

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140022.1 + phase: 0 /pseudo
(583 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
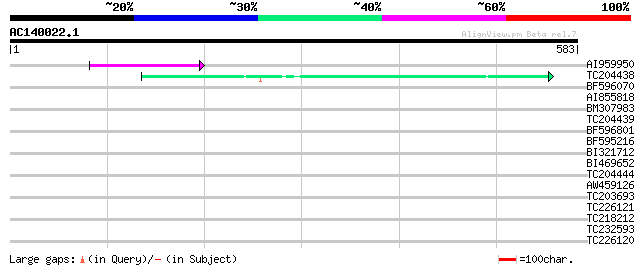
Score E
Sequences producing significant alignments: (bits) Value
AI959950 50 4e-06
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 43 4e-04
BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vi... 39 0.005
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 37 0.032
BM307983 37 0.032
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 37 0.032
BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Gl... 34 0.16
BF595216 33 0.27
BI321712 33 0.35
BI469652 weakly similar to GP|18149115|dbj| reverse transcriptas... 31 1.3
TC204444 homologue to UP|CADH_MEDSA (P31656) Cinnamyl-alcohol de... 29 5.1
AW459126 29 6.7
TC203693 similar to UP|Q949G4 (Q949G4) N3 like protein, partial ... 29 6.7
TC226121 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, ... 28 8.7
TC218212 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (11%) 28 8.7
TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (F... 28 8.7
TC226120 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, ... 28 8.7
>AI959950
Length = 466
Score = 49.7 bits (117), Expect = 4e-06
Identities = 43/118 (36%), Positives = 59/118 (49%)
Frame = -2
Query: 83 *KLWMKKSMQLRRIKHGS*LNYRKTRSQ*E*SGCTRQSTNPVVRLIAIKRGWWLKATNKN 142
*K K + +RI S LNY+K R E*+G + +VRL K+ LK T+
Sbjct: 396 *KRCKKNLISFKRIMSRSSLNYQKERR*LE*NGYFVTN*TRMVRL*DTKQD*LLKVTHNR 217
Query: 143 QVLIILKYLLLLQD*IQFACLFHSQLKITGKYIKWMLSLHFLMVLWKKKCMLSSLQDM 200
+V K L LL * +A FH Q + IKWM +HF M K+K ML++ D+
Sbjct: 216 KV*TTQKPLHLLHV*K*YASYFHLQPIVI*SCIKWM*KVHF*MA*SKRKFMLNNRLDL 43
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 42.7 bits (99), Expect = 4e-04
Identities = 116/432 (26%), Positives = 176/432 (39%), Gaps = 8/432 (1%)
Frame = +2
Query: 136 LKATNKNQVLIILKYLLLLQD*IQFACLFHSQLKITGKYIKWMLSLHFLMVLWKKKCMLS 195
LKAT + +V ++K LL D C +WM F M KK M S
Sbjct: 3404 LKATLRLKV*TLMKLSPLLLDLSPSDCYLV*LASSNSSCTRWM*RARF*MDT*MKKPMWS 3583
Query: 196 SLQDMWLEERRIKYID*RKHCMA*SRRQEHGTKR-LILILFKTVFRDVHSSTHSTSNSLI 254
S +D+ ++ +I Y R+ M *S+ QE G K +L K + R+ + S SN ++
Sbjct: 3584 SQRDL*IQLIQIMYTGSRRLSMD*SKLQELGMKG*QSSLLSKGIGRE-ELTRLSLSNKML 3760
Query: 255 LE-------MFLLCASMSMI*YSPVTIQR*SLNSGRL**VILK*QIWA*CPIFSALRSFN 307
+ LC + + R +LN L *V+L+ + IF +
Sbjct: 3761 KT***HRYMLMTLCLEGCRMRCFDILSNRCNLN---LR*VLLES*L-----IFWDSK*SR 3916
Query: 308 RRM*SLSLRRSMQVIF*RNLRWSIQSQFPRRLKKS*S*QEKAMVKG*TQLITKV*LEV*D 367
+ S + SMQ R+L W + + +*S Q+ + ++ T+ *L
Sbjct: 3917 WKTPYSSHKASMQRTLSRSLGWKMPAIKEHLHLLT*SCQKMKLAPVLIKVCTEA*LGAYY 4096
Query: 368 I*LQQGQI*YMELVYLANTWRIRVLVTCKEPRGFFVILKVL*PKEFFMVIIVM*SLLDIQ 427
I* M+ V++ + I VT + R F + VI+ + L I
Sbjct: 4097 I*QLADLTSPMQ*VFVQDIKPILR*VT*IK*REF*NM*MAPVTMGLCTVIVQIQCWLGIV 4276
Query: 428 IVIGQEIQKQEKARQGTHFI*EPVQYHGLRRNNLWSLFQQQK*NI**QPVVLLKQCG*EE 487
++IG E+Q EKA I EP+ +HG R+ + K +I Q + G*
Sbjct: 4277 MLIGLEVQMTEKALLVDVSIWEPILFHGSARSRTVCPYLLLKQSILQQEAAVHN*FG*SR 4456
Query: 488 F*K*CIMSRTLLQRYIVITSQQLH*AKIQFFMDGPSILTSGFTRYES*LLRKKW*SSTVP 547
*+ MS + V T L KI F PS LT T E L+ K S +
Sbjct: 4457 C*R-STMSNKMS*HCTVTT*VLLIFLKILFNTAEPSTLTLDITILEILLMIKLSHWSMLT 4633
Query: 548 LKSKLQIFLQSH 559
L++K QIF Q H
Sbjct: 4634 LRNK*QIFSQRH 4669
>BF596070 similar to GP|27901698|gb gag-pol polyprotein {Vitis vinifera},
partial (34%)
Length = 407
Score = 39.3 bits (90), Expect = 0.005
Identities = 25/76 (32%), Positives = 38/76 (49%)
Frame = -3
Query: 115 GCTRQSTNPVVRLIAIKRGWWLKATNKNQVLIILKYLLLLQD*IQFACLFHSQLKITGKY 174
G T P+VRLI ++ W LKAT++ LI + LL + F C L +TG
Sbjct: 366 GSTLLKLGPLVRLIGLRLVW*LKATHRYMALITVILSLLSPNSPLFVCFLLWLLFVTGPS 187
Query: 175 IKWMLSLHFLMVLWKK 190
I +L + F V+ ++
Sbjct: 186 ISLILRMPFSTVILRR 139
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 36.6 bits (83), Expect = 0.032
Identities = 25/73 (34%), Positives = 40/73 (54%)
Frame = -1
Query: 149 KYLLLLQD*IQFACLFHSQLKITGKYIKWMLSLHFLMVLWKKKCMLSSLQDMWLEERRIK 208
K++L LQD* C H IKWML + F MV +KKK ML++ Q + ++
Sbjct: 265 KHMLQLQD*KSLECF*HMYP**ILNSIKWMLKVLF*MV*FKKKYMLNNPQALKSRINQLM 86
Query: 209 YID*RKHCMA*SR 221
+I+ ++ M *++
Sbjct: 85 FINCKRLFMV*NK 47
>BM307983
Length = 406
Score = 36.6 bits (83), Expect = 0.032
Identities = 29/89 (32%), Positives = 42/89 (46%), Gaps = 1/89 (1%)
Frame = +3
Query: 130 IKRGWWLKATNKNQVLIILKYLLL-LQD*IQFACLFHSQLKITGKYIKWMLSLHFLMVLW 188
I+RGW + T+K I+ ++LL + Q L S+ + G+ I ML + M W
Sbjct: 60 IRRGWLQRDTSKPMGSIMRRHLLSGKNEYSQDHHLLSSKHNLVGRCINLMLKMPSYMEAW 239
Query: 189 KKKCMLSSLQDMWLEERRIKYID*RKHCM 217
KKK DM IK+ D*R+ M
Sbjct: 240 KKKYTWRFHLDMVPVMEEIKFAD*RRPYM 326
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 36.6 bits (83), Expect = 0.032
Identities = 47/142 (33%), Positives = 63/142 (44%)
Frame = +2
Query: 416 VIIVM*SLLDIQIVIGQEIQKQEKARQGTHFI*EPVQYHGLRRNNLWSLFQQQK*NI**Q 475
VI+ + L I ++IG E+Q EKA I E +HG R+ + QQK +I Q
Sbjct: 4238 VIVQIQCWLGIVMLIGLEVQMTEKALLVDASIWETTLFHGSARSRTVCPYLQQKPSILQQ 4417
Query: 476 PVVLLKQCG*EEF*K*CIMSRTLLQRYIVITSQQLH*AKIQFFMDGPSILTSGFTRYES* 535
+ G* *+ MS + V T L KI F PS LT T E
Sbjct: 4418 EAAVHS*FG*SRC*R-STMSNKMS*HCTVTT*VLLIFLKILFNTAEPSTLTLDITISEIL 4594
Query: 536 LLRKKW*SSTVPLKSKLQIFLQ 557
L+ K S + L++K QIF Q
Sbjct: 4595 LMIK*SH*SMLTLRNK*QIFSQ 4660
Score = 34.7 bits (78), Expect = 0.12
Identities = 34/115 (29%), Positives = 48/115 (41%)
Frame = +2
Query: 114 SGCTRQSTNPVVRLIAIKRGWWLKATNKNQVLIILKYLLLLQD*IQFACLFHSQLKITGK 173
SG +R V + W LKAT + +V +++ L L D +
Sbjct: 3335 SGSSRTKPMKKVS*PETRPDWLLKATLRLKV*TLMRLLPQLLDLSPSDYYLV*LVSSNSS 3514
Query: 174 YIKWMLSLHFLMVLWKKKCMLSSLQDMWLEERRIKYID*RKHCMA*SRRQEHGTK 228
+WM HF M KK M SS +D+ +I Y R+ M *S+ QE G K
Sbjct: 3515 CTRWM*RAHF*MDT*MKKSMWSSQRDLQTRLIQIMYTGSRRLSMD*SKLQELGMK 3679
>BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Glycine max},
partial (7%)
Length = 336
Score = 34.3 bits (77), Expect = 0.16
Identities = 35/106 (33%), Positives = 48/106 (45%)
Frame = +1
Query: 74 KKPRVMKIG*KLWMKKSMQLRRIKHGS*LNYRKTRSQ*E*SGCTRQSTNPVVRLIAIKRG 133
KKP M IG K L+ +G+* K E +G + +V L+ IK G
Sbjct: 19 KKP**MIIGSLSCKKN*TNLKETMYGN**KNLKIILLLEQNGFLEIN*MNMV*LLEIKPG 198
Query: 134 WWLKATNKNQVLIILKYLLLLQD*IQFACLFHSQLKITGKYIKWML 179
K K + + K++LLLQD* C +H T +IKWML
Sbjct: 199 **RKDIIKKRE*TMKKHMLLLQD*KPLECFWHMHP**TLNFIKWML 336
>BF595216
Length = 421
Score = 33.5 bits (75), Expect = 0.27
Identities = 16/33 (48%), Positives = 22/33 (66%)
Frame = +1
Query: 170 ITGKYIKWMLSLHFLMVLWKKKCMLSSLQDMWL 202
ITG + WM + FLMV +K+K + SSLQ + L
Sbjct: 280 ITGPFSSWM*IMPFLMVFYKRKSICSSLQVLTL 378
>BI321712
Length = 399
Score = 33.1 bits (74), Expect = 0.35
Identities = 38/115 (33%), Positives = 53/115 (46%)
Frame = -1
Query: 261 CASMSMI*YSPVTIQR*SLNSGRL**VILK*QIWA*CPIFSALRSFNRRM*SLSLRRSMQ 320
C M M * TIQ S +S ++ + L+*+IW I SA + + S +++M
Sbjct: 384 CVCM*MT*SLQGTIQACSKSSRKICQMNLR*RIWGSWHIISASK*NKKTKEFSSPKKAMP 205
Query: 321 VIF*RNLRWSIQSQFPRRLKKS*S*QEKAMVKG*TQLITKV*LEV*DI*LQQGQI 375
R+ RW Q R + S* + QL TKV*LEV * QG+I
Sbjct: 204 KKSLRSSRWMTPIQLAPRWNVAAS*ASMKKERMWIQLFTKV*LEVYVT*HVQGRI 40
>BI469652 weakly similar to GP|18149115|dbj| reverse transcriptase {Silene
noctiflora}, partial (60%)
Length = 427
Score = 31.2 bits (69), Expect = 1.3
Identities = 16/37 (43%), Positives = 22/37 (59%)
Frame = +1
Query: 164 FHSQLKITGKYIKWMLSLHFLMVLWKKKCMLSSLQDM 200
F QL T + KW+L + F M L K+KCM +LQ +
Sbjct: 220 FPLQLIKT*IFFKWILKVVF*MTLLKRKCMSDNLQTL 330
>TC204444 homologue to UP|CADH_MEDSA (P31656) Cinnamyl-alcohol dehydrogenase
(CAD) , partial (22%)
Length = 790
Score = 29.3 bits (64), Expect = 5.1
Identities = 17/62 (27%), Positives = 34/62 (54%), Gaps = 3/62 (4%)
Frame = +1
Query: 140 NKNQVLIILKYLLLLQD*IQFACLF---HSQLKITGKYIKWMLSLHFLMVLWKKKCMLSS 196
N N +++ KY++L+Q *+ F ++ + ITG +I M ++ WK+K + S
Sbjct: 391 NGNTIVLYQKYIILIQ-*LHFNDVYLGCEGRKSITGSFIGSMKETEEMLEFWKEKGLSSM 567
Query: 197 LQ 198
++
Sbjct: 568 IE 573
>AW459126
Length = 339
Score = 28.9 bits (63), Expect = 6.7
Identities = 16/59 (27%), Positives = 30/59 (50%)
Frame = +1
Query: 194 LSSLQDMWLEERRIKYID*RKHCMA*SRRQEHGTKRLILILFKTVFRDVHSSTHSTSNS 252
++ L WLE R+I+Y K A +++Q +K I+ K + + S +T+N+
Sbjct: 67 INDLTGKWLESRQIRYN*ATKRASASNKKQSSNSK----IIIKLINKSSEESXETTNNN 231
>TC203693 similar to UP|Q949G4 (Q949G4) N3 like protein, partial (90%)
Length = 1795
Score = 28.9 bits (63), Expect = 6.7
Identities = 19/60 (31%), Positives = 30/60 (49%)
Frame = +3
Query: 137 KATNKNQVLIILKYLLLLQD*IQFACLFHSQLKITGKYIKWMLSLHFLMVLWKKKCMLSS 196
K+T +N + + +L+LL +Q C + L GK + L L L +LW + LSS
Sbjct: 204 KSTRRNPLKVSSHFLMLLHCSVQ--CFGFTMLS*KGKLPSFSLPLTHLELLWSQFTFLSS 377
>TC226121 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, partial
(18%)
Length = 805
Score = 28.5 bits (62), Expect = 8.7
Identities = 13/27 (48%), Positives = 15/27 (55%)
Frame = -3
Query: 162 CLFHSQLKITGKYIKWMLSLHFLMVLW 188
C HS L I G W SLH+L+ LW
Sbjct: 698 CSIHSLLHILG----WHYSLHYLLALW 630
>TC218212 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (11%)
Length = 712
Score = 28.5 bits (62), Expect = 8.7
Identities = 12/49 (24%), Positives = 24/49 (48%)
Frame = +2
Query: 114 SGCTRQSTNPVVRLIAIKRGWWLKATNKNQVLIILKYLLLLQD*IQFAC 162
SGC + + L+ I GWWL+A + ++ + Y++ + + C
Sbjct: 518 SGCLQ*K*K*YINLVFINTGWWLEAKASHPFVVRMDYVVQKIEFVNIHC 664
>TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (Fragment),
partial (16%)
Length = 562
Score = 28.5 bits (62), Expect = 8.7
Identities = 36/106 (33%), Positives = 55/106 (50%), Gaps = 1/106 (0%)
Frame = +2
Query: 274 IQR*SLNSGRL**VILK*QIWA*CPIFSALRSFNRRM*SLSLRRSMQVIF*RNLRWSIQS 333
+Q * +S + * +LK* I IF LRS R S++ +MQ F*R+ +W +
Sbjct: 239 MQG*LRSSSKK*CKLLK*LILVS*LIFLELRSSKVRTKC*SVKGNMQKKF*RSFKWRNAN 418
Query: 334 QFPRR-LKKS*S*QEKAMVKG*TQLITKV*LEV*DI*LQQGQI*YM 378
+ +K+ S + ++K +I *L+V* I LQQGQ Y+
Sbjct: 419 LLAHQ*IKRRSSTR*TVLIKLMKDII-GA*LDV*CISLQQGQTFYL 553
>TC226120 similar to UP|Q9LL85 (Q9LL85) DNA-binding protein p24, partial
(50%)
Length = 642
Score = 28.5 bits (62), Expect = 8.7
Identities = 13/27 (48%), Positives = 15/27 (55%)
Frame = -1
Query: 162 CLFHSQLKITGKYIKWMLSLHFLMVLW 188
C HS L I G W SLH+L+ LW
Sbjct: 498 CSIHSLLHILG----WHYSLHYLLALW 430
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.369 0.164 0.579
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,042,367
Number of Sequences: 63676
Number of extensions: 345793
Number of successful extensions: 4653
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1977
Number of HSP's successfully gapped in prelim test: 232
Number of HSP's that attempted gapping in prelim test: 2651
Number of HSP's gapped (non-prelim): 2286
length of query: 583
length of database: 12,639,632
effective HSP length: 102
effective length of query: 481
effective length of database: 6,144,680
effective search space: 2955591080
effective search space used: 2955591080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC140022.1


