

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
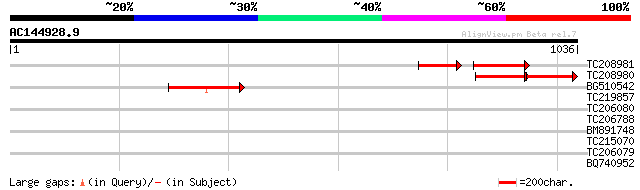
Query= AC144928.9 - phase: 2 /pseudo
(1036 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC208981 similar to UP|Q9FF46 (Q9FF46) ABC transporter-like prot... 135 8e-62
TC208980 similar to UP|Q9FF46 (Q9FF46) ABC transporter-like prot... 133 3e-57
BG510542 weakly similar to GP|9759338|dbj ABC transporter-like p... 127 2e-29
TC219857 similar to UP|Q9LVY7 (Q9LVY7) Cytochrome P450-like, par... 33 0.85
TC206080 weakly similar to UP|CSE1_ARATH (Q9ZPY7) Importin-alpha... 31 3.2
TC206788 similar to UP|Q940Y6 (Q940Y6) AT4g27600/T29A15_90, part... 30 5.5
BM891748 homologue to GP|15010748|gb| AT4g30260/F9N11_110 {Arabi... 30 7.2
TC215070 similar to UP|Q40336 (Q40336) Proline-rich cell wall pr... 30 7.2
TC206079 weakly similar to UP|CSE1_ARATH (Q9ZPY7) Importin-alpha... 29 9.3
BQ740952 29 9.3
>TC208981 similar to UP|Q9FF46 (Q9FF46) ABC transporter-like protein, partial
(16%)
Length = 555
Score = 135 bits (340), Expect(2) = 8e-62
Identities = 75/102 (73%), Positives = 80/102 (77%)
Frame = +1
Query: 848 LFYAK*RP*DHFLWINCTTGGRVILA*AA*LIFSRKTRWIILIQ*SNQWCIYLCSIS*PI 907
LF AK*RP*DHFLWIN T GGRVILA*AA LI S KT+ IILI *S+ WCI+LCSIS PI
Sbjct: 235 LFCAK*RP*DHFLWINYTIGGRVILA*AAWLISSLKTQLIILIH*SSLWCIFLCSISSPI 414
Query: 908 QDLLSQITILCCFVLYTV*LA*HMHYP*FLNLAQRSCTSSSC 949
+D L QI +LCCFVLYTV LA* MH P FLNLAQ SC C
Sbjct: 415 RDQLLQIIMLCCFVLYTVSLA*PMHCPYFLNLAQPSCGQYFC 540
Score = 121 bits (304), Expect(2) = 8e-62
Identities = 57/78 (73%), Positives = 61/78 (78%)
Frame = +2
Query: 748 KT*FFKIQGFIQPENSWCIQAI*VLSYTGWEAAITRSQDPGRRLSYFATCWSMLRINYQI 807
KT FF IQGFI P NSWC QAI VLSYTGWEAA TRS DP LSYF TCWS+LRI +QI
Sbjct: 2 KTQFF*IQGFI*PGNSWCFQAIQVLSYTGWEAATTRS*DPSH*LSYFVTCWSLLRITFQI 181
Query: 808 E*SNIWCIWLHLYGDCCI 825
+* NIWC W+H YGD CI
Sbjct: 182 K*PNIWCSWVHPYGDWCI 235
>TC208980 similar to UP|Q9FF46 (Q9FF46) ABC transporter-like protein, partial
(15%)
Length = 820
Score = 133 bits (335), Expect(2) = 3e-57
Identities = 68/92 (73%), Positives = 76/92 (81%)
Frame = +3
Query: 945 TSSSCFDSHCNTTKR**NFKGYS*FMLL*VGFTSIGHCKC*KVPRSVAHNSLRFTFKKWL 1004
TS+SCFDSHCNTTKR**+F+ YS*FMLL*VGFTS+ CKC*KV SVAHNSL FTF+ WL
Sbjct: 306 TSASCFDSHCNTTKR**SFEEYS*FMLL*VGFTSLSCCKC*KVSGSVAHNSLWFTFENWL 485
Query: 1005 *SS*LESLHIHPHSYGCYWSCNSVFLYGHLQK 1036
*SS LE +HI+PHS GC S+FLYGHL K
Sbjct: 486 *SSRLEFVHIYPHSNGCNLPGYSIFLYGHLPK 581
Score = 108 bits (270), Expect(2) = 3e-57
Identities = 63/99 (63%), Positives = 68/99 (68%)
Frame = +2
Query: 851 AK*RP*DHFLWINCTTGGRVILA*AA*LIFSRKTRWIILIQ*SNQWCIYLCSIS*PIQDL 910
AK RP D WIN T GRVILA*AA LI S KT+ I I *S+ WCI+LCSIS PI+
Sbjct: 17 AKXRPXDXXXWINYTMEGRVILA*AAWLISSLKTQLIXXIH*SSLWCIFLCSISSPIRYQ 196
Query: 911 LSQITILCCFVLYTV*LA*HMHYP*FLNLAQRSCTSSSC 949
L QI +LCCFVLYTV LA* MH P FLNL Q SC C
Sbjct: 197 LLQIIMLCCFVLYTVSLA*PMHCPYFLNLVQPSCGQYFC 313
>BG510542 weakly similar to GP|9759338|dbj ABC transporter-like protein
{Arabidopsis thaliana}, partial (6%)
Length = 439
Score = 127 bits (320), Expect = 2e-29
Identities = 84/145 (57%), Positives = 95/145 (64%), Gaps = 6/145 (4%)
Frame = -3
Query: 291 S*NFESRDF*NRC*IVAAFTT--FKHGCKFLCSAN*KGKNSKWTNAHDA*N*K*PAC*L* 348
S*NFES +F*+R *IV F T FKHGCKF+ A KGK + W+ A + *N*K* *
Sbjct: 437 S*NFESSNF*SRY*IVVTFRTYNFKHGCKFISGAKGKGKRT*WSYADNT*N*K*SRY**P 258
Query: 349 SKHRKRN*T*----KCYKGKTATN*HTNF*VCICSA*KRKGSATRKQEPYLLRSTKNGYK 404
+R RN KC KGKT T NF*VCI S *KR+G ATRKQE YLL NGYK
Sbjct: 257 PSYRNRNQRYRC*SKCAKGKTTTYS*PNF*VCIFST*KREG*ATRKQEAYLLGGN*NGYK 78
Query: 405 Y*EK*EAFY*DFFQRSDSYTESSKQ 429
+*+K +AF DFFQ SDSY ESSKQ
Sbjct: 77 H*KKEKAFNGDFFQGSDSYIESSKQ 3
>TC219857 similar to UP|Q9LVY7 (Q9LVY7) Cytochrome P450-like, partial (36%)
Length = 1146
Score = 32.7 bits (73), Expect = 0.85
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = -3
Query: 441 TWPYHRYHGPVRGWQDNISFSSSWKSTWMLGYWFNSDKW 479
TWP ++H +R W SSS+KS W+ W W
Sbjct: 673 TWPSTKWHISIRSW------SSSFKSAWIKFLWLREVLW 575
>TC206080 weakly similar to UP|CSE1_ARATH (Q9ZPY7) Importin-alpha re-exporter
(Cellular apoptosis susceptibility protein homolog),
partial (22%)
Length = 823
Score = 30.8 bits (68), Expect = 3.2
Identities = 20/55 (36%), Positives = 31/55 (56%), Gaps = 7/55 (12%)
Frame = -3
Query: 16 FLSSCIKKTKGDISNRL-CTAAEVKFYLNSLM-----EKSSSANYLKP-NRNCNL 63
FLSS +KK KG R+ C V+F LNS++ E + ++L P +R+C +
Sbjct: 467 FLSSIVKKNKGGAITRIFCYIWHVRFLLNSILFWPRKECDYAVHHLTPGHRSCRI 303
>TC206788 similar to UP|Q940Y6 (Q940Y6) AT4g27600/T29A15_90, partial (58%)
Length = 1182
Score = 30.0 bits (66), Expect = 5.5
Identities = 29/112 (25%), Positives = 48/112 (41%), Gaps = 4/112 (3%)
Frame = -1
Query: 99 CRACCEGFFCPHGITCMIPCPLGSYCPIATLNKTTG---VCEPYLYQLPPMQPNHTCGGA 155
CR C G+ IPC S + K++G +C+ ++ +P P
Sbjct: 1161 CRTICNGY*--RNKVAQIPCS*SSTFLCFKITKSSGLISICKD*IHTIPNNFPEIIKVSL 988
Query: 156 NVWADFSSSSETFCSAGSYCPTTTTKFP-CSSGLRKAEFM*F*HSNPKHACI 206
N D S +C+ G+ CP T F C +G+RK + + F + K+ C+
Sbjct: 987 NA-GDI*SC---YCNQGTVCPCFLTCFGYCFNGIRKFK*ISF---DDKYVCL 853
>BM891748 homologue to GP|15010748|gb| AT4g30260/F9N11_110 {Arabidopsis
thaliana}, partial (6%)
Length = 350
Score = 29.6 bits (65), Expect = 7.2
Identities = 11/19 (57%), Positives = 16/19 (83%)
Frame = -2
Query: 984 C*KVPRSVAHNSLRFTFKK 1002
C* VPR + H+++RFTF+K
Sbjct: 343 C*NVPRVMIHDTIRFTFRK 287
>TC215070 similar to UP|Q40336 (Q40336) Proline-rich cell wall protein,
partial (82%)
Length = 1597
Score = 29.6 bits (65), Expect = 7.2
Identities = 17/63 (26%), Positives = 27/63 (41%), Gaps = 2/63 (3%)
Frame = -1
Query: 442 WPYHRYHGPVRGWQDNISFSSSW--KSTWMLGYWFNSDKWKE*INSFL*ENYWLRATR*C 499
W +H + W +N F + W W +WF + W IN++ N W R C
Sbjct: 787 WCWHNW------WINNWWFHNGWFWHIRWCNNWWFWHNWW---INNWWFHNGWFWHIRWC 635
Query: 500 SSW 502
++W
Sbjct: 634 NNW 626
>TC206079 weakly similar to UP|CSE1_ARATH (Q9ZPY7) Importin-alpha re-exporter
(Cellular apoptosis susceptibility protein homolog),
partial (10%)
Length = 475
Score = 29.3 bits (64), Expect = 9.3
Identities = 15/32 (46%), Positives = 20/32 (61%), Gaps = 1/32 (3%)
Frame = -3
Query: 16 FLSSCIKKTKGDISNRL-CTAAEVKFYLNSLM 46
FLSS +KK KG R+ C V+F LNS++
Sbjct: 143 FLSSIVKKNKGGAITRIFCYIWHVRFLLNSIL 48
>BQ740952
Length = 429
Score = 29.3 bits (64), Expect = 9.3
Identities = 10/18 (55%), Positives = 10/18 (55%)
Frame = -1
Query: 66 WVSGCEPGWACSVPSGQK 83
WV GC P W C GQK
Sbjct: 399 WVEGCRPQWCCVQRRGQK 346
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.354 0.153 0.599
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,192,581
Number of Sequences: 63676
Number of extensions: 1184075
Number of successful extensions: 16143
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 8843
Number of HSP's successfully gapped in prelim test: 687
Number of HSP's that attempted gapping in prelim test: 6701
Number of HSP's gapped (non-prelim): 10609
length of query: 1036
length of database: 12,639,632
effective HSP length: 107
effective length of query: 929
effective length of database: 5,826,300
effective search space: 5412632700
effective search space used: 5412632700
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144928.9


