

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141109.6 - phase: 0
(348 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
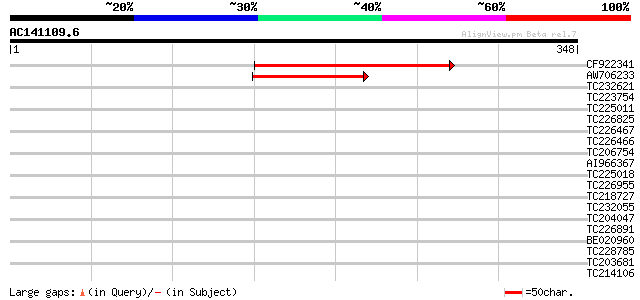
Score E
Sequences producing significant alignments: (bits) Value
CF922341 114 5e-26
AW706233 64 8e-11
TC232621 similar to UP|Q8LPN7 (Q8LPN7) AT3g19950/MPN9_19, partia... 33 0.15
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 33 0.25
TC225011 similar to UP|Q09083 (Q09083) Hydroxyproline-rich glyco... 33 0.25
TC226825 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydra... 32 0.33
TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 32 0.43
TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {... 32 0.43
TC206754 weakly similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-b... 32 0.43
AI966367 similar to GP|14335028|gb| AT4g37280/C7A10_80 {Arabidop... 32 0.43
TC225018 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich gly... 32 0.56
TC226955 similar to UP|Q9FQZ6 (Q9FQZ6) Avr9/Cf-9 rapidly elicite... 31 0.73
TC218727 similar to UP|Q8LSC5 (Q8LSC5) Geranylgeranyl pyrophosph... 31 0.73
TC232055 similar to UP|O49524 (O49524) Pherophorin - like protei... 31 0.73
TC204047 hydroxyproline-rich glycoprotein 31 0.73
TC226891 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-... 25 1.2
BE020960 30 1.2
TC228785 weakly similar to GB|AAP42749.1|30984572|BT008736 At3g5... 30 1.2
TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 30 1.2
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 30 1.2
>CF922341
Length = 675
Score = 114 bits (286), Expect = 5e-26
Identities = 56/124 (45%), Positives = 78/124 (62%), Gaps = 1/124 (0%)
Frame = +1
Query: 151 KYDELQRDMRALRGKEKFG-KTAYDLCLVLNVQIPHKFKVPYFEKYKGNSCPEEHLKMYV 209
K D L+ +RA+ G E + +L LV N+ P KFKV F+KYKG +CP+ HLKMY
Sbjct: 304 KLDHLEEGLRAIEGGEDYAFANLEELFLVPNIITPPKFKVLDFDKYKGTTCPKNHLKMYC 483
Query: 210 RRMSTYARNDQVFIYYFQESLASPASKWYTNLDKTRIQTFCDLSEVFVEQYNYNVDMAPD 269
++M YA+++++ I+ FQESL A WYTNL+ +R+ ++ DL FV QY YN DM
Sbjct: 484 QKMGAYAKDEELLIHSFQESLTGVAVTWYTNLEPSRVHSWKDLMVAFVRQYQYNFDMVLS 663
Query: 270 RSDL 273
R L
Sbjct: 664 RMQL 675
>AW706233
Length = 376
Score = 64.3 bits (155), Expect = 8e-11
Identities = 30/72 (41%), Positives = 46/72 (63%), Gaps = 1/72 (1%)
Frame = -3
Query: 150 EKYDELQRDMRALRGKEKFG-KTAYDLCLVLNVQIPHKFKVPYFEKYKGNSCPEEHLKMY 208
EK D L+ +A+ G + + +L LV N+ P KFKV F+KYKG +CP+ HLKMY
Sbjct: 347 EKLDHLKERFKAIEGGQDYAFANLEELFLVXNIISPPKFKVLNFDKYKGTTCPKNHLKMY 168
Query: 209 VRRMSTYARNDQ 220
++M YA++++
Sbjct: 167 CQKMGAYAKDEK 132
>TC232621 similar to UP|Q8LPN7 (Q8LPN7) AT3g19950/MPN9_19, partial (33%)
Length = 1315
Score = 33.5 bits (75), Expect = 0.15
Identities = 22/84 (26%), Positives = 34/84 (40%), Gaps = 4/84 (4%)
Frame = +1
Query: 30 AQAHAQAQVPPPP----PVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEI 85
A A +Q +PPPP P+ T + +S T+ P S+P P PPF +
Sbjct: 85 ATAPSQFPLPPPPISSAPLATAASSKNSNSQSPTLTLTLPIPSSPTSPSPAPPPFPSSSP 264
Query: 86 LCPIACEAQMPTHQYVAHVPPPPV 109
P +++ PPP+
Sbjct: 265 APPPPLPSRISPPSSETDPTPPPM 336
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 32.7 bits (73), Expect = 0.25
Identities = 27/92 (29%), Positives = 37/92 (39%), Gaps = 3/92 (3%)
Frame = -2
Query: 22 AQVLAQAQAQAHAQAQVPPPPPVRTQVEASSST--ISEWTICADTPTHSAPQHSMPWFPP 79
AQ LA H + PPPPP + T ++ W + P P P PP
Sbjct: 273 AQSLAMCPNPPHLKQDPPPPPPPMPTLPPPPRTQSLARWP---NPPQLKQP----PPLPP 115
Query: 80 FTTGEILCPIACEAQMPTHQYVAH-VPPPPVR 110
E+L + A+ P + H PPPP+R
Sbjct: 114 PPPPELLLLLQSLARCPNSPHW*HAAPPPPLR 19
>TC225011 similar to UP|Q09083 (Q09083) Hydroxyproline-rich glycoprotein
precursor, partial (20%)
Length = 964
Score = 32.7 bits (73), Expect = 0.25
Identities = 29/99 (29%), Positives = 36/99 (36%), Gaps = 1/99 (1%)
Frame = +2
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQ-HSMPWFPPFTTGEILCPIACEAQMPT 97
PPPPP + S P H P H P P+ P P
Sbjct: 257 PPPPPKKPYYYHSPPP----------PVHPHPYPHPHPHPHPY-------PHPHPHPHPH 385
Query: 98 HQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHS 136
YV H PPPP + P + P +H P I P++HS
Sbjct: 386 PPYVYHSPPPPPKKPYKYPS-PPPPVHP-PYIPHPVYHS 496
>TC226825 similar to UP|Q9ZW85 (Q9ZW85) 3-isopropylmalate dehydratase, small
subunit, partial (71%)
Length = 1121
Score = 32.3 bits (72), Expect = 0.33
Identities = 36/111 (32%), Positives = 44/111 (39%), Gaps = 6/111 (5%)
Frame = +3
Query: 23 QVLAQA-QAQAHAQAQVPP----PPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWF 77
Q LA A Q+ + A PP PPP T SS+TIS TPT S F
Sbjct: 135 QTLATASQSLSKPHALNPPRPLLPPPPSTASATSSATIS-------TPTRS--------F 269
Query: 78 PPFTTGEIL-CPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVP 127
PP T+ L P + + PT + P P V + S P VP
Sbjct: 270 PPSTSPSSLPSPTSTRSSAPTPSSASPPPTPRVSSNPARSKPSTPSSSAVP 422
>TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (6%)
Length = 1017
Score = 32.0 bits (71), Expect = 0.43
Identities = 27/82 (32%), Positives = 30/82 (35%)
Frame = +3
Query: 40 PPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQ 99
PPPPV T I+ A TP AP + P P T P
Sbjct: 351 PPPPVSTPPPLPPPKIAPTP--AKTPPAPAPAKATPAPAPAPTKPAPTP----------- 491
Query: 100 YVAHVPPPPVRAPLVVMTYSAP 121
A VPPPP AP V+ AP
Sbjct: 492 --APVPPPPTLAPTPVVEVPAP 551
Score = 31.2 bits (69), Expect = 0.73
Identities = 32/123 (26%), Positives = 45/123 (36%), Gaps = 13/123 (10%)
Frame = +3
Query: 23 QVLAQAQAQAHAQAQVPPPPPVRTQVEASS--STISEWTICADTPTHSAPQHSMPWFPPF 80
Q+ + AQA A A A PP P T + + + T+ A PT S P + PP
Sbjct: 30 QLPSNAQAPATAPANSPPTTPAATSPSSGTPLAVTLPPTVVASPPTTSTPPTTSQ--PPA 203
Query: 81 TTGEILCPIACEAQMPTHQYVAHVPP-------PPVRAPLVVMTYSAPM----IHTVPQI 129
P+ A PP PP +P+V T P+ + T P +
Sbjct: 204 IVAPKPAPVTPPAPKVAPASSPKAPPPQTPQPQPPKVSPVVTPTSPPPIPPPPVSTPPPL 383
Query: 130 EEP 132
P
Sbjct: 384 PPP 392
>TC226466 weakly similar to GB|AAD49970.1|5734705|F24J5 F24J5.4 {Arabidopsis
thaliana;} , partial (22%)
Length = 978
Score = 32.0 bits (71), Expect = 0.43
Identities = 36/128 (28%), Positives = 43/128 (33%), Gaps = 13/128 (10%)
Frame = +1
Query: 18 AQAQAQVLAQAQAQAHAQAQVPP--PPPVRTQVEASSSTISEWTICADTPTHSAPQHSMP 75
A A + Q + A A A VP PP SS T T A PT S P
Sbjct: 13 ALALVSLCCQLPSNAQAPASVPANSPPTTLAATSPSSGTPPAVTQAASPPTTSNPPTISQ 192
Query: 76 WFPPFTTGEILCPI---------ACEAQMPTHQYVAHVPP--PPVRAPLVVMTYSAPMIH 124
PP P+ A + PT Q PP PV AP + P +
Sbjct: 193 --PPAIVAPKPAPVTPPAPKVAPASSPKAPTPQTPPPQPPKVSPVAAPTLPPPLPPPPVA 366
Query: 125 TVPQIEEP 132
T P + P
Sbjct: 367 TPPPLPPP 390
Score = 28.1 bits (61), Expect = 6.2
Identities = 24/82 (29%), Positives = 29/82 (35%)
Frame = +1
Query: 40 PPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQ 99
PPPPV T I+ A T AP + P P T P
Sbjct: 349 PPPPVATPPPLPPPKIAPTP--AKTSPAPAPAKATPAPAPAPTKPAPTP----------- 489
Query: 100 YVAHVPPPPVRAPLVVMTYSAP 121
A +PPPP AP ++ AP
Sbjct: 490 --APIPPPPTLAPTPIVEVPAP 549
>TC206754 weakly similar to UP|GUN4_ARATH (Q9LX31) Tetrapyrrole-binding
protein, chloroplast precursor (Genomes uncoulped 4),
partial (62%)
Length = 1115
Score = 32.0 bits (71), Expect = 0.43
Identities = 20/64 (31%), Positives = 28/64 (43%), Gaps = 3/64 (4%)
Frame = +1
Query: 52 SSTISEWTICADTPTHSAPQH---SMPWFPPFTTGEILCPIACEAQMPTHQYVAHVPPPP 108
S+T + + A TP P H S P PP+TT + + P H+PPPP
Sbjct: 79 SNTTNVHSPNATTPQTPHPPHHFSSNPLHPPYTTTSLSRTLLQTHISPIPSLKPHLPPPP 258
Query: 109 VRAP 112
+ P
Sbjct: 259 PKPP 270
>AI966367 similar to GP|14335028|gb| AT4g37280/C7A10_80 {Arabidopsis
thaliana}, partial (28%)
Length = 491
Score = 32.0 bits (71), Expect = 0.43
Identities = 19/65 (29%), Positives = 29/65 (44%), Gaps = 2/65 (3%)
Frame = +3
Query: 66 THSAPQHSMPW--FPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMI 123
THS P+HS+ + FPP + ++ H PPPP P T+S P+
Sbjct: 36 THSLPKHSVLFSRFPPIVPFSLSRSLSLHHPWEIHPRTTTPPPPPTPPP---ATFSPPIP 206
Query: 124 HTVPQ 128
+ P+
Sbjct: 207 PSTPR 221
>TC225018 homologue to UP|Q39865 (Q39865) Hydroxyproline-rich glycoprotein
(Fragment), partial (19%)
Length = 434
Score = 31.6 bits (70), Expect = 0.56
Identities = 23/85 (27%), Positives = 34/85 (39%), Gaps = 2/85 (2%)
Frame = +1
Query: 96 PTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHSG--NMEVCDRVNDLQEKYD 153
P +Y H PPPP +P ++H +P + HS +C+ EK +
Sbjct: 4 PPPKYYYHSPPPPSPSPPRSHIIITLLLHLLPLLTTIS*HSSMDMKNMCNE*KKAIEKEE 183
Query: 154 ELQRDMRALRGKEKFGKTAYDLCLV 178
E RD R K + G CL+
Sbjct: 184 EALRDAR----KRRIGSKVSAFCLL 246
>TC226955 similar to UP|Q9FQZ6 (Q9FQZ6) Avr9/Cf-9 rapidly elicited protein
146, partial (20%)
Length = 934
Score = 31.2 bits (69), Expect = 0.73
Identities = 23/97 (23%), Positives = 31/97 (31%), Gaps = 6/97 (6%)
Frame = +3
Query: 19 QAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFP 78
Q A+ QA+ Q+H Q V PPPP W P
Sbjct: 201 QHDAETPRQARRQSHCQLNVVPPPP-----------------------------QPRWVP 293
Query: 79 PFTTGEILCPIACEAQMPTH------QYVAHVPPPPV 109
+ +LCP Q+ H + PPPP+
Sbjct: 294 QLSPP*VLCPKGVRVQLQQHAPQLLLPHRREAPPPPL 404
>TC218727 similar to UP|Q8LSC5 (Q8LSC5) Geranylgeranyl pyrophosphate
synthase, partial (55%)
Length = 892
Score = 31.2 bits (69), Expect = 0.73
Identities = 29/95 (30%), Positives = 37/95 (38%), Gaps = 2/95 (2%)
Frame = -1
Query: 40 PPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQ 99
P P TQ +S IS+ + P Q +P PPF E + P Q
Sbjct: 379 PQPETTTQCTWRTSLISQPPRRSHLPLLPPTQPQLPATPPFGARE*I---------PASQ 227
Query: 100 YVAHVPPPPVRAPLVVMTYSAPMIHT--VPQIEEP 132
H PPPP P +V + P T +PQ EP
Sbjct: 226 -APHSPPPPNLYPPLVPPQALPYRSTSLIPQSLEP 125
>TC232055 similar to UP|O49524 (O49524) Pherophorin - like protein, partial
(7%)
Length = 448
Score = 31.2 bits (69), Expect = 0.73
Identities = 27/100 (27%), Positives = 39/100 (39%)
Frame = +2
Query: 37 QVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMP 96
+VP PPP T +SSS S ++ + P P PP + P
Sbjct: 146 RVPNPPPKPT---SSSSPSSPSSVSSGETERRIPPPPPP--PP--------------RKP 268
Query: 97 THQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHS 136
A PPPP P + + + VP++EE +HS
Sbjct: 269 ASNAAAAPPPPPPPPPAKAVRVAPAKVRRVPEVEE-FYHS 385
>TC204047 hydroxyproline-rich glycoprotein
Length = 887
Score = 31.2 bits (69), Expect = 0.73
Identities = 26/104 (25%), Positives = 35/104 (33%)
Frame = +1
Query: 33 HAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACE 92
H PPPPP + S + P HS P P P +
Sbjct: 82 HPVYHSPPPPPKKPYKYPSPPP----PVKKPYPHPHPVYHSPPPPPKKPYKYPSPPPPVK 249
Query: 93 AQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIFHS 136
P V H PPPP + P + P+ P + P++HS
Sbjct: 250 KPYPHPHPVYHSPPPPPKKPYKYPSPPPPVHTYPPHVPTPVYHS 381
>TC226891 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-binding
protein {Arabidopsis thaliana;} , partial (41%)
Length = 1187
Score = 25.0 bits (53), Expect(2) = 1.2
Identities = 22/74 (29%), Positives = 30/74 (39%)
Frame = +3
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTH 98
PPPPP ++ + ++ S A + P H P PP TG+ A +P H
Sbjct: 432 PPPPPPPSRPPSPTTPASPTARSALSSLPPPPSHLSP--PP--TGK------SAATLPPH 581
Query: 99 QYVAHVPPPPVRAP 112
P PP AP
Sbjct: 582 S----PPNPPATAP 611
Score = 23.9 bits (50), Expect(2) = 1.2
Identities = 12/21 (57%), Positives = 12/21 (57%)
Frame = +2
Query: 27 QAQAQAHAQAQVPPPPPVRTQ 47
QAQAQA AQ Q P TQ
Sbjct: 290 QAQAQAQAQTQTSLQPQPETQ 352
>BE020960
Length = 388
Score = 30.4 bits (67), Expect = 1.2
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = +2
Query: 93 AQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVP 127
A P+H H PPP R P+++ T S +IHT P
Sbjct: 74 APQPSHP---HQPPP--RQPIIIYTVSPKVIHTTP 163
>TC228785 weakly similar to GB|AAP42749.1|30984572|BT008736 At3g52060
{Arabidopsis thaliana;} , partial (65%)
Length = 860
Score = 30.4 bits (67), Expect = 1.2
Identities = 23/85 (27%), Positives = 31/85 (36%), Gaps = 8/85 (9%)
Frame = +1
Query: 36 AQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEA-- 93
A PPP S S+ S ++ +PT ++P S P P T +L P + A
Sbjct: 271 ASSPPPSSTTPPTPTSPSSPSTASLSTPSPTPTSPSSSPPPSTPKTPNLLLPPASASASN 450
Query: 94 ------QMPTHQYVAHVPPPPVRAP 112
PT PP V P
Sbjct: 451 TSPSSRSSPTLPSSGSATPPGVATP 525
>TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (40%)
Length = 913
Score = 30.4 bits (67), Expect = 1.2
Identities = 30/119 (25%), Positives = 43/119 (35%)
Frame = +3
Query: 14 TMAEAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHS 73
T A + A A + ++ P P + ASS + T A +PT ++P
Sbjct: 99 TSPPAATTTPFASPATAPSKPKSPAPVASPTSSSPPASSPNAATATPPASSPTVASPPSK 278
Query: 74 MPWFPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
P T P A T V V PP AP+ V + AP+ + P P
Sbjct: 279 AAAPAPVATPPAATPPAATPPAATPPAVTPVSSPP--APVPVSSPPAPVPVSSPPALAP 449
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 30.4 bits (67), Expect = 1.2
Identities = 22/77 (28%), Positives = 29/77 (37%), Gaps = 3/77 (3%)
Frame = +1
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQ---HSMPWFPPFTTGEILCPIACEAQM 95
P PPP V + + + P HS P HS P PP + ++
Sbjct: 136 PSPPPPPPPVFSPPPPVQYYYSSPPPPQHSPPPPPPHSPPPPPPSPPPPVYPYLSPPPPP 315
Query: 96 PTHQYVAHVPPPPVRAP 112
P H PPPPV +P
Sbjct: 316 PVHS-----PPPPVYSP 351
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.130 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,191,774
Number of Sequences: 63676
Number of extensions: 280318
Number of successful extensions: 2819
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 2522
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2751
length of query: 348
length of database: 12,639,632
effective HSP length: 98
effective length of query: 250
effective length of database: 6,399,384
effective search space: 1599846000
effective search space used: 1599846000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC141109.6


