

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144723.3 + phase: 0
(127 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
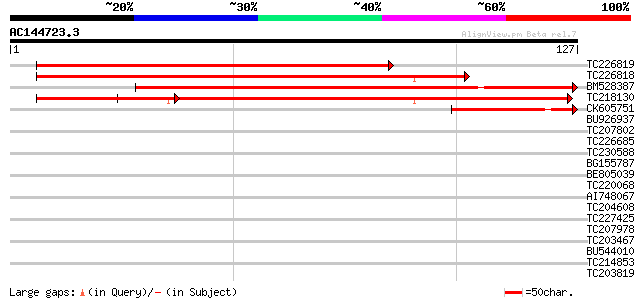
Score E
Sequences producing significant alignments: (bits) Value
TC226819 homologue to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, p... 167 1e-42
TC226818 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, par... 156 2e-39
BM528387 112 5e-26
TC218130 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, par... 101 8e-23
CK605751 43 4e-05
BU926937 27 1.9
TC207802 similar to UP|Q6R3L0 (Q6R3L0) Metal-nicotianamine trans... 27 1.9
TC226685 similar to UP|Q9FRY9 (Q9FRY9) Response regulator 6, par... 27 3.3
TC230588 weakly similar to UP|O14047 (O14047) SPAC2C4.14c protei... 27 3.3
BG155787 weakly similar to GP|28798549|gb 60S ribosomal protein ... 27 3.3
BE805039 26 4.3
TC220068 similar to GB|AAL31246.1|16974485|AY061919 AT4g32480/F8... 26 4.3
AI748067 similar to GP|28874734|emb progesterone 5-beta-reductas... 26 4.3
TC204608 similar to UP|Q9LVI4 (Q9LVI4) Gb|AAF09073.1 (AT3g17860/... 26 4.3
TC227425 similar to UP|Q8HWA9 (Q8HWA9) Mus musculus 0 day neonat... 26 5.6
TC207978 weakly similar to UP|Q6YU70 (Q6YU70) Cyclin G-associate... 26 5.6
TC203467 similar to UP|O82462 (O82462) Glutamyl-tRNA synthetase ... 25 7.3
BU544010 similar to GP|22795261|gb| nodulin-like protein {Oryza ... 25 7.3
TC214853 similar to UP|STO_ARATH (Q96288) Salt-tolerance protein... 25 7.3
TC203819 25 7.3
>TC226819 homologue to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, partial (67%)
Length = 646
Score = 167 bits (423), Expect = 1e-42
Identities = 72/80 (90%), Positives = 78/80 (97%)
Frame = +1
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A GCSVNFEFLNYT+ITSKCKGP+YPPKECCG+FKEFACPYADV+NDLTN+CASTMFSY
Sbjct: 403 AKKGCSVNFEFLNYTVITSKCKGPRYPPKECCGAFKEFACPYADVLNDLTNECASTMFSY 582
Query: 67 INLYGRYPPGLFASECREGK 86
INLYG+YPPGLFASECREGK
Sbjct: 583 INLYGKYPPGLFASECREGK 642
>TC226818 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, partial (65%)
Length = 729
Score = 156 bits (395), Expect = 2e-39
Identities = 71/102 (69%), Positives = 82/102 (79%), Gaps = 5/102 (4%)
Frame = +3
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A C VNFEFLNYT+ITSKCKGP+YPPK+CC +FKEFACPYA+ +ND TNDCA+ MFSY
Sbjct: 186 AKKSCPVNFEFLNYTVITSKCKGPQYPPKQCCAAFKEFACPYANALNDSTNDCAAIMFSY 365
Query: 67 INLYGRYPPGLFASECREGKEGL-----ACDALPPSVSADDT 103
INLYG+YP G FAS CR+GK+G+ A A PPSVS DDT
Sbjct: 366 INLYGKYPFGFFASVCRQGKKGITINCPASAASPPSVSEDDT 491
>BM528387
Length = 319
Score = 112 bits (280), Expect = 5e-26
Identities = 48/99 (48%), Positives = 71/99 (71%)
Frame = +1
Query: 29 GPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLYGRYPPGLFASECREGKEG 88
GP+YPPK CC +FK+FACP+++ +ND+T DCA+ MFSYIN+YG+YPPGLFA++C+EG +G
Sbjct: 1 GPQYPPKVCCEAFKQFACPFSNEVNDMTTDCATVMFSYINIYGKYPPGLFANQCKEGNQG 180
Query: 89 LACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILLF 127
L + P+ ++ +T + + P + T FL LF
Sbjct: 181 LDFFLVTPTNTSSNTIH-VAAPPHFFSLATISAFLGFLF 294
>TC218130 similar to UP|Q9ZWN0 (Q9ZWN0) GPI-anchored protein, partial (54%)
Length = 595
Score = 101 bits (252), Expect = 8e-23
Identities = 52/108 (48%), Positives = 66/108 (60%), Gaps = 6/108 (5%)
Frame = +1
Query: 25 SKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLYGRYPPGLFASECRE 84
+ K P CG + ACPYA+V+NDLTNDCA+ MFS+INLYG+YPPG FAS C E
Sbjct: 178 ANAKEPSTLQATLCGIERVVACPYANVLNDLTNDCATIMFSFINLYGKYPPGFFASMCHE 357
Query: 85 GKEGL------ACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILL 126
GK+G+ A A PPSVSADDT + V V+ F +++
Sbjct: 358 GKKGITINCLPAAAASPPSVSADDTGMKTHQMRVSVTVVVVVYFHVVM 501
Score = 51.6 bits (122), Expect = 1e-07
Identities = 24/39 (61%), Positives = 25/39 (63%), Gaps = 7/39 (17%)
Frame = +3
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPP-------KECC 38
A C VNFEFLNYTIIT KCKG +YPP K CC
Sbjct: 123 AKKSCPVNFEFLNYTIITCKCKGAQYPPSNAVRH*KSCC 239
>CK605751
Length = 724
Score = 42.7 bits (99), Expect = 4e-05
Identities = 22/28 (78%), Positives = 24/28 (85%)
Frame = -3
Query: 100 ADDTANQIVHTPSLVLVLTACIFLILLF 127
ADDTANQ VH PSL+L+L AC FLILLF
Sbjct: 722 ADDTANQAVHCPSLLLMLIAC-FLILLF 642
>BU926937
Length = 451
Score = 27.3 bits (59), Expect = 1.9
Identities = 14/41 (34%), Positives = 19/41 (46%), Gaps = 4/41 (9%)
Frame = -3
Query: 61 STMFSYINLYGRY----PPGLFASECREGKEGLACDALPPS 97
STM IN+ G + ++ S+C G L C PPS
Sbjct: 431 STMMQPINILGNH*VDSASSIYCSQCSMGSRWLCCCKSPPS 309
>TC207802 similar to UP|Q6R3L0 (Q6R3L0) Metal-nicotianamine transporter YSL1,
partial (29%)
Length = 1137
Score = 27.3 bits (59), Expect = 1.9
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = -2
Query: 25 SKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLYG 71
+K K PK+ K+CCG+ ++ P+ +I MF IN YG
Sbjct: 317 AKAKNPKHSCKQCCGNAEK---PWTPMI---------AMFL*INAYG 213
>TC226685 similar to UP|Q9FRY9 (Q9FRY9) Response regulator 6, partial (42%)
Length = 1190
Score = 26.6 bits (57), Expect = 3.3
Identities = 15/46 (32%), Positives = 17/46 (36%), Gaps = 9/46 (19%)
Frame = +2
Query: 4 FHIAPAGCSVNFEFLNYTIITSKCKGP---------KYPPKECCGS 40
F + C V FL T SKC P +PPK GS
Sbjct: 797 FDVVG*SCQVGLSFLTTTKFISKCNNPPGIIFPSAISFPPKNKSGS 934
>TC230588 weakly similar to UP|O14047 (O14047) SPAC2C4.14c protein, partial
(21%)
Length = 893
Score = 26.6 bits (57), Expect = 3.3
Identities = 15/41 (36%), Positives = 20/41 (48%)
Frame = +2
Query: 37 CCGSFKEFACPYADVINDLTNDCASTMFSYINLYGRYPPGL 77
CC +FK F CP A S ++ +I+ Y R P GL
Sbjct: 299 CCSTFKPFICPKATF---------SVLWFFISSYSR-PEGL 391
>BG155787 weakly similar to GP|28798549|gb 60S ribosomal protein {Medicago
sativa}, partial (62%)
Length = 331
Score = 26.6 bits (57), Expect = 3.3
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = +3
Query: 88 GLACDALPPSVSADDTANQIVHTPSLVLV 116
GL D +PP + +IVHT +L+LV
Sbjct: 219 GLLLDTIPPLLEGS*MLGKIVHTVALLLV 305
>BE805039
Length = 427
Score = 26.2 bits (56), Expect = 4.3
Identities = 10/34 (29%), Positives = 23/34 (67%)
Frame = +1
Query: 94 LPPSVSADDTANQIVHTPSLVLVLTACIFLILLF 127
L PS+ + DT + +V + S ++++ AC +++L+
Sbjct: 34 LLPSIVSKDTEHTLVFSKSPLILVVACKLMVILW 135
>TC220068 similar to GB|AAL31246.1|16974485|AY061919 AT4g32480/F8B4_180
{Arabidopsis thaliana;} , partial (10%)
Length = 654
Score = 26.2 bits (56), Expect = 4.3
Identities = 13/39 (33%), Positives = 20/39 (50%), Gaps = 1/39 (2%)
Frame = -2
Query: 10 GCSVNFEFLNYTII-TSKCKGPKYPPKECCGSFKEFACP 47
GCS++F + T++ S + P PP CC S + P
Sbjct: 638 GCSLHFPLPSQTLL*NSAPQSPSPPPSLCCLSHPKNCVP 522
>AI748067 similar to GP|28874734|emb progesterone 5-beta-reductase {Digitalis
purpurea}, partial (13%)
Length = 421
Score = 26.2 bits (56), Expect = 4.3
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = -2
Query: 8 PAGCSVNFEFLNYTIITSKCKGPKYP 33
PAGC+VNF + + TI +C G P
Sbjct: 135 PAGCAVNFPWASETIGFLQCFG*AQP 58
>TC204608 similar to UP|Q9LVI4 (Q9LVI4) Gb|AAF09073.1 (AT3g17860/MEB5_8),
partial (22%)
Length = 1439
Score = 26.2 bits (56), Expect = 4.3
Identities = 10/37 (27%), Positives = 14/37 (37%)
Frame = +3
Query: 29 GPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFS 65
G P + C + E+ C Y + N C FS
Sbjct: 360 GTSIPKESFCNCWSEYECCYCEATTTRRNTCYGASFS 470
>TC227425 similar to UP|Q8HWA9 (Q8HWA9) Mus musculus 0 day neonate cerebellum
cDNA, RIKEN full-length enriched library,
clone:C230098O20 product:histocompatibility 13, full
insert sequence, partial (6%)
Length = 2073
Score = 25.8 bits (55), Expect = 5.6
Identities = 13/30 (43%), Positives = 20/30 (66%), Gaps = 3/30 (10%)
Frame = +2
Query: 101 DDTANQIVHTPS---LVLVLTACIFLILLF 127
DD+ N+IV+ S ++ V+ A FL+LLF
Sbjct: 857 DDSDNEIVNIDSKGAVIFVIAASTFLVLLF 946
>TC207978 weakly similar to UP|Q6YU70 (Q6YU70) Cyclin G-associated
kinase-like protein, partial (8%)
Length = 1099
Score = 25.8 bits (55), Expect = 5.6
Identities = 17/43 (39%), Positives = 19/43 (43%), Gaps = 4/43 (9%)
Frame = -2
Query: 82 CREGKEGLACDALPPSVSADDTANQIVHTPSL----VLVLTAC 120
C G G AC L ADD V PSL VL ++AC
Sbjct: 465 CC*GSSGKACHDLFADADADDVTAGEVFKPSLIGEGVLAVSAC 337
>TC203467 similar to UP|O82462 (O82462) Glutamyl-tRNA synthetase , partial
(88%)
Length = 2333
Score = 25.4 bits (54), Expect = 7.3
Identities = 9/22 (40%), Positives = 14/22 (62%)
Frame = +2
Query: 43 EFACPYADVINDLTNDCASTMF 64
+FACPY D I +T+ S+ +
Sbjct: 1211 DFACPYVDAIEGITHALRSSEY 1276
>BU544010 similar to GP|22795261|gb| nodulin-like protein {Oryza sativa
(japonica cultivar-group)}, partial (8%)
Length = 554
Score = 25.4 bits (54), Expect = 7.3
Identities = 12/34 (35%), Positives = 19/34 (55%)
Frame = -3
Query: 1 MTLFHIAPAGCSVNFEFLNYTIITSKCKGPKYPP 34
+T+ + + +F FL++ I K KGP YPP
Sbjct: 438 ITIVYSGALATAASFCFLSWAI---KIKGPSYPP 346
>TC214853 similar to UP|STO_ARATH (Q96288) Salt-tolerance protein, partial
(65%)
Length = 2471
Score = 25.4 bits (54), Expect = 7.3
Identities = 11/31 (35%), Positives = 17/31 (54%)
Frame = +1
Query: 4 FHIAPAGCSVNFEFLNYTIITSKCKGPKYPP 34
+ + +GC + LNY +S C+ PKY P
Sbjct: 652 YEASVSGCPRDLNPLNYPESSSACQFPKYWP 744
>TC203819
Length = 632
Score = 25.4 bits (54), Expect = 7.3
Identities = 9/24 (37%), Positives = 15/24 (62%)
Frame = +1
Query: 5 HIAPAGCSVNFEFLNYTIITSKCK 28
H+A G ++ ++LN + T KCK
Sbjct: 421 HLAAEGATILVKYLNPVLCTQKCK 492
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.141 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,495,190
Number of Sequences: 63676
Number of extensions: 114610
Number of successful extensions: 837
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 835
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 835
length of query: 127
length of database: 12,639,632
effective HSP length: 86
effective length of query: 41
effective length of database: 7,163,496
effective search space: 293703336
effective search space used: 293703336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144723.3


