

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.3 + phase: 0
(85 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
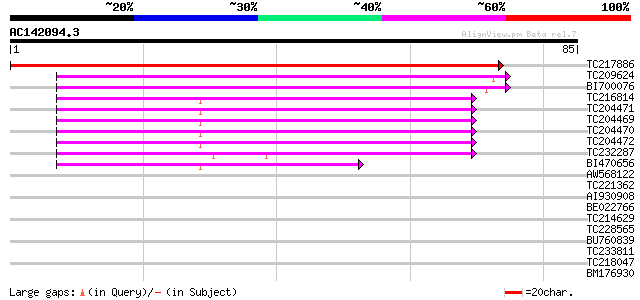
Sequences producing significant alignments: (bits) Value
TC217886 132 3e-32
TC209624 75 8e-15
BI700076 similar to GP|9755373|gb|A F17F8.4 {Arabidopsis thalian... 74 1e-14
TC216814 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (53%) 55 5e-09
TC204471 52 4e-08
TC204469 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (62%) 52 4e-08
TC204470 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (62%) 52 7e-08
TC204472 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (62%) 52 7e-08
TC232287 weakly similar to UP|Q93V70 (Q93V70) At1g21000/F9H16_1,... 47 1e-06
BI470656 38 8e-04
AW568122 32 0.043
TC221362 29 0.36
AI930908 similar to GP|18252504|gb| adenosine 5'-phosphosulfate ... 27 1.4
BE022766 27 1.4
TC214629 UP|Q9XQB2 (Q9XQB2) Chlorophyll a/b binding protein CP29... 27 2.4
TC228565 similar to UP|Q9XF14 (Q9XF14) Bundle sheath defective p... 27 2.4
BU760839 homologue to GP|19911573|dbj syringolide-induced protei... 26 3.1
TC233811 similar to GB|AAB61094.1|2194119|F20P5 F20P5.5 {Arabido... 26 3.1
TC218047 similar to UP|Q8GT81 (Q8GT81) Fiber protein Fb12 (Fragm... 26 3.1
BM176930 26 4.0
>TC217886
Length = 1093
Score = 132 bits (332), Expect = 3e-32
Identities = 57/74 (77%), Positives = 64/74 (86%)
Frame = +3
Query: 1 MNSSSGYCDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSEL 60
MNSS GYC H DLRSNE N FCVDCA+R CRHCKEAHS+HRRFQIY+YSYQDVFRH+EL
Sbjct: 174 MNSSFGYCTYHHDLRSNEMNVFCVDCALRMCRHCKEAHSLHRRFQIYKYSYQDVFRHAEL 353
Query: 61 QKHFDCSNIQVTLN 74
QK+FDCS IQ ++
Sbjct: 354 QKYFDCSKIQTYIS 395
>TC209624
Length = 860
Score = 74.7 bits (182), Expect = 8e-15
Identities = 32/69 (46%), Positives = 41/69 (59%), Gaps = 1/69 (1%)
Frame = +1
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H + + NEKN C+DC C HC +H HR Q+ RY Y DV R +LQK DCS
Sbjct: 139 CSYHENAKKNEKNVCCLDCCTSICPHCLPSHRFHRLLQVRRYVYHDVVRLEDLQKLIDCS 318
Query: 68 NIQV-TLNA 75
N+Q T+N+
Sbjct: 319 NVQAYTINS 345
>BI700076 similar to GP|9755373|gb|A F17F8.4 {Arabidopsis thaliana}, partial
(36%)
Length = 421
Score = 74.3 bits (181), Expect = 1e-14
Identities = 32/69 (46%), Positives = 41/69 (59%), Gaps = 1/69 (1%)
Frame = +1
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H + + NEKN C+DC C HC +H HR Q+ RY Y DV R +LQK DCS
Sbjct: 208 CSHHENAKKNEKNICCLDCCTSICPHCLPSHRCHRLLQVRRYVYHDVVRLEDLQKLIDCS 387
Query: 68 NIQ-VTLNA 75
N+Q T+N+
Sbjct: 388 NVQPYTINS 414
>TC216814 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (53%)
Length = 797
Score = 55.5 bits (132), Expect = 5e-09
Identities = 27/64 (42%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Frame = +3
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C +H HR QI R SY DV R SE+QK D
Sbjct: 303 CKVHADSHKSECNMYCLDCVNGALCSACLSSHKEHRIIQIRRSSYHDVIRVSEIQKFLDI 482
Query: 67 SNIQ 70
+ +Q
Sbjct: 483 TGVQ 494
>TC204471
Length = 509
Score = 52.4 bits (124), Expect = 4e-08
Identities = 27/64 (42%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Frame = +3
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H HR QI R SY DV R SE+QK D
Sbjct: 303 CKLHADSHKSECNMYCLDCMNGPLCSLCLAHHKDHRAIQIRRSSYHDVIRVSEIQKVLDI 482
Query: 67 SNIQ 70
+ +Q
Sbjct: 483 TGVQ 494
>TC204469 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (62%)
Length = 1382
Score = 52.4 bits (124), Expect = 4e-08
Identities = 27/64 (42%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Frame = +1
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H HR QI R SY DV R SE+QK D
Sbjct: 292 CKLHADSHKSECNMYCLDCMNGPLCSLCLAHHKDHRAIQIRRSSYHDVIRVSEIQKVLDI 471
Query: 67 SNIQ 70
+ +Q
Sbjct: 472 TGVQ 483
>TC204470 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (62%)
Length = 1349
Score = 51.6 bits (122), Expect = 7e-08
Identities = 26/64 (40%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = +1
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H HR QI R SY DV R +E+QK D
Sbjct: 331 CKLHADSHKSECNMYCLDCMNGALCSLCLAYHKDHRAIQIRRSSYHDVIRVNEIQKVLDI 510
Query: 67 SNIQ 70
+ +Q
Sbjct: 511 TGVQ 522
>TC204472 similar to UP|Q9LPJ4 (Q9LPJ4) F6N18.8, partial (62%)
Length = 1513
Score = 51.6 bits (122), Expect = 7e-08
Identities = 26/64 (40%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = +1
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H HR QI R SY DV R +E+QK D
Sbjct: 469 CKLHADSHKSECNMYCLDCMNGALCSLCLAYHKDHRAIQIRRSSYHDVIRVNEIQKVLDI 648
Query: 67 SNIQ 70
+ +Q
Sbjct: 649 TGVQ 660
>TC232287 weakly similar to UP|Q93V70 (Q93V70) At1g21000/F9H16_1, partial
(52%)
Length = 793
Score = 47.4 bits (111), Expect = 1e-06
Identities = 26/65 (40%), Positives = 35/65 (53%), Gaps = 2/65 (3%)
Frame = +2
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEA-HSIHRRFQIYRYSYQDVFRHSELQKHFD 65
C H D +E N FC+DC C +C+ + H H+ QI R SY DV R +E+Q D
Sbjct: 185 CRIHGDAARSECNMFCLDCNENACCFYCRSSKHKDHQVIQIRRSSYHDVVRVAEIQTVLD 364
Query: 66 CSNIQ 70
S +Q
Sbjct: 365 ISGVQ 379
>BI470656
Length = 423
Score = 38.1 bits (87), Expect = 8e-04
Identities = 19/47 (40%), Positives = 22/47 (46%), Gaps = 1/47 (2%)
Frame = +2
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQD 53
C H D +E N +C+DC C C H HR QI R SY D
Sbjct: 281 CKLHADSHKSECNMYCLDCMNGALCSLCLAYHKDHRAIQIRRSSYHD 421
>AW568122
Length = 430
Score = 32.3 bits (72), Expect = 0.043
Identities = 14/40 (35%), Positives = 19/40 (47%), Gaps = 1/40 (2%)
Frame = +1
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQI 46
C H D +E N +C+DC C C +H HR Q+
Sbjct: 307 CKVHADSHKSECNMYCLDCMNGALCSTCLASHREHRAIQV 426
>TC221362
Length = 694
Score = 29.3 bits (64), Expect = 0.36
Identities = 19/57 (33%), Positives = 30/57 (52%), Gaps = 6/57 (10%)
Frame = -2
Query: 34 CKEAHSIHRRFQIYRYSYQDVFRHSELQKHFD------CSNIQVTLNALLQVIDSFS 84
CK++H HR+ R S Q + S LQ F+ CS+ Q TL+ ++ + +FS
Sbjct: 639 CKQSHDCHRKSHKKRLSKQHRYLRSSLQYRFNHLS**ACSS*Q-TLHKIMFITITFS 472
>AI930908 similar to GP|18252504|gb| adenosine 5'-phosphosulfate reductase
{Glycine max}, partial (16%)
Length = 404
Score = 27.3 bits (59), Expect = 1.4
Identities = 11/21 (52%), Positives = 13/21 (61%)
Frame = -1
Query: 33 HCKEAHSIHRRFQIYRYSYQD 53
HC EAHSI RF + Y +D
Sbjct: 329 HCLEAHSISGRFHVLYYLCED 267
>BE022766
Length = 181
Score = 27.3 bits (59), Expect = 1.4
Identities = 14/33 (42%), Positives = 15/33 (45%), Gaps = 3/33 (9%)
Frame = -3
Query: 23 CVDCAVRFCRHCKEAHS---IHRRFQIYRYSYQ 52
C C C C H+ IHRR Q R SYQ
Sbjct: 173 CSPCKPS*CHTCTSQHAQLMIHRRHQYLRNSYQ 75
>TC214629 UP|Q9XQB2 (Q9XQB2) Chlorophyll a/b binding protein CP29, partial
(42%)
Length = 779
Score = 26.6 bits (57), Expect = 2.4
Identities = 11/41 (26%), Positives = 20/41 (47%)
Frame = +2
Query: 11 HRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSY 51
H L+ +K C+ CA F +CK + R + + ++ Y
Sbjct: 632 HPPLKKKQKKKNCLSCAWSF*LYCKSLFLLLRLWVVIKHFY 754
>TC228565 similar to UP|Q9XF14 (Q9XF14) Bundle sheath defective protein 2,
partial (54%)
Length = 632
Score = 26.6 bits (57), Expect = 2.4
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = -1
Query: 4 SSGYCDDHRDLRSNEKNTFCVDCAVRFC 31
S+ YC +R+N FC C+V FC
Sbjct: 335 ST*YCTISITIRTNNAPNFCAFCSVIFC 252
>BU760839 homologue to GP|19911573|dbj syringolide-induced protein 19-1-5
{Glycine max}, partial (25%)
Length = 447
Score = 26.2 bits (56), Expect = 3.1
Identities = 10/32 (31%), Positives = 14/32 (43%)
Frame = +1
Query: 26 CAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRH 57
C +FCRHC + R Y Y+ +H
Sbjct: 322 CTRKFCRHCHRLLCKY*RLMYYVYACLSAIKH 417
>TC233811 similar to GB|AAB61094.1|2194119|F20P5 F20P5.5 {Arabidopsis
thaliana;} , partial (18%)
Length = 411
Score = 26.2 bits (56), Expect = 3.1
Identities = 12/40 (30%), Positives = 19/40 (47%)
Frame = -2
Query: 18 EKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRH 57
E ++ C+ H AHS+H Q+ + SY+D H
Sbjct: 251 ESHSMCL*VQASTNEHNHSAHSLHSHLQMEKDSYKDQGGH 132
>TC218047 similar to UP|Q8GT81 (Q8GT81) Fiber protein Fb12 (Fragment),
partial (91%)
Length = 833
Score = 26.2 bits (56), Expect = 3.1
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = -3
Query: 45 QIYRYSYQDVFRHSELQKHFDCSNIQV 71
Q+Y+YSYQ +R+S+ CSN ++
Sbjct: 465 QVYKYSYQHPYRNSQ-----PCSNSKI 400
>BM176930
Length = 315
Score = 25.8 bits (55), Expect = 4.0
Identities = 8/20 (40%), Positives = 13/20 (65%)
Frame = +2
Query: 31 CRHCKEAHSIHRRFQIYRYS 50
C+ CKE H H + +Y+Y+
Sbjct: 176 CKVCKETHESHSQ*MMYKYN 235
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.329 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,891,563
Number of Sequences: 63676
Number of extensions: 71258
Number of successful extensions: 514
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 514
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 514
length of query: 85
length of database: 12,639,632
effective HSP length: 61
effective length of query: 24
effective length of database: 8,755,396
effective search space: 210129504
effective search space used: 210129504
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC142094.3


