

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
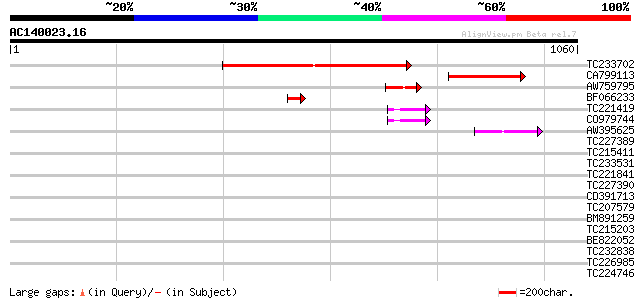
Query= AC140023.16 - phase: 0
(1060 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC233702 similar to UP|Q7X5W7 (Q7X5W7) Homeobox transcription fa... 536 e-152
CA799113 274 2e-73
AW759795 67 4e-11
BF066233 57 4e-08
TC221419 52 1e-06
CO979744 50 7e-06
AW395625 45 1e-04
TC227389 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacety... 35 0.17
TC215411 similar to UP|Q8RVH8 (Q8RVH8) Aux/IAA protein, partial ... 33 0.87
TC233531 similar to UP|Q84U29 (Q84U29) Zinc finger-like protein ... 32 1.1
TC221841 similar to UP|IF39_TOBAC (P56821) Eukaryotic translatio... 32 1.9
TC227390 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacety... 32 1.9
CD391713 31 2.5
TC207579 similar to GB|AAN15531.1|23198008|BT000212 expressed pr... 31 2.5
BM891259 31 3.3
TC215203 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly prote... 30 4.3
BE822052 similar to GP|4557063|gb|A expressed protein {Arabidops... 30 4.3
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 30 4.3
TC226985 similar to UP|Q9LPN1 (Q9LPN1) F2J10.2 protein, partial ... 29 9.6
TC224746 UP|O81975 (O81975) Polyubiquitin (Fragment), complete 29 9.6
>TC233702 similar to UP|Q7X5W7 (Q7X5W7) Homeobox transcription factor
Hox7-like protein, partial (21%)
Length = 1129
Score = 536 bits (1381), Expect = e-152
Identities = 277/353 (78%), Positives = 301/353 (84%)
Frame = +1
Query: 398 RRSLNPLTWIEILRQVLVAAGFGSKQGAFQREGLGKELDILVNYGLCPGTLKCELFKILS 457
RRSLN LTWIEIL QVLVA+GFGSKQG+ + E L KEL++LVNYGLCPGTLK ELF ILS
Sbjct: 7 RRSLNSLTWIEILHQVLVASGFGSKQGSLRGEVLNKELNLLVNYGLCPGTLKSELFNILS 186
Query: 458 ERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITLFEKISSSAYRLRMSTV 517
ERGN GCKV+ELAKSMQIAELNL+ST EELESLI STLSSDITLFEKISS+AYRLRMSTV
Sbjct: 187 ERGNIGCKVAELAKSMQIAELNLASTPEELESLICSTLSSDITLFEKISSTAYRLRMSTV 366
Query: 518 AKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHNSRKAKHNKLKV 577
KD D+S SDTED GSVDDELND+DTCSSGDDF S I+S+ RKL+R NS K+N LKV
Sbjct: 367 MKDGDESHSDTEDFGSVDDELNDTDTCSSGDDFESDPINSSKRKLKRANSH--KNNMLKV 540
Query: 578 YTEIDESHAGEVWLLGLMDSEYSDLKIEEKLNALAALTGLLSSGSSIRMKDPVKVTADCS 637
YTEIDESH GE WLLGLM+SEYSDL IEEKLNALAALT L+SSGSSIRMKD KVTADC+
Sbjct: 541 YTEIDESHPGEAWLLGLMESEYSDLNIEEKLNALAALTDLVSSGSSIRMKDSTKVTADCN 720
Query: 638 SSIQLRGSGAKIKRSVNPIEQMQCTKEVHMNSHACPVDSSLLVSKFHIQEASLEKRKVSA 697
S IQLRGSGAKIKRS ++VH+NS C VDSS L+S+FH EAS K KVS
Sbjct: 721 SGIQLRGSGAKIKRSAVKKPGPLWNQKVHLNSDPCAVDSSSLISRFHTHEASFGKGKVSF 900
Query: 698 YSHPIQSVFLGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEE 750
SHPIQSVFLGSDRRYNRYWLFLGPCN+DDPGHRR+YFESSEDGHWEVIDTEE
Sbjct: 901 ISHPIQSVFLGSDRRYNRYWLFLGPCNVDDPGHRRIYFESSEDGHWEVIDTEE 1059
>CA799113
Length = 435
Score = 274 bits (700), Expect = 2e-73
Identities = 130/145 (89%), Positives = 136/145 (93%), Gaps = 1/145 (0%)
Frame = +1
Query: 820 DNLNLTEIT-DYLPSPGAVVIEAGKKEEEQLHKWIRVQEYDSWIWNSFYLDLNVVKYGRR 878
DNLNLTE D LPS GAVVIEAGKK EEQ+ KWIRVQEYDSWIWNSFYLDLNVVKYG+R
Sbjct: 1 DNLNLTETAEDSLPSAGAVVIEAGKKGEEQIQKWIRVQEYDSWIWNSFYLDLNVVKYGKR 180
Query: 879 SYLDSLARCRSCHDLYWRDERHCKICHMTFELDFDLEEKYAIHIAMCREKEDSNTFPNHK 938
SYLDSLARC+SCHDLYWRDERHCKICHMTFELDFDLEE+YAIHIA CREKEDSNTFPNHK
Sbjct: 181 SYLDSLARCKSCHDLYWRDERHCKICHMTFELDFDLEERYAIHIATCREKEDSNTFPNHK 360
Query: 939 VLPSQIQSLKAAIYAIESVMPEDAL 963
VL SQIQSLKAA+YAIESVMPEDA+
Sbjct: 361 VLSSQIQSLKAAVYAIESVMPEDAM 435
>AW759795
Length = 311
Score = 67.0 bits (162), Expect = 4e-11
Identities = 34/67 (50%), Positives = 45/67 (66%)
Frame = +2
Query: 703 QSVFLGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDR 762
+S+ LG DRR NRYW F+ + +DPG R++ E DG W +ID+EEA ALL+ LD R
Sbjct: 56 RSLPLGQDRRRNRYWQFVASASSNDPGSGRIFVE-YHDGKWRLIDSEEAFDALLTSLDSR 232
Query: 763 GKREALL 769
G RE+ L
Sbjct: 233 GIRESHL 253
>BF066233
Length = 103
Score = 57.0 bits (136), Expect = 4e-08
Identities = 26/33 (78%), Positives = 28/33 (84%)
Frame = +2
Query: 520 DDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGS 552
D D+S SDTEDSGSVDDE N +DTCSSGDDF S
Sbjct: 2 DGDESDSDTEDSGSVDDEFNVADTCSSGDDFES 100
>TC221419
Length = 686
Score = 52.4 bits (124), Expect = 1e-06
Identities = 27/80 (33%), Positives = 47/80 (58%)
Frame = +2
Query: 707 LGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKRE 766
LG DR +NRYW F C + R++ ES + +W ++E L AL+S L+ +G+RE
Sbjct: 122 LGKDRDHNRYWWF---CR-----YGRIFVESCDSKNWGYYSSKEELDALMSSLNCKGERE 277
Query: 767 ALLIESLERRQTSLCRSMSR 786
+L + LE+ +++C + +
Sbjct: 278 RVLRKHLEKYYSTICSELQK 337
>CO979744
Length = 795
Score = 49.7 bits (117), Expect = 7e-06
Identities = 26/80 (32%), Positives = 44/80 (54%)
Frame = -2
Query: 707 LGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKRE 766
LG DR +NRYW F C + R++ ES + +W ++E L L+S L+ +G+RE
Sbjct: 431 LGKDRDHNRYWWF---CR-----YGRIFVESCDSKNWGYYSSKEELDTLMSSLNCKGERE 276
Query: 767 ALLIESLERRQTSLCRSMSR 786
L + LE+ ++C + +
Sbjct: 275 RALQKHLEKYYNTICSELQK 216
>AW395625
Length = 450
Score = 45.4 bits (106), Expect = 1e-04
Identities = 33/128 (25%), Positives = 56/128 (42%), Gaps = 2/128 (1%)
Frame = +3
Query: 870 LNVVKYGRRSYLDSLARCRSCHDLYWRDERHCKICHMTFEL--DFDLEEKYAIHIAMCRE 927
L+ +K+G++ + C C Y+ C C TF K+ +H + +
Sbjct: 18 LSAMKFGKKWCHQLQSICDLCLHAYFSGGAPCSSCCRTFSACKSNPSSSKHIVH-SEGKV 194
Query: 928 KEDSNTFPNHKVLPSQIQSLKAAIYAIESVMPEDALVGAWRKSAHNLWIKRLRRTSTLVE 987
K D + F L +I+ LK + +E +P +AL WR S W +L +S+ +
Sbjct: 195 KIDIDCFHASSSLSLRIRLLKILLSIVEVTLPLEALQPLWRDSCRKSWSTKLDASSSSED 374
Query: 988 LLQVLADF 995
LLQ + F
Sbjct: 375 LLQDIISF 398
>TC227389 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (63%)
Length = 1144
Score = 35.0 bits (79), Expect = 0.17
Identities = 21/52 (40%), Positives = 31/52 (59%), Gaps = 6/52 (11%)
Frame = +3
Query: 504 KISSSAYRLRMSTVAKD---DDDSQSDTEDSGSVDDELNDSDTCS---SGDD 549
K S SA ++++ KD D D SD +D GS D+E+ D+D+ S SGD+
Sbjct: 492 KASGSAKQVKVVEPKKDNEEDSDDDSDDDDFGSSDEEMEDADSDSDDESGDE 647
>TC215411 similar to UP|Q8RVH8 (Q8RVH8) Aux/IAA protein, partial (87%)
Length = 1723
Score = 32.7 bits (73), Expect = 0.87
Identities = 25/91 (27%), Positives = 37/91 (40%)
Frame = +1
Query: 742 HWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSMSRIKVSNIGMGCMSHSD 801
+W V D LL + ++ G+ L+ S S+C + S++K N M SD
Sbjct: 169 NWVVFD*TVMSKPLLGIAEEEGQSNVTLLVSSSATMESVCLNSSKLKERNY----MGLSD 336
Query: 802 QSELDRVAEDSCSPVSDVDNLNLTEITDYLP 832
S +D A NL TE+ LP
Sbjct: 337 CSSVDSSAPSFSDETKSNLNLKATELRLGLP 429
>TC233531 similar to UP|Q84U29 (Q84U29) Zinc finger-like protein (Fragment),
partial (68%)
Length = 926
Score = 32.3 bits (72), Expect = 1.1
Identities = 29/100 (29%), Positives = 47/100 (47%), Gaps = 13/100 (13%)
Frame = -3
Query: 462 NGCKVSELAKSMQIAELN-----LSSTTEELESLIYS----TLSSDITLFEKISSSAYRL 512
+G + L ++M +L +SS+T S ++S + SS +TL++ IS SA
Sbjct: 879 HGINAAPLCRTMHTQQLYSTTLPISSSTPPFSSSLFSYSATSSSSSLTLYKSISCSA--- 709
Query: 513 RMSTVAKDDDDSQSDTEDSGS----VDDELNDSDTCSSGD 548
+ S SDT + S +EL + TCS+GD
Sbjct: 708 -------SSNKSISDTPTTASWLFSPSEELPTASTCSTGD 610
>TC221841 similar to UP|IF39_TOBAC (P56821) Eukaryotic translation initiation
factor 3 subunit 9 (eIF-3 eta) (eIF3 p110) (eIF3b),
partial (50%)
Length = 1207
Score = 31.6 bits (70), Expect = 1.9
Identities = 23/104 (22%), Positives = 47/104 (45%), Gaps = 3/104 (2%)
Frame = +1
Query: 491 IYSTLSSDITLFEKISSSAYRLRMSTVAKDDDDSQSDT-EDSGSV--DDELNDSDTCSSG 547
+ T+ +D+ + ++I ++ RL + D D + T ED G V D+E+ +
Sbjct: 55 VLPTVMADVMIMKEIEDTSLRLGVDLSTLDLDAIRLPTGEDCGIVSDDEEVYQEENLEFE 234
Query: 548 DDFGSGSIHSNIRKLRRHNSRKAKHNKLKVYTEIDESHAGEVWL 591
FG+ + N+ + R K + K+Y++I +W+
Sbjct: 235 SGFGNIIVVDNLPVVPREKFEKLEGVVRKIYSQIGVIKEDGLWM 366
>TC227390 similar to UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase
HD2-P39, partial (24%)
Length = 808
Score = 31.6 bits (70), Expect = 1.9
Identities = 19/50 (38%), Positives = 30/50 (60%), Gaps = 5/50 (10%)
Frame = +1
Query: 504 KISSSAYRLRMSTVAKD-----DDDSQSDTEDSGSVDDELNDSDTCSSGD 548
K S+SA ++++ KD DDDS D +D GS D+E+ D+D+ S +
Sbjct: 121 KASASAKQVKIVEPKKDNEEDSDDDSDED-DDFGSSDEEMEDADSDSDDE 267
>CD391713
Length = 594
Score = 31.2 bits (69), Expect = 2.5
Identities = 16/50 (32%), Positives = 29/50 (58%)
Frame = +1
Query: 477 ELNLSSTTEELESLIYSTLSSDITLFEKISSSAYRLRMSTVAKDDDDSQS 526
E+ L +T L+S +YST ++ I +F+K SS++ + DD D+ +
Sbjct: 19 EIRL*VSTRHLQSSMYSTAAASIYIFQKNSSNSEI*KKKKKTSDDTDTNT 168
>TC207579 similar to GB|AAN15531.1|23198008|BT000212 expressed protein
{Arabidopsis thaliana;} , partial (57%)
Length = 1249
Score = 31.2 bits (69), Expect = 2.5
Identities = 26/133 (19%), Positives = 58/133 (43%), Gaps = 13/133 (9%)
Frame = +2
Query: 448 LKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTEEL-------ESLIYSTLSSDIT 500
L+ E + +E + ++EL ++ + LS ++L E+L+ S T
Sbjct: 152 LQSESSALRAELADRNRLIAELQSQVESIDAALSEAADKLARADQDKENLLKENASLSNT 331
Query: 501 LFEKISSSAYRLR------MSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGS 554
+ K++ +L M ++ +D+D S+ + + + + + + T GDD S +
Sbjct: 332 V-RKLTRDVSKLETFRKTLMKSLREDEDTSEGTADTAAKLHSQASFTSTSQFGDDDASST 508
Query: 555 IHSNIRKLRRHNS 567
+ S +R + S
Sbjct: 509 LSSRTSSMRINTS 547
>BM891259
Length = 437
Score = 30.8 bits (68), Expect = 3.3
Identities = 21/90 (23%), Positives = 40/90 (44%), Gaps = 1/90 (1%)
Frame = +1
Query: 479 NLSSTTEELESLIYSTLSSDITLFEKISSSAYRL-RMSTVAKDDDDSQSDTEDSGSVDDE 537
+L +E S ++S+ + +L ++ L R+S+ DD+ + SD SVD
Sbjct: 37 DLPEIHQERTSTLHSSSTPSGSLNGMVNEQHVNLERISSTGSDDESASSDEGSVDSVDQS 216
Query: 538 LNDSDTCSSGDDFGSGSIHSNIRKLRRHNS 567
L T S + S +++R +N+
Sbjct: 217 LMSPSTRESSNALSSEQSQTDVRTTSHNNT 306
>TC215203 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly protein 1-like
protein 1, partial (31%)
Length = 729
Score = 30.4 bits (67), Expect = 4.3
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 2/67 (2%)
Frame = +2
Query: 485 EELESLIYSTLSSDITLFEKISSSAYRLRMSTVAKDD--DDSQSDTEDSGSVDDELNDSD 542
EELE+L+ T+ +KI A A+ D D ++D ED DD+ D D
Sbjct: 113 EELENLMEHDYDIGSTIRDKIIPHAVSWFTGEAAQSDFEDIDEADYEDEEGDDDDDEDED 292
Query: 543 TCSSGDD 549
G+D
Sbjct: 293 EDEDGED 313
>BE822052 similar to GP|4557063|gb|A expressed protein {Arabidopsis
thaliana}, partial (28%)
Length = 668
Score = 30.4 bits (67), Expect = 4.3
Identities = 12/40 (30%), Positives = 19/40 (47%)
Frame = -1
Query: 510 YRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDD 549
YRL D+D+ + D D+ +DE D D G++
Sbjct: 545 YRLNKDASDSDEDEDEEDDHDTNDQNDEEGDEDFSGEGEE 426
Score = 29.3 bits (64), Expect = 9.6
Identities = 15/49 (30%), Positives = 21/49 (42%)
Frame = -1
Query: 518 AKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHN 566
+ DDDDS D +D DD+ D D D+ + K R+ N
Sbjct: 380 SNDDDDSDEDDDDDNDGDDDGEDEDEKEGEDEEDEEEVPQPPTKKRK*N 234
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 30.4 bits (67), Expect = 4.3
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +2
Query: 520 DDDDSQSDTEDSGSVDDELNDSDTCSSGDD 549
+DD+ + D + G DD+ NDSD S D+
Sbjct: 434 EDDEEEGDVQRGGEPDDDDNDSDDDSDDDE 523
>TC226985 similar to UP|Q9LPN1 (Q9LPN1) F2J10.2 protein, partial (9%)
Length = 797
Score = 29.3 bits (64), Expect = 9.6
Identities = 13/45 (28%), Positives = 21/45 (45%)
Frame = -2
Query: 910 LDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLKAAIYAI 954
+ LE+K+A +C E N PNH+ + I SL ++
Sbjct: 286 ISLSLEQKFAYLC*ICSEGYSGNRLPNHQTVHLHISSLSPCFLSV 152
>TC224746 UP|O81975 (O81975) Polyubiquitin (Fragment), complete
Length = 1000
Score = 29.3 bits (64), Expect = 9.6
Identities = 17/65 (26%), Positives = 34/65 (52%), Gaps = 1/65 (1%)
Frame = +3
Query: 738 SEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQT-SLCRSMSRIKVSNIGMGC 796
+ DG W+V+ EEA+ + + D + L+ E+ +R + ++ + + +SN G
Sbjct: 288 ASDGLWDVVSNEEAVAMIKPIEDAEEAAKRLMQEAYQRGSSDNITCVVDAVALSN--QGA 461
Query: 797 MSHSD 801
SHS+
Sbjct: 462 SSHSN 476
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 49,755,650
Number of Sequences: 63676
Number of extensions: 733502
Number of successful extensions: 3784
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 3579
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3754
length of query: 1060
length of database: 12,639,632
effective HSP length: 107
effective length of query: 953
effective length of database: 5,826,300
effective search space: 5552463900
effective search space used: 5552463900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC140023.16


