

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
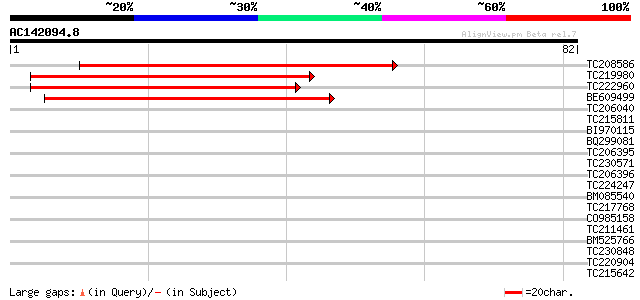
Query= AC142094.8 - phase: 0 /pseudo
(82 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC208586 similar to UP|Q43323 (Q43323) Membrane protein, partial... 49 3e-07
TC219980 47 2e-06
TC222960 similar to UP|Q43323 (Q43323) Membrane protein, partial... 45 5e-06
BE609499 39 4e-04
TC206040 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (... 30 0.28
TC215811 similar to UP|O49529 (O49529) Predicted protein, partia... 28 1.1
BI970115 27 2.4
BQ299081 similar to GP|2827699|emb predicted protein {Arabidopsi... 26 4.1
TC206395 26 4.1
TC230571 similar to UP|Q41645 (Q41645) Extensin (Fragment), part... 25 5.4
TC206396 25 5.4
TC224247 similar to UP|Q9FF84 (Q9FF84) ATP-dependent RNA helicas... 25 7.0
BM085540 25 7.0
TC217768 25 7.0
CO985158 25 7.0
TC211461 weakly similar to UP|DPD4_SCHPO (O59835) DNA polymerase... 25 9.1
BM525766 SP|P17817|PROC Pyrroline-5-carboxylate reductase (EC 1.... 25 9.1
TC230848 similar to UP|Q8LPT2 (Q8LPT2) At1g10410/F14N23_31, part... 25 9.1
TC220904 homologue to GB|AAS47665.1|44681450|BT011659 At5g03110 ... 25 9.1
TC215642 similar to UP|Q84MC1 (Q84MC1) At4g31290, partial (80%) 25 9.1
>TC208586 similar to UP|Q43323 (Q43323) Membrane protein, partial (66%)
Length = 893
Score = 49.3 bits (116), Expect = 3e-07
Identities = 24/46 (52%), Positives = 31/46 (67%)
Frame = +2
Query: 11 VFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLH 56
V NL IG++P V+NF IGG + GFLLGF+LL + L Q+LH
Sbjct: 443 VINLAIGMLPHVDNFAHIGGFLSGFLLGFILLLRPQFGWLEQQRLH 580
>TC219980
Length = 1021
Score = 46.6 bits (109), Expect = 2e-06
Identities = 21/41 (51%), Positives = 30/41 (72%)
Frame = +3
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ ++ V NL +GI+P V+NF IGG + GFLLGFV L +
Sbjct: 273 LTLVIIIVINLAVGILPHVDNFAHIGGFLTGFLLGFVFLIR 395
>TC222960 similar to UP|Q43323 (Q43323) Membrane protein, partial (52%)
Length = 731
Score = 45.4 bits (106), Expect = 5e-06
Identities = 21/39 (53%), Positives = 30/39 (76%)
Frame = +1
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
+++ ++ V NL IGI+P V+NF IGG + GFLLGF+LL
Sbjct: 190 ITLLVIIVINLGIGILPHVDNFAHIGGFLVGFLLGFILL 306
>BE609499
Length = 442
Score = 39.3 bits (90), Expect = 4e-04
Identities = 19/42 (45%), Positives = 27/42 (64%)
Frame = +2
Query: 6 IGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDP 47
+G + VF LT GI+P V +GGL+ GFL+GFV + + P
Sbjct: 167 LGFLLVFILTGGILPPVGGSAHMGGLLAGFLVGFVFVIRAPP 292
>TC206040 similar to UP|Q9XFB4 (Q9XFB4) PR1a precursor, partial (90%)
Length = 772
Score = 29.6 bits (65), Expect = 0.28
Identities = 18/61 (29%), Positives = 30/61 (48%), Gaps = 7/61 (11%)
Frame = -3
Query: 18 IVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKR-------CLPIICFILLST 70
I+P++ + L F L ++LCK V+P Q L+ C+ +IC+ +LS
Sbjct: 338 ILPVLATTTRVDQLTITFTLISIVLCKSSNRVIPSQVLNTHL*SCSVVCIDVICW*VLSI 159
Query: 71 G 71
G
Sbjct: 158 G 156
>TC215811 similar to UP|O49529 (O49529) Predicted protein, partial (5%)
Length = 629
Score = 27.7 bits (60), Expect = 1.1
Identities = 11/31 (35%), Positives = 17/31 (54%)
Frame = +1
Query: 35 FLLGFVLLCKKDPFVLPDQKLHKRCLPIICF 65
FL F +C + + + LH+RC+P I F
Sbjct: 55 FLSYFFWVCMMNMYAYSNTVLHRRCIPFISF 147
>BI970115
Length = 449
Score = 26.6 bits (57), Expect = 2.4
Identities = 13/43 (30%), Positives = 23/43 (53%)
Frame = -1
Query: 12 FNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQK 54
++L G VPI +GG++ V+ K DPF+ P+++
Sbjct: 377 WSLLTGPVPI------LGGVVVAIAAANVIFVKNDPFLQPEER 267
>BQ299081 similar to GP|2827699|emb predicted protein {Arabidopsis
thaliana}, partial (5%)
Length = 422
Score = 25.8 bits (55), Expect = 4.1
Identities = 9/27 (33%), Positives = 15/27 (55%)
Frame = +2
Query: 39 FVLLCKKDPFVLPDQKLHKRCLPIICF 65
F +C + + + LH+RC+P I F
Sbjct: 2 FFWVCMMNMYAYSNTVLHRRCIPFISF 82
>TC206395
Length = 522
Score = 25.8 bits (55), Expect = 4.1
Identities = 8/31 (25%), Positives = 19/31 (60%)
Frame = +2
Query: 27 LIGGLIPGFLLGFVLLCKKDPFVLPDQKLHK 57
++GG++ V+ K DPF+ P+++ ++
Sbjct: 128 ILGGVVVAIAAANVIFVKNDPFLKPEERKYE 220
>TC230571 similar to UP|Q41645 (Q41645) Extensin (Fragment), partial (7%)
Length = 667
Score = 25.4 bits (54), Expect = 5.4
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = -2
Query: 30 GLIPGFLLGFVLLCKKDPFV 49
GL+PG L+G +LL + PF+
Sbjct: 60 GLLPGTLMGVLLLLSRLPFL 1
>TC206396
Length = 795
Score = 25.4 bits (54), Expect = 5.4
Identities = 8/28 (28%), Positives = 17/28 (60%)
Frame = +2
Query: 27 LIGGLIPGFLLGFVLLCKKDPFVLPDQK 54
++GG++ V+ K DPF+ P+++
Sbjct: 398 ILGGVVVAIAAANVIFVKNDPFLQPEER 481
>TC224247 similar to UP|Q9FF84 (Q9FF84) ATP-dependent RNA helicase A-like
protein, partial (7%)
Length = 706
Score = 25.0 bits (53), Expect = 7.0
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = +2
Query: 16 IGIVPIVNNFGLIGGLIPGFLLGFV 40
IGI + + +IG LIP F LGF+
Sbjct: 392 IGI*VTFHQYMVIGELIPSFHLGFI 466
>BM085540
Length = 391
Score = 25.0 bits (53), Expect = 7.0
Identities = 12/26 (46%), Positives = 15/26 (57%)
Frame = -3
Query: 34 GFLLGFVLLCKKDPFVLPDQKLHKRC 59
G+ L VLLC+K P LP + L C
Sbjct: 341 GYELELVLLCRKGPLELP*RGLLGTC 264
>TC217768
Length = 1516
Score = 25.0 bits (53), Expect = 7.0
Identities = 17/55 (30%), Positives = 23/55 (40%), Gaps = 6/55 (10%)
Frame = +2
Query: 33 PGFLLGFVLLCKKDPFVLPDQKLHKRCLPII------CFILLSTGWTGVLTKGSE 81
P F+ GF C K P K ++ P + C STGW G +KG +
Sbjct: 1037 PEFISGFPARCPKWP------KFSRKLSPAMEFRNGWCAF*TSTGWYGCGSKGGQ 1183
>CO985158
Length = 780
Score = 25.0 bits (53), Expect = 7.0
Identities = 7/25 (28%), Positives = 17/25 (68%)
Frame = +3
Query: 40 VLLCKKDPFVLPDQKLHKRCLPIIC 64
++ C++D F+ Q+++ LP++C
Sbjct: 135 LICCRRDSFLFSKQRIYIFILPLLC 209
>TC211461 weakly similar to UP|DPD4_SCHPO (O59835) DNA polymerase delta
subunit 4 , partial (15%)
Length = 459
Score = 24.6 bits (52), Expect = 9.1
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = -1
Query: 28 IGGLIPGFLLGFVLLCKKDPFVLPD 52
IG + FL+ V C PF+ PD
Sbjct: 117 IGDFLEDFLVAVVFFCL*KPFMFPD 43
>BM525766 SP|P17817|PROC Pyrroline-5-carboxylate reductase (EC 1.5.1.2)
(P5CR) (P5C reductase). [Soybean] {Glycine max}, partial
(4%)
Length = 428
Score = 24.6 bits (52), Expect = 9.1
Identities = 9/19 (47%), Positives = 11/19 (57%)
Frame = +2
Query: 54 KLHKRCLPIICFILLSTGW 72
KLH C +CF+L GW
Sbjct: 239 KLHMICYNSLCFVLHYEGW 295
>TC230848 similar to UP|Q8LPT2 (Q8LPT2) At1g10410/F14N23_31, partial (24%)
Length = 1400
Score = 24.6 bits (52), Expect = 9.1
Identities = 10/26 (38%), Positives = 14/26 (53%)
Frame = -2
Query: 27 LIGGLIPGFLLGFVLLCKKDPFVLPD 52
+ G I FL+ V C+ PF+ PD
Sbjct: 142 MAGERIGDFLVAMVFFCR*KPFMFPD 65
>TC220904 homologue to GB|AAS47665.1|44681450|BT011659 At5g03110 {Arabidopsis
thaliana;} , partial (7%)
Length = 708
Score = 24.6 bits (52), Expect = 9.1
Identities = 11/20 (55%), Positives = 14/20 (70%)
Frame = -3
Query: 24 NFGLIGGLIPGFLLGFVLLC 43
NF + GL GF+ GF+LLC
Sbjct: 220 NFSVRPGLEMGFVGGFMLLC 161
>TC215642 similar to UP|Q84MC1 (Q84MC1) At4g31290, partial (80%)
Length = 1146
Score = 24.6 bits (52), Expect = 9.1
Identities = 11/20 (55%), Positives = 13/20 (65%)
Frame = +3
Query: 28 IGGLIPGFLLGFVLLCKKDP 47
IGG G+LLG LLC + P
Sbjct: 303 IGGERRGYLLGSCLLCSRRP 362
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.154 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,700,352
Number of Sequences: 63676
Number of extensions: 75599
Number of successful extensions: 472
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 472
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 472
length of query: 82
length of database: 12,639,632
effective HSP length: 58
effective length of query: 24
effective length of database: 8,946,424
effective search space: 214714176
effective search space used: 214714176
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC142094.8


