECFYESSGRISFRSNVTLS 45
K +S +LE F D CF+ SSG +SF N S
Sbjct: 378 KTLSLDLEFF---DNFCFFTSSGSLSFEGNFLFS 286
>TC207848 homologue to UP|Q84RS0 (Q84RS0) ZR4 protein (Fragment), partial
(92%)
Length = 1403
Score = 26.9 bits (58), Expect = 3.3
Identities = 15/50 (30%), Positives = 29/50 (58%), Gaps = 1/50 (2%)
Frame = +1
Query: 11 LKLISYELEKFVEADEECFY-ESSGRISFRSNVTLSRKKTDGLETEDLGN 59
LK + + ++F E E ++ E+ GR+ + NV + K + G+ +EDL +
Sbjct: 862 LKRVRFSRKRFSEKQAEQWWAENRGRVYEQYNVRMIDKSSVGVGSEDLAH 1011
>BU579083
Length = 376
Score = 26.6 bits (57), Expect = 4.3
Identities = 14/34 (41%), Positives = 18/34 (52%)
Frame = -2
Query: 12 KLISYELEKFVEADEECFYESSGRISFRSNVTLS 45
K +S +LE F D CF+ SSG +S N S
Sbjct: 369 KTLSLDLEFF---DNFCFFSSSGSLSLEGNFLFS 277
>CA800962
Length = 409
Score = 26.2 bits (56), Expect = 5.7
Identities = 13/24 (54%), Positives = 17/24 (70%)
Frame = +3
Query: 2 KDAAVTETQLKLISYELEKFVEAD 25
K+ VTE LKLI+ ELEK + A+
Sbjct: 333 KEDEVTENDLKLINDELEKVLGAE 404
>TC207505
Length = 1191
Score = 26.2 bits (56), Expect = 5.7
Identities = 17/50 (34%), Positives = 20/50 (40%), Gaps = 5/50 (10%)
Frame = -2
Query: 98 RRSALHIIKKMSNMV-----LSSSKSCNTYGNTADHATSTNEKLCKVGPY 142
R LH SN S S S N NT + T + LCK+ PY
Sbjct: 533 RTRLLHFAPTSSNSYGSIPQASVSSSSNKLDNTEQESGPTCKTLCKLSPY 384
>TC218821 weakly similar to UP|Q7PBB4 (Q7PBB4) NADH dehydrogenase I chain N,
partial (8%)
Length = 1164
Score = 25.8 bits (55), Expect = 7.4
Identities = 28/102 (27%), Positives = 44/102 (42%)
Frame = -3
Query: 20 KFVEADEECFYESSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREK 79
K VE D C Y + ISFR L KK L K IC L +S + R K
Sbjct: 892 KVVEKDASCLYHNVCMISFRKK--LIYKKGTIL-------KFIC------LFASSQTRLK 758
Query: 80 TEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNT 121
T + I +++ + ++ R + K+ +++LS+ T
Sbjct: 757 T*INIWLLILQQNKQERRRLCVAKLH*KLEDIILSNKNKLLT 632
>TC216752 similar to UP|Q8LA49 (Q8LA49) Globulin-like protein, partial (53%)
Length = 748
Score = 25.4 bits (54), Expect = 9.7
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -1
Query: 106 KKMSNMVLSSSKSCNTYGNTADHATST 132
KKM++ VL SC+T G+T A T
Sbjct: 577 KKMASAVLFERNSCSTSGSTLKAACKT 497
>TC223305 similar to GB|AAQ89640.1|37202050|BT010618 At4g09550 {Arabidopsis
thaliana;} , partial (82%)
Length = 559
Score = 25.4 bits (54), Expect = 9.7
Identities = 11/36 (30%), Positives = 22/36 (60%)
Frame = -3
Query: 18 LEKFVEADEECFYESSGRISFRSNVTLSRKKTDGLE 53
L +FV+ + CF +G+ F S ++L+ + +G+E
Sbjct: 164 LVQFVQPPDPCFKLRTGQSMFSSTISLN*RGHNGIE 57
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.128 0.345
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,109,421
Number of Sequences: 63676
Number of extensions: 50194
Number of successful extensions: 227
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 226
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 226
length of query: 143
length of database: 12,639,632
effective HSP length: 88
effective length of query: 55
effective length of database: 7,036,144
effective search space: 386987920
effective search space used: 386987920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144929.16


