

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144483.8 + phase: 0
(253 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
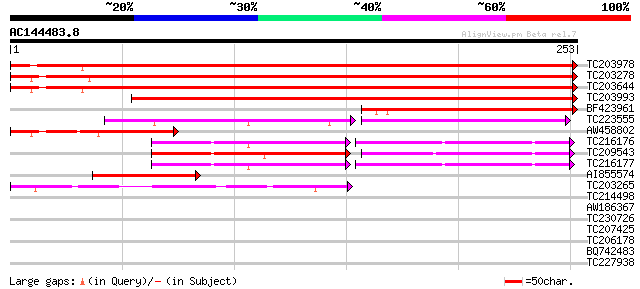
Score E
Sequences producing significant alignments: (bits) Value
TC203978 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, ... 411 e-115
TC203278 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, ... 389 e-109
TC203644 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, ... 385 e-107
TC203993 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, ... 360 e-100
BF423961 weakly similar to GP|7331143|gb|A chaperonin 21 precurs... 115 1e-26
TC223555 similar to UP|Q7PXP0 (Q7PXP0) AgCP11981 (Fragment), par... 72 3e-13
AW458802 GP|7264742|gb| alcohol dehydrogenase 7 {Vitis vinifera}... 64 9e-11
TC216176 similar to UP|Q9JI95 (Q9JI95) CPN10-like protein, parti... 61 4e-10
TC209543 similar to UP|CH10_ARATH (P34893) 10 kDa chaperonin (Pr... 61 4e-10
TC216177 similar to UP|Q9JI95 (Q9JI95) CPN10-like protein, parti... 61 4e-10
AI855574 similar to GP|21780187|gb| cp10-like protein {Gossypium... 55 3e-08
TC203265 similar to GB|AAO64777.1|29028796|BT005842 At3g60210 {A... 52 2e-07
TC214498 weakly similar to UP|PTH2_YEAST (P34222) Peptidyl-tRNA ... 29 2.3
AW186367 similar to GP|23534479|g cellulose synthase {Populus tr... 28 3.1
TC230726 28 3.1
TC207425 similar to UP|Q9FJ82 (Q9FJ82) Dihydrodipicolinate reduc... 28 3.1
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 28 4.0
BQ742483 similar to GP|21780187|gb| cp10-like protein {Gossypium... 28 5.2
TC227938 homologue to UP|Q9LK77 (Q9LK77) Similarity to acyl-CoA ... 27 8.9
>TC203978 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, partial
(83%)
Length = 1070
Score = 411 bits (1057), Expect = e-115
Identities = 211/254 (83%), Positives = 234/254 (92%), Gaps = 1/254 (0%)
Frame = +2
Query: 1 MATTQLTASSISTRNLSSFERLRTSSIQFPR-NNARIAILTQWSLPSMVVKAATVVAPKH 59
MAT+Q+T+S RNL +FER + S+++FP + RI LTQ S PS VVKAATVVAPKH
Sbjct: 146 MATSQITSS---LRNLPAFERFQPSTLRFPSAGHVRIGTLTQTSFPSFVVKAATVVAPKH 316
Query: 60 TTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKA 119
TTVKPLGDRVLVKIKDAEEKT GGILLP+TAQ KPQGGEVVA+GDGK+VGK+KVD+SVK
Sbjct: 317 TTVKPLGDRVLVKIKDAEEKTAGGILLPATAQGKPQGGEVVAVGDGKSVGKSKVDVSVKT 496
Query: 120 GAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTA 179
G QVVYSKYAGTEVEF+GSKHLILKD+DIVGILET+EVKDLKPLNDRVLIKVAEA+EKTA
Sbjct: 497 GLQVVYSKYAGTEVEFNGSKHLILKDDDIVGILETDEVKDLKPLNDRVLIKVAEADEKTA 676
Query: 180 GGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSD 239
GGLLLTEATK+KPSIGTVIAVGPGP+D+EGNRKPLS++PGNTVLYSKYAGNDFKGKDGSD
Sbjct: 677 GGLLLTEATKEKPSIGTVIAVGPGPLDEEGNRKPLSVVPGNTVLYSKYAGNDFKGKDGSD 856
Query: 240 YIALRASDVMAILS 253
YIALRASDVMAILS
Sbjct: 857 YIALRASDVMAILS 898
>TC203278 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, partial
(86%)
Length = 1099
Score = 389 bits (998), Expect = e-109
Identities = 202/256 (78%), Positives = 229/256 (88%), Gaps = 3/256 (1%)
Frame = +1
Query: 1 MATTQLTA--SSISTRNLSSFERLRTSSIQFPRNNA-RIAILTQWSLPSMVVKAATVVAP 57
MA+ QLTA SSIST +SFE LR S++QF RI L+Q S +VVKAATVVAP
Sbjct: 124 MASAQLTAIASSIST---ASFEGLRPSAVQFASTGRIRIGNLSQRSFRGLVVKAATVVAP 294
Query: 58 KHTTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISV 117
K+T +KPLGDRVL+KIK+AEEKT+GGILLPSTAQ+KPQGGEVVA+G+GKTVGKN V+ISV
Sbjct: 295 KYTAIKPLGDRVLIKIKEAEEKTEGGILLPSTAQTKPQGGEVVAVGEGKTVGKNNVEISV 474
Query: 118 KAGAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEK 177
K GAQVVYSKYAGTEV+FDG+KHLI+KD+DIVGILET+++KDLKPLNDRVLIKVAEAEEK
Sbjct: 475 KTGAQVVYSKYAGTEVDFDGTKHLIVKDDDIVGILETDDIKDLKPLNDRVLIKVAEAEEK 654
Query: 178 TAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDG 237
T+GGLLLTEATKDKPSIGTVIAVGPG +D+EGNRKPLS+ PGNTVLYSKYAGNDFKGKDG
Sbjct: 655 TSGGLLLTEATKDKPSIGTVIAVGPGHLDEEGNRKPLSVTPGNTVLYSKYAGNDFKGKDG 834
Query: 238 SDYIALRASDVMAILS 253
SDYI LR SDVMA+LS
Sbjct: 835 SDYITLRVSDVMAVLS 882
>TC203644 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, partial
(87%)
Length = 1177
Score = 385 bits (988), Expect = e-107
Identities = 199/256 (77%), Positives = 229/256 (88%), Gaps = 3/256 (1%)
Frame = +3
Query: 1 MATTQLTA--SSISTRNLSSFERLRTSSIQFPR-NNARIAILTQWSLPSMVVKAATVVAP 57
MA+ QL+A SSIST +SFE LR S++QF RI L+Q S +VVKAATVVAP
Sbjct: 153 MASAQLSAISSSIST---ASFEGLRPSAVQFASAGRIRIGNLSQRSFRGLVVKAATVVAP 323
Query: 58 KHTTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISV 117
K+T +KPLGDRVL+KIK+AEEKT+GGILLPSTAQ+KPQGGEVVA+G+GKT+GKN V+ISV
Sbjct: 324 KYTAIKPLGDRVLIKIKEAEEKTEGGILLPSTAQTKPQGGEVVAVGEGKTIGKNNVEISV 503
Query: 118 KAGAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEK 177
K GAQVVYSKYAGTEV+F+G+KHLI+KD+DIVGILET+++KDLKPLNDRVLIKVAEAEEK
Sbjct: 504 KTGAQVVYSKYAGTEVDFNGTKHLIVKDDDIVGILETDDIKDLKPLNDRVLIKVAEAEEK 683
Query: 178 TAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDG 237
T+GGLLLTEATKDKPSIGTVIAVGPG +D+EGNRKPLS+ PGNTVLYSKYAGNDFKGKDG
Sbjct: 684 TSGGLLLTEATKDKPSIGTVIAVGPGHLDEEGNRKPLSVTPGNTVLYSKYAGNDFKGKDG 863
Query: 238 SDYIALRASDVMAILS 253
SDYI LR SDVMA+LS
Sbjct: 864 SDYITLRVSDVMAVLS 911
>TC203993 similar to UP|Q9M5A8 (Q9M5A8) Chaperonin 21 precursor, partial
(79%)
Length = 783
Score = 360 bits (923), Expect = e-100
Identities = 179/199 (89%), Positives = 194/199 (96%)
Frame = +1
Query: 55 VAPKHTTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVD 114
VAPKHTTVKPLGDRVLVKIKDAEEKT GGILLP+ AQ KPQGGEVVA+G+GK+VGK KVD
Sbjct: 22 VAPKHTTVKPLGDRVLVKIKDAEEKTAGGILLPANAQGKPQGGEVVAVGEGKSVGKCKVD 201
Query: 115 ISVKAGAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEA 174
+SVK GAQVV+SKYAGTEVEF+GSKHLILKD+DIVGILET+EVKDLKPLNDRVLIKVAEA
Sbjct: 202 VSVKTGAQVVHSKYAGTEVEFNGSKHLILKDDDIVGILETDEVKDLKPLNDRVLIKVAEA 381
Query: 175 EEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKG 234
EEKTAGGLLLTEATK+KPSIGTVIAVGPGP+D+EGNRKPLS++PGNTVLYS+YAGNDFKG
Sbjct: 382 EEKTAGGLLLTEATKEKPSIGTVIAVGPGPLDEEGNRKPLSVMPGNTVLYSRYAGNDFKG 561
Query: 235 KDGSDYIALRASDVMAILS 253
KDGSDYIALRASDVMAILS
Sbjct: 562 KDGSDYIALRASDVMAILS 618
>BF423961 weakly similar to GP|7331143|gb|A chaperonin 21 precursor
{Lycopersicon esculentum}, partial (33%)
Length = 301
Score = 115 bits (289), Expect = 1e-26
Identities = 65/100 (65%), Positives = 75/100 (75%), Gaps = 4/100 (4%)
Frame = +2
Query: 158 KDLKP-LNDRV---LIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKP 213
KD+K LN R+ LI +E EE+T G LLLT AT D PSIGTVIA GPG +D+EG RKP
Sbjct: 2 KDIKDTLNTRLTKYLIVGSEPEEETCGRLLLTSATHDMPSIGTVIAGGPGHLDEEGARKP 181
Query: 214 LSILPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAILS 253
L+ PGNTVLYS AG DFKG DGSDYI+LR DV+A+LS
Sbjct: 182 LAWTPGNTVLYSACAGRDFKG*DGSDYISLRVLDVLAVLS 301
>TC223555 similar to UP|Q7PXP0 (Q7PXP0) AgCP11981 (Fragment), partial (74%)
Length = 723
Score = 71.6 bits (174), Expect = 3e-13
Identities = 37/93 (39%), Positives = 56/93 (59%)
Frame = +2
Query: 158 KDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSIL 217
K L PL DRVLI+ A+A KT GG+++ E + K GTV+AVGPG + +G P+S+
Sbjct: 152 KRLVPLFDRVLIQRADAVTKTKGGIVIPEKAQGKVLTGTVVAVGPGSRNSDGQHVPVSVK 331
Query: 218 PGNTVLYSKYAGNDFKGKDGSDYIALRASDVMA 250
G+ VL +Y G + + ++ R SD++A
Sbjct: 332 VGDQVLLPEYGGTKVELSESKEFHLFRESDMLA 430
Score = 69.7 bits (169), Expect = 1e-12
Identities = 44/117 (37%), Positives = 66/117 (55%), Gaps = 5/117 (4%)
Frame = +2
Query: 43 SLPSMVVKAATVVAPKHTTVK---PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEV 99
S P ++ A+ V + K PL DRVL++ DA KT+GGI++P AQ K G V
Sbjct: 92 SCPFILEMASAVAQSALASAKRLVPLFDRVLIQRADAVTKTKGGIVIPEKAQGKVLTGTV 271
Query: 100 VAIGDG-KTVGKNKVDISVKAGAQVVYSKYAGTEVEFDGSKHL-ILKDEDIVGILET 154
VA+G G + V +SVK G QV+ +Y GT+VE SK + ++ D++ +E+
Sbjct: 272 VAVGPGSRNSDGQHVPVSVKVGDQVLLPEYGGTKVELSESKEFHLFRESDMLAKIES 442
>AW458802 GP|7264742|gb| alcohol dehydrogenase 7 {Vitis vinifera}, partial
(5%)
Length = 313
Score = 63.5 bits (153), Expect = 9e-11
Identities = 42/79 (53%), Positives = 54/79 (68%), Gaps = 4/79 (5%)
Frame = +2
Query: 1 MATTQLTA--SSISTRNLSSFERLRTSSIQFPRNNARIAI--LTQWSLPSMVVKAATVVA 56
MA+ QLTA SSIST +SFE L+ S++ F + RI I L+Q S +VVK TVVA
Sbjct: 89 MASAQLTAIASSIST---ASFEGLQPSTVXFA-STGRIXIGNLSQRSFQGLVVKXTTVVA 256
Query: 57 PKHTTVKPLGDRVLVKIKD 75
PK+T + PLGD VL+KIK+
Sbjct: 257 PKYTAIXPLGDXVLIKIKE 313
>TC216176 similar to UP|Q9JI95 (Q9JI95) CPN10-like protein, partial (36%)
Length = 592
Score = 61.2 bits (147), Expect = 4e-10
Identities = 39/98 (39%), Positives = 52/98 (52%)
Frame = +1
Query: 155 EEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPL 214
E K L P +R+LI+ KT+ G+LL E T S G VIAVGPG D GN P+
Sbjct: 109 EMAKRLIPCFNRILIEKIVPPSKTSAGILLPEKTSQLNS-GKVIAVGPGSRDKAGNLIPV 285
Query: 215 SILPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
S+ G+ VL +Y G K D ++ R D++ IL
Sbjct: 286 SVKEGDHVLLPEYGGTQIK-LDDKEFHLFRDEDILGIL 396
Score = 60.5 bits (145), Expect = 7e-10
Identities = 33/90 (36%), Positives = 53/90 (58%), Gaps = 1/90 (1%)
Frame = +1
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDG-KTVGKNKVDISVKAGAQ 122
P +R+L++ KT GILLP S+ G+V+A+G G + N + +SVK G
Sbjct: 130 PCFNRILIEKIVPPSKTSAGILLPEKT-SQLNSGKVIAVGPGSRDKAGNLIPVSVKEGDH 306
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V+ +Y GT+++ D + + +DEDI+GIL
Sbjct: 307 VLLPEYGGTQIKLDDKEFHLFRDEDILGIL 396
>TC209543 similar to UP|CH10_ARATH (P34893) 10 kDa chaperonin (Protein CPN10)
(Protein groES), partial (98%)
Length = 487
Score = 61.2 bits (147), Expect = 4e-10
Identities = 36/90 (40%), Positives = 54/90 (60%), Gaps = 1/90 (1%)
Frame = +3
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNK-VDISVKAGAQ 122
PL +RVLV+ KT GILLP + SK G+V+A+G G K + ++VK G
Sbjct: 90 PLFNRVLVEKIVPPSKTNAGILLPEKS-SKLNSGKVIAVGPGFHSKDGKLIPVAVKEGDT 266
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V+ +Y GTEV+ D ++ + +D+DI+G L
Sbjct: 267 VLLPEYGGTEVKLDNKEYHLFRDDDILGTL 356
Score = 57.0 bits (136), Expect = 8e-09
Identities = 36/95 (37%), Positives = 52/95 (53%)
Frame = +3
Query: 158 KDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSIL 217
K L PL +RVL++ KT G+LL E + K + G VIAVGPG +G P+++
Sbjct: 78 KRLIPLFNRVLVEKIVPPSKTNAGILLPEKSS-KLNSGKVIAVGPGFHSKDGKLIPVAVK 254
Query: 218 PGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
G+TVL +Y G + K D +Y R D++ L
Sbjct: 255 EGDTVLLPEYGGTEVK-LDNKEYHLFRDDDILGTL 356
>TC216177 similar to UP|Q9JI95 (Q9JI95) CPN10-like protein, partial (36%)
Length = 618
Score = 61.2 bits (147), Expect = 4e-10
Identities = 34/90 (37%), Positives = 54/90 (59%), Gaps = 1/90 (1%)
Frame = +1
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDG-KTVGKNKVDISVKAGAQ 122
P +R+LV+ KT GILLP + S+ G+V+A+G G + N + +SVK G
Sbjct: 169 PCFNRILVEKIVPPSKTSAGILLPEKS-SQLNSGKVIAVGPGSRDQAGNLIPVSVKEGDH 345
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V+ +Y GT+++ D + + +DEDI+GIL
Sbjct: 346 VLLPEYGGTQIKLDDKEFHLFRDEDILGIL 435
Score = 59.7 bits (143), Expect = 1e-09
Identities = 37/98 (37%), Positives = 52/98 (52%)
Frame = +1
Query: 155 EEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPL 214
E K L P +R+L++ KT+ G+LL E + S G VIAVGPG D GN P+
Sbjct: 148 EMAKRLIPCFNRILVEKIVPPSKTSAGILLPEKSSQLNS-GKVIAVGPGSRDQAGNLIPV 324
Query: 215 SILPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
S+ G+ VL +Y G K D ++ R D++ IL
Sbjct: 325 SVKEGDHVLLPEYGGTQIK-LDDKEFHLFRDEDILGIL 435
>AI855574 similar to GP|21780187|gb| cp10-like protein {Gossypium
hirsutum}, partial (17%)
Length = 145
Score = 55.1 bits (131), Expect = 3e-08
Identities = 28/48 (58%), Positives = 35/48 (72%)
Frame = +2
Query: 38 ILTQWSLPSMVVKAATVVAPKHTTVKPLGDRVLVKIKDAEEKTQGGIL 85
IL+Q S +VVKA TVVAPK+T +KPL DR L+KI + +E T GG L
Sbjct: 2 ILSQRSFRGLVVKADTVVAPKYTAIKPL*DRALIKIIERQEITDGGFL 145
Score = 30.8 bits (68), Expect = 0.62
Identities = 18/43 (41%), Positives = 25/43 (57%)
Frame = +2
Query: 141 LILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAGGLL 183
L++K + +V T +KPL DR LIK+ E +E T GG L
Sbjct: 29 LVVKADTVVAPKYTA----IKPL*DRALIKIIERQEITDGGFL 145
>TC203265 similar to GB|AAO64777.1|29028796|BT005842 At3g60210 {Arabidopsis
thaliana;} , partial (75%)
Length = 720
Score = 52.4 bits (124), Expect = 2e-07
Identities = 46/156 (29%), Positives = 78/156 (49%), Gaps = 3/156 (1%)
Frame = +3
Query: 1 MATTQLTASS--ISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPK 58
MA+T LT + + N +F R S +Q R++ +I +T+ P+ VV
Sbjct: 126 MASTFLTLPTPFLHKTNAITFSDKRPSFLQ--RSSLKINAITKKWEPTKVV--------- 272
Query: 59 HTTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVK 118
P DRVLV++++ +KT GG+LLP +A E +G+ TVG ++ K
Sbjct: 273 -----PQADRVLVRLEELSDKTVGGVLLPKSAVK----FERYLVGEILTVGAEAGEL--K 419
Query: 119 AGAQVVYSKYAGTEVEF-DGSKHLILKDEDIVGILE 153
AG +V+++ EV+ +KH K D++ ++E
Sbjct: 420 AGTKVLFTDMNAYEVDLGTDAKHCFCKASDLLAVVE 527
Score = 34.7 bits (78), Expect = 0.043
Identities = 27/107 (25%), Positives = 52/107 (48%), Gaps = 2/107 (1%)
Frame = +3
Query: 148 IVGILETEEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATK--DKPSIGTVIAVGPGPV 205
I I + E + P DRVL+++ E +KT GG+LL ++ ++ +G ++ VG
Sbjct: 231 INAITKKWEPTKVVPQADRVLVRLEELSDKTVGGVLLPKSAVKFERYLVGEILTVGA--- 401
Query: 206 DDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
+ G K G VL++ + + + +ASD++A++
Sbjct: 402 -EAGELK-----AGTKVLFTDMNAYEVDLGTDAKHCFCKASDLLAVV 524
>TC214498 weakly similar to UP|PTH2_YEAST (P34222) Peptidyl-tRNA hydrolase 2
(PTH 2) , partial (16%)
Length = 976
Score = 28.9 bits (63), Expect = 2.3
Identities = 22/57 (38%), Positives = 31/57 (53%), Gaps = 6/57 (10%)
Frame = +2
Query: 164 NDRVLIKVAEAEEKTA-GGLLLTEATKDKPSIG--TVIAVGPGP---VDDEGNRKPL 214
N + + K+AEA E ++ +A + + S G TV+AVGPGP VD R PL
Sbjct: 530 NQQEMNKLAEAAESIGLPTFVVADAGRTQVSAGSKTVLAVGPGPKSSVDSVTGRLPL 700
>AW186367 similar to GP|23534479|g cellulose synthase {Populus tremuloides},
partial (13%)
Length = 431
Score = 28.5 bits (62), Expect = 3.1
Identities = 25/91 (27%), Positives = 39/91 (42%), Gaps = 7/91 (7%)
Frame = +1
Query: 166 RVLIKVAEAEEKTAGGLLLTEAT-------KDKPSIGTVIAVGPGPVDDEGNRKPLSILP 218
R+ VA+A++ GG ++ + T KD P + V G +D EGN+ P +
Sbjct: 37 RINALVAKAQKVPQGGWIMQDGTPWPGNNTKDHPGMIQVFLGSSGGLDTEGNQLPRLVY- 213
Query: 219 GNTVLYSKYAGNDFKGKDGSDYIALRASDVM 249
V K G K G+ +R S V+
Sbjct: 214 ---VSREKRPGFQHHKKAGAMNALVRVSAVL 297
>TC230726
Length = 554
Score = 28.5 bits (62), Expect = 3.1
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = -2
Query: 197 VIAVGPGPVDDEGNRKPLSILPG 219
V+ VGP P+++E P ++LPG
Sbjct: 394 VVVVGPAPLEEEEGASPPALLPG 326
>TC207425 similar to UP|Q9FJ82 (Q9FJ82) Dihydrodipicolinate reductase-like
protein, partial (42%)
Length = 555
Score = 28.5 bits (62), Expect = 3.1
Identities = 28/96 (29%), Positives = 42/96 (43%)
Frame = +1
Query: 85 LLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAGAQVVYSKYAGTEVEFDGSKHLILK 144
L+ +AQ +VV G K +GK A V +K G EV G+
Sbjct: 157 LISCSAQPTQNNIKVVINGATKEIGK---------AAVVAVTKARGMEVA--GAVDTCHI 303
Query: 145 DEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAG 180
EDI I EE+ ++ +ND +I + ++ K AG
Sbjct: 304 GEDIGKICGMEELLEIPIINDLTMILGSISQTKAAG 411
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 28.1 bits (61), Expect = 4.0
Identities = 21/58 (36%), Positives = 30/58 (51%), Gaps = 3/58 (5%)
Frame = -1
Query: 2 ATTQLTASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAA---TVVA 56
ATT A++ +T +S RT S PRN IA +T W + ++ + A TVVA
Sbjct: 814 ATTSTAAATTATTAVSKNIGFRTLSSHMPRN---IAQITNWIVRTITCQVACLPTVVA 650
>BQ742483 similar to GP|21780187|gb| cp10-like protein {Gossypium hirsutum},
partial (5%)
Length = 406
Score = 27.7 bits (60), Expect = 5.2
Identities = 12/14 (85%), Positives = 12/14 (85%)
Frame = +2
Query: 117 VKAGAQVVYSKYAG 130
VK G QVVYSKYAG
Sbjct: 365 VKTGLQVVYSKYAG 406
>TC227938 homologue to UP|Q9LK77 (Q9LK77) Similarity to acyl-CoA
thioesterase, partial (16%)
Length = 542
Score = 26.9 bits (58), Expect = 8.9
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = +1
Query: 202 PGPVDDEGNRKPLSILPG 219
P P D G+RKP+++ PG
Sbjct: 121 PPPADSSGDRKPIALWPG 174
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.131 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,761,459
Number of Sequences: 63676
Number of extensions: 84860
Number of successful extensions: 361
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 332
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 341
length of query: 253
length of database: 12,639,632
effective HSP length: 95
effective length of query: 158
effective length of database: 6,590,412
effective search space: 1041285096
effective search space used: 1041285096
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144483.8


