

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.8 + phase: 0 /pseudo
(359 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
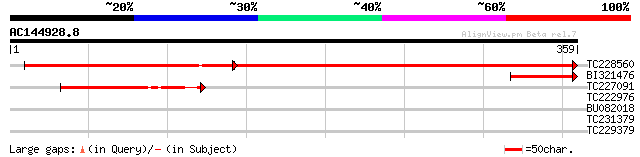
TC228560 similar to UP|Q9ST64 (Q9ST64) ATPase (Fragment), partia... 315 e-145
BI321476 64 8e-11
TC227091 similar to UP|Q7Q6L1 (Q7Q6L1) AgCP5440 (Fragment), part... 64 8e-11
TC222976 similar to UP|ARS1_HUMAN (O43681) Arsenical pump-drivin... 32 0.58
BU082018 weakly similar to GP|23093517|gb CG32133-PA {Drosophila... 31 0.76
TC231379 similar to UP|Q9FI52 (Q9FI52) Nucleotide-binding protei... 31 0.76
TC229379 similar to UP|O49472 (O49472) ATP binding protein-like,... 29 3.8
>TC228560 similar to UP|Q9ST64 (Q9ST64) ATPase (Fragment), partial (82%)
Length = 1503
Score = 315 bits (807), Expect(2) = e-145
Identities = 164/217 (75%), Positives = 180/217 (82%)
Frame = +3
Query: 143 VRRAKTGRVTGYTSSWTG*SYCDFQGYTIS*ITGI*YVHSNCI*YGTYWSYITFIVLA*F 202
V R KTGR+TG+TSSWTG*SYCD QGYTIS ITGI*+V+SN I*Y TY S+IT IVL F
Sbjct: 603 VGRTKTGRITGFTSSWTG*SYCDLQGYTISRITGI*HVYSNSI*YSTYGSHITIIVLTRF 782
Query: 203 LGCIHREDTKAKTKDSFCYLCN*ICVWARKSSSRCY*QIGEA*GEDDKSARAFS*H*FDR 262
LGCIH EDT+AKTK+SFCYL N*I VWAR++ + C *QIGE EDDKSAR S*+*F+R
Sbjct: 783 LGCIHWEDTEAKTKNSFCYLSN*IRVWARRNPTECC*QIGETQREDDKSARTLS*Y*FNR 962
Query: 263 ICYCYNPHGDGS**VIQIECLIEERKCSCEETHCQSTSSTICI*LQVLCNEKKGSNTCS* 322
IC+CYNPHGDG+**VI IEC EE KCSCEET+CQ SSTI I*LQVLC EKKGSN CS*
Sbjct: 963 ICHCYNPHGDGN**VI*IECFPEEGKCSCEETYCQPDSSTIYI*LQVLCYEKKGSNACS* 1142
Query: 323 YDSE*SRTLQPINDPSTTC*C*DKRCSCSQISG*HYL 359
Y SE SRTLQ I+DP TT C*DKRCS SQISG*HYL
Sbjct: 1143YSSERSRTLQLIDDPGTTHRC*DKRCSSSQISG*HYL 1253
Score = 218 bits (554), Expect(2) = e-145
Identities = 107/135 (79%), Positives = 122/135 (90%)
Frame = +1
Query: 10 VKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
V+S+A TE+ S FD+MV+GT+RKYYMLGGKGGV KTSCAASLAVKFANNGHPTLVVSTD
Sbjct: 217 VRSIAGTTEAASGFDEMVSGTKRKYYMLGGKGGVRKTSCAASLAVKFANNGHPTLVVSTD 396
Query: 70 PAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMD 129
PAHSLSDSFAQDLTGGALV V+GPDYPLFALEINPEK RE+F++ A++ GG TGVKDFMD
Sbjct: 397 PAHSLSDSFAQDLTGGALVPVEGPDYPLFALEINPEKFREEFQNAAQKKGG-TGVKDFMD 573
Query: 130 GMGLGMIVDQTAKVR 144
GMGLGMI DQ +++
Sbjct: 574 GMGLGMIADQLGELK 618
>BI321476
Length = 455
Score = 64.3 bits (155), Expect = 8e-11
Identities = 34/42 (80%), Positives = 34/42 (80%)
Frame = +3
Query: 318 NTCS*YDSE*SRTLQPINDPSTTC*C*DKRCSCSQISG*HYL 359
N CS*Y SE*SRTLQ INDP TT C*DKRCS SQIS *HYL
Sbjct: 3 NACS*YSSE*SRTLQFINDPGTTRRC*DKRCSSSQISW*HYL 128
>TC227091 similar to UP|Q7Q6L1 (Q7Q6L1) AgCP5440 (Fragment), partial (58%)
Length = 1610
Score = 64.3 bits (155), Expect = 8e-11
Identities = 37/92 (40%), Positives = 56/92 (60%)
Frame = +2
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ +GGKGGVGKT+C++ L++ A L++STDPAH+LSD+F Q T + V+G
Sbjct: 335 KWIFVGGKGGVGKTTCSSILSILLATVRSSVLIISTDPAHNLSDAFQQRFTKTPTL-VNG 511
Query: 93 PDYPLFALEINPEKAREDFRDVAKQNGGSTGV 124
L+A+E++P ED GG+ G+
Sbjct: 512 FS-NLYAMEVDPTVEHEDM-------GGADGM 583
>TC222976 similar to UP|ARS1_HUMAN (O43681) Arsenical pump-driving ATPase
(Arsenite-translocating ATPase) (Arsenical resistance
ATPase) (Arsenite-transporting ATPase) (ARSA) (ASNA-I) ,
partial (9%)
Length = 425
Score = 31.6 bits (70), Expect = 0.58
Identities = 15/37 (40%), Positives = 22/37 (58%), Gaps = 8/37 (21%)
Frame = +3
Query: 22 VFDDMVAGTER--------KYYMLGGKGGVGKTSCAA 50
+ D V GT + K+ +GGKGGVGKT+C++
Sbjct: 312 IMGDQVEGTVQNVLEQETLKWVFVGGKGGVGKTTCSS 422
>BU082018 weakly similar to GP|23093517|gb CG32133-PA {Drosophila
melanogaster}, partial (1%)
Length = 421
Score = 31.2 bits (69), Expect = 0.76
Identities = 14/30 (46%), Positives = 17/30 (56%)
Frame = +3
Query: 58 NNGHPTLVVSTDPAHSLSDSFAQDLTGGAL 87
+NG+PT P HS+ DSF QD T L
Sbjct: 219 SNGNPTTTAIVSPLHSILDSFPQDDTSHLL 308
>TC231379 similar to UP|Q9FI52 (Q9FI52) Nucleotide-binding protein, partial
(59%)
Length = 653
Score = 31.2 bits (69), Expect = 0.76
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = +2
Query: 16 PTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFA 57
P + + +A + K +L GKGGVGK++ +A LA A
Sbjct: 212 PDPDLVAIAERMATVKHKILVLSGKGGVGKSTFSAQLAFALA 337
>TC229379 similar to UP|O49472 (O49472) ATP binding protein-like, partial
(73%)
Length = 1527
Score = 28.9 bits (63), Expect = 3.8
Identities = 13/31 (41%), Positives = 19/31 (60%)
Frame = +2
Query: 27 VAGTERKYYMLGGKGGVGKTSCAASLAVKFA 57
+ G + + GKGGVGK++ A +LAV A
Sbjct: 146 IDGVKNTIAVASGKGGVGKSTTAVNLAVALA 238
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.347 0.152 0.531
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,188,107
Number of Sequences: 63676
Number of extensions: 268912
Number of successful extensions: 2011
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1995
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2008
length of query: 359
length of database: 12,639,632
effective HSP length: 98
effective length of query: 261
effective length of database: 6,399,384
effective search space: 1670239224
effective search space used: 1670239224
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144928.8


