

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139882.5 - phase: 0
(379 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
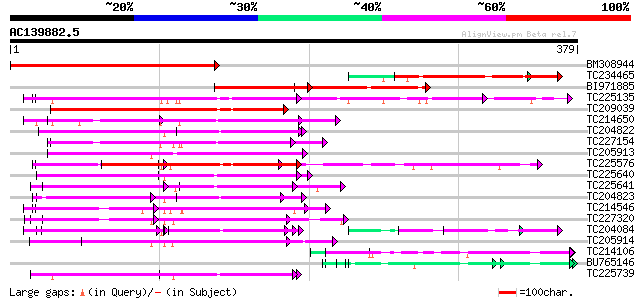
Score E
Sequences producing significant alignments: (bits) Value
BM308944 279 2e-75
TC234465 similar to GB|AAA91085.1|561756|MUSHDH05 Huntington dis... 152 2e-37
BI971885 137 6e-33
TC225135 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, ... 123 1e-28
TC209039 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, ... 115 3e-26
TC214650 similar to UP|Q9LJH8 (Q9LJH8) RNA binding protein nucle... 110 1e-24
TC204822 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, part... 101 5e-22
TC227154 similar to UP|Q9LJH8 (Q9LJH8) RNA binding protein nucle... 100 8e-22
TC205913 weakly similar to UP|Q9LEB4 (Q9LEB4) RNA Binding Protei... 100 1e-21
TC225576 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, ... 98 7e-21
TC225640 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, part... 97 1e-20
TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding pr... 94 1e-19
TC204823 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, part... 92 5e-19
TC214546 similar to PIR|T01932|T01932 RNA binding protein homolo... 92 5e-19
TC227320 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding prot... 91 1e-18
TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, co... 88 7e-18
TC205914 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, ... 87 9e-18
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 87 1e-17
BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvo... 85 6e-17
TC225739 homologue to UP|Q41124 (Q41124) Chloroplast RNA binding... 84 8e-17
>BM308944
Length = 429
Score = 279 bits (713), Expect = 2e-75
Identities = 138/140 (98%), Positives = 139/140 (98%)
Frame = +3
Query: 1 MTTRIAPGVGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDR 60
MTTRIAPGVGANLLGQH+AERNQDATAYVGNLDPQI EELLWELFVQAGPVVNVYVPKDR
Sbjct: 9 MTTRIAPGVGANLLGQHAAERNQDATAYVGNLDPQICEELLWELFVQAGPVVNVYVPKDR 188
Query: 61 VTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLD 120
VTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLD
Sbjct: 189 VTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLD 368
Query: 121 PDVDEKLLYDTFSAFGVIVT 140
PDVDEKLLYDTFSAFGVIVT
Sbjct: 369 PDVDEKLLYDTFSAFGVIVT 428
>TC234465 similar to GB|AAA91085.1|561756|MUSHDH05 Huntington disease protein
{Mus musculus;} , partial (12%)
Length = 545
Score = 152 bits (385), Expect = 2e-37
Identities = 77/115 (66%), Positives = 84/115 (72%), Gaps = 3/115 (2%)
Frame = +3
Query: 258 IHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPP-MQQFRPPH-SMQMPP 315
I ++PPPPQA AFQPM P PGQ WHQQQ GQ M+ MPPP +QQFRPPH MQ P
Sbjct: 6 IPTLRPPPPQAAAFQPM--PVPGQPAWHQQQPGQTMLSPAMPPPQIQQFRPPHPGMQQMP 179
Query: 316 PPPPQGMQGPPRPLPPPSVMAGQ-QQVWRPPPPPQQQGGHPMGYPQNSMPPPPPP 369
PPP Q PPRPLPPPSVMA Q VWR PPPPQQQGG P+ YPQ++M PPPPP
Sbjct: 180 PPP----QAPPRPLPPPSVMASQPPPVWR-PPPPQQQGGRPLPYPQSAMLPPPPP 329
Score = 52.8 bits (125), Expect = 3e-07
Identities = 43/143 (30%), Positives = 51/143 (35%), Gaps = 20/143 (13%)
Frame = +3
Query: 227 PPSLPNAPQANGTIPAPVPPRP-----------FANGVAPPPIHVIQPP---------PP 266
P P PQA P PVP +P + + PP I +PP PP
Sbjct: 9 PTLRPPPPQAAAFQPMPVPGQPAWHQQQPGQTMLSPAMPPPQIQQFRPPHPGMQQMPPPP 188
Query: 267 QAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPP 326
QAP P +PPP + PPP+ + PPPPQ G P
Sbjct: 189 QAP---PRPLPPPS-------------VMASQPPPVWR-----------PPPPQQQGGRP 287
Query: 327 RPLPPPSVMAGQQQVWRPPPPPQ 349
P P Q PPPPPQ
Sbjct: 288 LPYP--------QSAMLPPPPPQ 332
Score = 38.5 bits (88), Expect = 0.005
Identities = 35/120 (29%), Positives = 46/120 (38%), Gaps = 3/120 (2%)
Frame = +3
Query: 202 PAERVLAASNPTAQKSR---PHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPI 258
P + +L+ + P Q + PH PP P AP P P+PP P PPP+
Sbjct: 93 PGQTMLSPAMPPPQIQQFRPPHPGMQQMPPP-PQAP------PRPLPP-PSVMASQPPPV 248
Query: 259 HVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPP 318
+PPPP QQQ G+P+ P S +PPPPP
Sbjct: 249 W--RPPPP-------------------QQQGGRPLPY------------PQSAMLPPPPP 329
>BI971885
Length = 421
Score = 137 bits (346), Expect = 6e-33
Identities = 65/65 (100%), Positives = 65/65 (100%)
Frame = +1
Query: 138 IVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGE 197
IVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGE
Sbjct: 1 IVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGE 180
Query: 198 RHGTP 202
RHGTP
Sbjct: 181 RHGTP 195
Score = 128 bits (322), Expect = 4e-30
Identities = 65/91 (71%), Positives = 71/91 (77%)
Frame = +2
Query: 191 KKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFA 250
+K +G AERVLAASNPT QKSRPHTLFASGPP+LP+ PQANG APVPPRPFA
Sbjct: 161 RKTLRGNGMVPRAERVLAASNPTTQKSRPHTLFASGPPTLPSVPQANGV--APVPPRPFA 334
Query: 251 NGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQ 281
NGVAPP I ++PPPPQA AFQP MP PGQ
Sbjct: 335 NGVAPPAIPALRPPPPQAAAFQP--MPVPGQ 421
>TC225135 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(97%)
Length = 2392
Score = 123 bits (309), Expect = 1e-28
Identities = 73/189 (38%), Positives = 111/189 (58%), Gaps = 3/189 (1%)
Frame = +3
Query: 10 GANLLGQHSAERNQDATA--YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQG 67
G N G NQ T YVG+LDP +++ L++LF Q G VV+V V +D + + G
Sbjct: 129 GPNGGGGAGGAGNQFVTTSLYVGDLDPNVTDAQLYDLFNQLGQVVSVRVCRDLTSRRSLG 308
Query: 68 YGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEK 126
YG+V F + +DA A+ VLN L +PIR+ + +D G N+FI NLD +D K
Sbjct: 309 YGYVNFSNPQDAARALDVLNFTPLNNRPIRIMYSHRDPSIRKSGQGNIFIKNLDRAIDHK 488
Query: 127 LLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITV 186
L+DTFS FG I+ + K+ D +G S+G+GF+ +D+ E++ AIE +NG L ++Q+ V
Sbjct: 489 ALHDTFSTFGNIL-SCKVATD-SSGQSKGYGFVQFDNEESAQKAIEKLNGMLLNDKQVYV 662
Query: 187 SYAYKKDTK 195
+K +
Sbjct: 663 GPFLRKQER 689
Score = 98.6 bits (244), Expect = 4e-21
Identities = 91/335 (27%), Positives = 151/335 (44%), Gaps = 31/335 (9%)
Frame = +3
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
+ +A++ + +V NL +++ L F + G + + V +D + + +GFV F +
Sbjct: 690 ESAADKAKFNNVFVKNLSESTTDDELKNTFGEFGTITSAVVMRDG-DGKSKCFGFVNFEN 866
Query: 76 EEDADYAIKVLNMIKLYGKPIRVNKASQD---------------KKSLDV--GANLFIGN 118
+DA A++ LN K K V KA + K++ D GANL++ N
Sbjct: 867 ADDAARAVEALNGKKFDDKEWYVGKAQKKSERENELKQRFEQSMKEAADKYQGANLYVKN 1046
Query: 119 LDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQY 178
LD + ++ L + FS FG I T+ K+MRDP+ G SRG GF+++ + E + A+ MNG+
Sbjct: 1047 LDDSIGDEKLKELFSPFGTI-TSCKVMRDPN-GLSRGSGFVAFSTPEEASRALLEMNGKM 1220
Query: 179 LCNRQITVSYAYKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANG 238
+ ++ + V+ A +K+ + R ++ P + R ++ G P + +
Sbjct: 1221 VVSKPLYVTLAQRKEDRRARLQAQFAQMRPVGMPPSVGPRV-PMYPPGGPGIGQQLFYSQ 1397
Query: 239 TIPAPVPPRP-----------FANGVAPPPIHVIQPPPPQAPAFQPMQMP---PPGQQVW 284
PA +P +P G AP P + P Q Q P PG
Sbjct: 1398 GPPAIIPSQPGFGYQQQLMPGMRPGAAPVPNFFV----PMVQQGQQGQRPGGRRPG--AV 1559
Query: 285 HQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPP 319
Q QQ PMM Q M P + +R P +P P P
Sbjct: 1560 QQSQQPVPMMPQQMLPRGRVYRYPPGRGIPDVPIP 1664
Score = 94.4 bits (233), Expect = 8e-20
Identities = 107/379 (28%), Positives = 162/379 (42%), Gaps = 20/379 (5%)
Frame = +3
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S ++ ++ NLD I + L + F G +++ V D + Q +GYGFV+F +EE
Sbjct: 423 SIRKSGQGNIFIKNLDRAIDHKALHDTFSTFGNILSCKVATDS-SGQSKGYGFVQFDNEE 599
Query: 78 DADYAIKVLNMIKLYGKPIRVNK--ASQDKKSLDVGA---NLFIGNLDPDVDEKLLYDTF 132
A AI+ LN + L K + V Q+++S A N+F+ NL + L +TF
Sbjct: 600 SAQKAIEKLNGMLLNDKQVYVGPFLRKQERESAADKAKFNNVFVKNLSESTTDDELKNTF 779
Query: 133 SAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
FG I T+ +MRD D G S+ FGF+++++ + + A+EA+NG+ +++ V A KK
Sbjct: 780 GEFGTI-TSAVVMRDGD-GKSKCFGFVNFENADDAARAVEALNGKKFDDKEWYVGKAQKK 953
Query: 193 DTKGERHGTPAERVLAASNPTAQKSRPHTLFAS------GPPSLPNAPQANGTIPAPVPP 246
ER +R + A K + L+ G L GTI +
Sbjct: 954 S---ERENELKQRFEQSMKEAADKYQGANLYVKNLDDSIGDEKLKELFSPFGTITSCKVM 1124
Query: 247 RPFANGVAPPPIHVIQPPPPQAPAFQ-----PMQMPPPGQQVWHQQQQGQPMMQQGMPPP 301
R NG++ V P +A M + P Q+++ + Q
Sbjct: 1125RD-PNGLSRGSGFVAFSTPEEASRALLEMNGKMVVSKPLYVTLAQRKEDRRARLQA---- 1289
Query: 302 MQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMA-GQQQVWRPPPP---PQQQGGHPMG 357
QF + MPP GP P+ PP GQQ + PP P Q G G
Sbjct: 1290--QFAQMRPVGMPPSV------GPRVPMYPPGGPGIGQQLFYSQGPPAIIPSQPG---FG 1436
Query: 358 YPQNSMPPPPPPNHHGMRP 376
Y Q MP GMRP
Sbjct: 1437YQQQLMP--------GMRP 1469
>TC209039 similar to UP|Q9M549 (Q9M549) Poly(A)-binding protein, partial
(25%)
Length = 800
Score = 115 bits (288), Expect = 3e-26
Identities = 66/160 (41%), Positives = 100/160 (62%), Gaps = 1/160 (0%)
Frame = +1
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YVG+L +++ L++LF Q VV+V + +D T Q GYG+V F + DA AI VLN
Sbjct: 316 YVGDLHHDVNDPQLYDLFNQVAQVVSVRICRDVATQQSLGYGYVNFSNAHDAAKAIDVLN 495
Query: 88 MIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMR 146
L GK IR+ + +D + G AN+FI NLD +D K LYDTFSAFG I+ + K+
Sbjct: 496 FTPLNGKIIRIMYSIRDPSARKSGAANVFIKNLDKAIDHKALYDTFSAFGNIL-SCKVAT 672
Query: 147 DPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITV 186
D +G S+G GF+ ++S E++ +AI+ +NG + ++ + V
Sbjct: 673 DA-SGQSKGHGFVQFESEESAQNAIDKLNGMLINDKXVFV 789
>TC214650 similar to UP|Q9LJH8 (Q9LJH8) RNA binding protein nucleolysin;
oligouridylate binding protein (AT3g14100/MAG2_5),
partial (93%)
Length = 1684
Score = 110 bits (275), Expect = 1e-24
Identities = 64/199 (32%), Positives = 111/199 (55%), Gaps = 3/199 (1%)
Frame = +2
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
+ YVGN+ PQ+++ LL ELF AG + + + + YGFV++ A +AI
Sbjct: 347 SVYVGNIHPQVTDSLLQELFSTAGALEGCKL----IRKEKSSYGFVDYFDRSSAAFAIVT 514
Query: 86 LNMIKLYGKPIRVNKASQDKKSLDVGA--NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
LN ++G+PI+VN A + D N+F+G+L P+V + LY FS + ++ +
Sbjct: 515 LNGRNIFGQPIKVNWAYASSQREDTSGHFNIFVGDLSPEVTDATLYACFSVYPSC-SDAR 691
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYK-KDTKGERHGTP 202
+M D TG SRGFGF+S+ + + + SAI + G++L +RQI ++A K E+ +
Sbjct: 692 VMWDQKTGRSRGFGFVSFRNQQDAQSAINDLTGKWLGSRQIRCNWATKGASASDEKQTSD 871
Query: 203 AERVLAASNPTAQKSRPHT 221
+ V+ +N +++ + T
Sbjct: 872 SRSVVELTNGSSEDGQETT 928
Score = 63.2 bits (152), Expect = 2e-10
Identities = 55/233 (23%), Positives = 100/233 (42%), Gaps = 46/233 (19%)
Frame = +2
Query: 10 GANLLGQ-------HSAERNQDATA----YVGNLDPQISEELLWELFVQAGPVVNVYVPK 58
G N+ GQ +++ + +D + +VG+L P++++ L+ F + V
Sbjct: 521 GRNIFGQPIKVNWAYASSQREDTSGHFNIFVGDLSPEVTDATLYACFSVYPSCSDARVMW 700
Query: 59 DRVTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVN------KASQDKKSLDVGA 112
D+ T + +G+GFV FR+++DA AI L L + IR N AS +K++ D +
Sbjct: 701 DQKTGRSRGFGFVSFRNQQDAQSAINDLTGKWLGSRQIRCNWATKGASASDEKQTSDSRS 880
Query: 113 ----------------------------NLFIGNLDPDVDEKLLYDTFSAFGV-IVTNPK 143
+++GNL P+V L+ F + + + +
Sbjct: 881 VVELTNGSSEDGQETTNDDTPEKNPQYTTVYVGNLAPEVTSVDLHQHFHSLNAGTIEDVR 1060
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKG 196
+ RD +GFGF+ Y + + AI+ N + L + I S+ K G
Sbjct: 1061VQRD------KGFGFVRYSTHAEAALAIQMGNARILFGKPIKCSWGSKPTPPG 1201
Score = 48.5 bits (114), Expect = 5e-06
Identities = 32/97 (32%), Positives = 53/97 (53%), Gaps = 3/97 (3%)
Frame = +2
Query: 10 GANLLGQHSAERN-QDATAYVGNLDPQISEELLWELF--VQAGPVVNVYVPKDRVTNQHQ 66
G + E+N Q T YVGNL P+++ L + F + AG + +V V +D+
Sbjct: 914 GQETTNDDTPEKNPQYTTVYVGNLAPEVTSVDLHQHFHSLNAGTIEDVRVQRDK------ 1075
Query: 67 GYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQ 103
G+GFV + + +A AI++ N L+GKPI+ + S+
Sbjct: 1076 GFGFVRYSTHAEAALAIQMGNARILFGKPIKCSWGSK 1186
>TC204822 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (67%)
Length = 1232
Score = 101 bits (252), Expect = 5e-22
Identities = 62/186 (33%), Positives = 101/186 (53%), Gaps = 9/186 (4%)
Frame = +3
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
E ++A +VGNL + + L LF QAG V V +R T+Q +G+GFV + E+A
Sbjct: 315 EPPEEAKLFVGNLPYDVDSQKLAMLFEQAGTVEIAEVIYNRETDQSRGFGFVTMSTVEEA 494
Query: 80 DYAIKVLNMIKLYGKPIRVNKAS---------QDKKSLDVGANLFIGNLDPDVDEKLLYD 130
+ A++ N + G+ + VNKAS ++S + ++++GNL DVD L
Sbjct: 495 ESAVEKFNRYDIDGRLLTVNKASPRGTRPERPPPRRSFESSLSIYVGNLPWDVDNTRLKQ 674
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
FS G +V N +++ D ++G SRGFGF++ + A+ A++G+ L R I VS A
Sbjct: 675 IFSKHGNVV-NARVVYDRESGRSRGFGFVTMSDETEMNDAVAALDGESLDGRAIKVSVAE 851
Query: 191 KKDTKG 196
+ +G
Sbjct: 852 DRPRRG 869
Score = 55.8 bits (133), Expect = 3e-08
Identities = 31/87 (35%), Positives = 50/87 (56%)
Frame = +3
Query: 112 ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAI 171
A LF+GNL DVD + L F G + +++ + +T SRGFGF++ + E ++SA+
Sbjct: 330 AKLFVGNLPYDVDSQKLAMLFEQAGTVEI-AEVIYNRETDQSRGFGFVTMSTVEEAESAV 506
Query: 172 EAMNGQYLCNRQITVSYAYKKDTKGER 198
E N + R +TV+ A + T+ ER
Sbjct: 507 EKFNRYDIDGRLLTVNKASPRGTRPER 587
>TC227154 similar to UP|Q9LJH8 (Q9LJH8) RNA binding protein nucleolysin;
oligouridylate binding protein (AT3g14100/MAG2_5),
partial (92%)
Length = 1698
Score = 100 bits (250), Expect = 8e-22
Identities = 60/175 (34%), Positives = 95/175 (54%), Gaps = 9/175 (5%)
Frame = +1
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
+ YVGN+ Q++E LL E+F GPV + + YGF+ + A AI
Sbjct: 310 SVYVGNIHTQVTEPLLQEVFSGTGPVEGCKL----IRKDKSSYGFIHYFDRRSAALAILS 477
Query: 86 LNMIKLYGKPIRVNKASQDKKSLDVGA---------NLFIGNLDPDVDEKLLYDTFSAFG 136
LN L+G+PI+VN A + D A N+F+G+L P+V + L+ FS +
Sbjct: 478 LNGRHLFGQPIKVNWAYASGQREDTSAFFF*FAGHYNIFVGDLSPEVTDATLFACFSVYP 657
Query: 137 VIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYK 191
++ ++M D TG SRGFGF+S+ + + + SAI + G++L +RQI ++A K
Sbjct: 658 TC-SDARVMWDQKTGRSRGFGFVSFRNQQDAQSAINDLTGKWLGSRQIRCNWATK 819
Score = 67.8 bits (164), Expect = 8e-12
Identities = 60/220 (27%), Positives = 96/220 (43%), Gaps = 35/220 (15%)
Frame = +1
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
+VG+L P++++ L+ F + V D+ T + +G+GFV FR+++DA AI L
Sbjct: 592 FVGDLSPEVTDATLFACFSVYPTCSDARVMWDQKTGRSRGFGFVSFRNQQDAQSAINDLT 771
Query: 88 MIKLYGKPIRVN-----------KASQDKKSL----------------DVGAN------L 114
L + IR N K + D KS+ D N +
Sbjct: 772 GKWLGSRQIRCNWATKGAGGTEEKQNSDAKSVVELTYGSSDGKETSNSDAPENNPQYTTV 951
Query: 115 FIGNLDPDVDEKLLYDTFSAFGV-IVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEA 173
++GNL P+ + L+ F + G ++ ++ RD +GFGF+ Y + + AI+
Sbjct: 952 YVGNLAPEATQLDLHHHFHSLGAGVIEEVRVQRD------KGFGFVRYSTHAEAALAIQM 1113
Query: 174 MNGQ-YLCNRQITVSYAYKKDTKGERHGTPAERVLAASNP 212
N Q LC +QI S+ K T P AAS P
Sbjct: 1114GNAQSLLCGKQIKCSWG-SKPTPAGTASNPLPPPAAASLP 1230
>TC205913 weakly similar to UP|Q9LEB4 (Q9LEB4) RNA Binding Protein 45,
partial (55%)
Length = 886
Score = 100 bits (249), Expect = 1e-21
Identities = 58/176 (32%), Positives = 95/176 (53%), Gaps = 2/176 (1%)
Frame = +1
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
T ++G+L + E L + F G VV++ + ++++T Q +GYGFVEF S A+ ++
Sbjct: 67 TLWIGDLQYWVDESYLSQCFAHNGEVVSIKIIRNKLTGQPEGYGFVEFVSHASAEAFLRT 246
Query: 86 LNMIKLYG--KPIRVNKASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
N ++ G + R+N AS D ++F+G+L PDV + LL +TF A V K
Sbjct: 247 YNGAQMPGTEQTFRLNWASFGDSGPD--HSIFVGDLAPDVTDFLLQETFRAHYPSVKGAK 420
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERH 199
++ DP TG S+G+GF+ + + A+ MNG Y R + +S A K +H
Sbjct: 421 VVTDPATGRSKGYGFVKFADEAQRNRAMTEMNGVYCSTRPMRISAATPKKNASFQH 588
Score = 37.7 bits (86), Expect = 0.009
Identities = 21/61 (34%), Positives = 35/61 (56%)
Frame = +1
Query: 24 DATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAI 83
+ T +GNLD ++EE L + F+Q G +V V + + GYG+V+F + A+ AI
Sbjct: 673 NTTVCIGNLDLNVTEEELKQTFMQFGDIVLVKIYAGK------GYGYVQFGTRVSAEDAI 834
Query: 84 K 84
+
Sbjct: 835 Q 837
>TC225576 similar to UP|Q9AT32 (Q9AT32) Poly(A)-binding protein, partial
(93%)
Length = 2491
Score = 97.8 bits (242), Expect = 7e-21
Identities = 53/135 (39%), Positives = 86/135 (63%), Gaps = 1/135 (0%)
Frame = +3
Query: 62 TNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLD 120
T + GYG+V F + +DA A+ VLN L +PIR+ + +D G AN+FI NLD
Sbjct: 471 TRRSLGYGYVNFSNPQDAARALDVLNFTPLNNRPIRIMYSHRDPSLRKSGTANIFIKNLD 650
Query: 121 PDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLC 180
+D K L+DTFS+FG+I++ KI D +G S+G+GF+ +D+ EA+ +AI+ +NG +
Sbjct: 651 KAIDHKALHDTFSSFGLILSC-KIATDA-SGLSKGYGFVQFDNEEAAQNAIDKLNGMLIN 824
Query: 181 NRQITVSYAYKKDTK 195
++Q+ V + +K +
Sbjct: 825 DKQVYVGHFLRKQDR 869
Score = 77.4 bits (189), Expect = 1e-14
Identities = 78/273 (28%), Positives = 116/273 (41%), Gaps = 16/273 (5%)
Frame = +1
Query: 100 KASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFI 159
K S DK G NL++ NLD + ++ L + F+ +G I T+ K+MRDP TG RG GF+
Sbjct: 1180 KESADKYQ---GVNLYLKNLDDTISDEKLKEMFAEYGTI-TSCKVMRDP-TGIGRGSGFV 1344
Query: 160 SYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERHGTPAERVLAASNPTAQKSRP 219
++ + E + A+ MNG+ + + V+ A +K+ + R + RP
Sbjct: 1345 AFSTPEEASRALGEMNGKMFAGKPLYVALAQRKEERRAR-----------LQAQFSQMRP 1491
Query: 220 HTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQA---------PA 270
+ S P +P P AP + F G PP + PPQA P
Sbjct: 1492 VAITPSVAPRMPLYPPG-----APGLGQQFLYGQGPPAM-----MPPQAGFGYQQQLVPG 1641
Query: 271 FQPMQMPPPG--QQVWHQQQQGQ-PMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPP- 326
+P P P + Q QQGQ P+ ++G P Q +P MQ P + + PP
Sbjct: 1642 MRPGGGPMPSFFVPMVQQGQQGQRPVGRRGTGPVQQPQQPMPMMQQQMLPRGRVYRYPPG 1821
Query: 327 ---RPLPPPSVMAGQQQVWRPPPPPQQQGGHPM 356
+ +P G V P GG PM
Sbjct: 1822 RNMQDVPLQVAAGGMLSV------PYDMGGLPM 1902
Score = 73.9 bits (180), Expect = 1e-13
Identities = 51/171 (29%), Positives = 91/171 (52%), Gaps = 5/171 (2%)
Frame = +3
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S ++ A ++ NLD I + L + F G +++ + D + +GYGFV+F +EE
Sbjct: 603 SLRKSGTANIFIKNLDKAIDHKALHDTFSSFGLILSCKIATD-ASGLSKGYGFVQFDNEE 779
Query: 78 DADYAIKVLNMIKLYGKPIRVNK--ASQDKK---SLDVGANLFIGNLDPDVDEKLLYDTF 132
A AI LN + + K + V QD++ S N+++ NL ++ L F
Sbjct: 780 AAQNAIDKLNGMLINDKQVYVGHFLRKQDRENALSKTKFNNVYVKNLSESTTDEELMKFF 959
Query: 133 SAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQ 183
+G I T+ IMRD D G SR FGF+++++ + + A+E +NG+ + +++
Sbjct: 960 GEYGTI-TSAVIMRDAD-GKSRCFGFVNFENPDDAAKAVEGLNGKKVDDKE 1106
Score = 51.6 bits (122), Expect = 6e-07
Identities = 31/91 (34%), Positives = 50/91 (54%)
Frame = +1
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
+ SA++ Q Y+ NLD IS+E L E+F + G + + V +D T +G GFV F +
Sbjct: 1180 KESADKYQGVNLYLKNLDDTISDEKLKEMFAEYGTITSCKVMRD-PTGIGRGSGFVAFST 1356
Query: 76 EEDADYAIKVLNMIKLYGKPIRVNKASQDKK 106
E+A A+ +N GKP+ V A + ++
Sbjct: 1357 PEEASRALGEMNGKMFAGKPLYVALAQRKEE 1449
>TC225640 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (66%)
Length = 1290
Score = 97.1 bits (240), Expect = 1e-20
Identities = 62/187 (33%), Positives = 102/187 (54%), Gaps = 10/187 (5%)
Frame = +3
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
AE+ ++ +VGNL I E L LF QAG V V +R T++ +G+GFV + E+
Sbjct: 423 AEKAEEDKIFVGNLPFDIDSENLASLFGQAGTVEVAEVIYNRATDRSRGFGFVTMSTLEE 602
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS----------QDKKSLDVGANLFIGNLDPDVDEKLL 128
A+++ + +L G+ + VNKA+ + +S G +++GNL +VD+ L
Sbjct: 603 LKKAVEMFSGYELNGRVLTVNKAAPKGAQPERPPRPPRSFSSGLRVYVGNLPWEVDDARL 782
Query: 129 YDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSY 188
FS G V + +++ D +TG SRGFGF++ S + AI A++GQ L R I V+
Sbjct: 783 EQIFSEHGK-VEDARVVYDRETGRSRGFGFVTMSSETDMNDAIAALDGQSLDGRAIRVNV 959
Query: 189 AYKKDTK 195
A + ++
Sbjct: 960 AQDRPSR 980
Score = 52.0 bits (123), Expect = 4e-07
Identities = 33/103 (32%), Positives = 51/103 (49%)
Frame = +3
Query: 100 KASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFI 159
KA +DK +F+GNL D+D + L F G + +++ + T SRGFGF+
Sbjct: 429 KAEEDK--------IFVGNLPFDIDSENLASLFGQAGTVEV-AEVIYNRATDRSRGFGFV 581
Query: 160 SYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERHGTP 202
+ + E A+E +G L R +TV+ A K + ER P
Sbjct: 582 TMSTLEELKKAVEMFSGYELNGRVLTVNKAAPKGAQPERPPRP 710
>TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding protein 2
{Arabidopsis thaliana;} , partial (59%)
Length = 1223
Score = 94.0 bits (232), Expect = 1e-19
Identities = 61/184 (33%), Positives = 97/184 (52%), Gaps = 6/184 (3%)
Frame = +3
Query: 15 GQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFR 74
G + E ++ +VGNL E L LF QAG V V +R T++ +G+GFV
Sbjct: 396 GGFAEEEAEEVKIFVGNLPFDFDSEKLASLFEQAGTVEVAEVIYNRATDRSRGFGFVTMS 575
Query: 75 SEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLD------VGANLFIGNLDPDVDEKLL 128
+ E+ + A+K+ + +L G+ + VNKA+ + +++GNL DVD L
Sbjct: 576 TIEELEKAVKMFSGYELNGRVLTVNKAAPKGAQPERPPRPPQSFRVYVGNLPWDVDNSRL 755
Query: 129 YDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSY 188
FS G V + +++ D +TG SRGFGF++ S + AI A++GQ L R I V+
Sbjct: 756 EQIFSEHGK-VEDARVVYDRETGRSRGFGFVTMSSETDMNDAIAALDGQSLDGRAIRVNV 932
Query: 189 AYKK 192
A ++
Sbjct: 933 AAQR 944
Score = 55.5 bits (132), Expect = 4e-08
Identities = 33/84 (39%), Positives = 45/84 (53%)
Frame = +3
Query: 23 QDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYA 82
Q YVGNL + L ++F + G V + V DR T + +G+GFV SE D + A
Sbjct: 699 QSFRVYVGNLPWDVDNSRLEQIFSEHGKVEDARVVYDRETGRSRGFGFVTMSSETDMNDA 878
Query: 83 IKVLNMIKLYGKPIRVNKASQDKK 106
I L+ L G+ IRVN A+Q K
Sbjct: 879 IAALDGQSLDGRAIRVNVAAQRPK 950
Score = 50.1 bits (118), Expect = 2e-06
Identities = 34/114 (29%), Positives = 56/114 (48%), Gaps = 3/114 (2%)
Frame = +3
Query: 114 LFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEA 173
+F+GNL D D + L F G + +++ + T SRGFGF++ + E + A++
Sbjct: 432 IFVGNLPFDFDSEKLASLFEQAGTVEV-AEVIYNRATDRSRGFGFVTMSTIEELEKAVKM 608
Query: 174 MNGQYLCNRQITVSYAYKKDTKGERHGTPAE--RVLAASNP-TAQKSRPHTLFA 224
+G L R +TV+ A K + ER P + RV + P SR +F+
Sbjct: 609 FSGYELNGRVLTVNKAAPKGAQPERPPRPPQSFRVYVGNLPWDVDNSRLEQIFS 770
>TC204823 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (66%)
Length = 1596
Score = 91.7 bits (226), Expect = 5e-19
Identities = 58/175 (33%), Positives = 92/175 (52%), Gaps = 9/175 (5%)
Frame = +2
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
AE ++A +VGNL + + L LF QAG V V +R T+Q +G+GFV + E+
Sbjct: 398 AEPPEEAKLFVGNLPYDVDSQKLAMLFEQAGTVEIAEVIYNRETDQSRGFGFVTMSTVEE 577
Query: 79 ADYAIKVLNMIKLYGKPIRVNKAS---------QDKKSLDVGANLFIGNLDPDVDEKLLY 129
A+ A++ + G+ + VNKAS + S + ++++GNL DVD L
Sbjct: 578 AENAVEKFSRYDFDGRLLTVNKASPRGTRPERPPPRHSFEPSLSIYVGNLPWDVDNTRLE 757
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQI 184
FS G +V N +++ D +T SRGFGF++ A+ A++GQ + R I
Sbjct: 758 QIFSEHGNVV-NARVVYDRETRRSRGFGFVTMSDETEMKDAVAALDGQSMDERPI 919
Score = 52.0 bits (123), Expect = 4e-07
Identities = 29/87 (33%), Positives = 49/87 (55%)
Frame = +2
Query: 112 ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAI 171
A LF+GNL DVD + L F G + +++ + +T SRGFGF++ + E +++A+
Sbjct: 416 AKLFVGNLPYDVDSQKLAMLFEQAGTVEI-AEVIYNRETDQSRGFGFVTMSTVEEAENAV 592
Query: 172 EAMNGQYLCNRQITVSYAYKKDTKGER 198
E + R +TV+ A + T+ ER
Sbjct: 593 EKFSRYDFDGRLLTVNKASPRGTRPER 673
Score = 45.8 bits (107), Expect = 3e-05
Identities = 28/82 (34%), Positives = 43/82 (52%)
Frame = +2
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
+HS E + + YVGNL + L ++F + G VVN V DR T + +G+GFV
Sbjct: 683 RHSFEPS--LSIYVGNLPWDVDNTRLEQIFSEHGNVVNARVVYDRETRRSRGFGFVTMSD 856
Query: 76 EEDADYAIKVLNMIKLYGKPIR 97
E + A+ L+ + +PIR
Sbjct: 857 ETEMKDAVAALDGQSMDERPIR 922
>TC214546 similar to PIR|T01932|T01932 RNA binding protein homolog - common
tobacco (fragment) {Nicotiana tabacum;} , partial (68%)
Length = 1756
Score = 91.7 bits (226), Expect = 5e-19
Identities = 59/204 (28%), Positives = 106/204 (51%), Gaps = 7/204 (3%)
Frame = +3
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
+A ++ T ++G+L + E L F G VV+ V +++ T Q +GYGFVEF S
Sbjct: 468 AASSDEIRTVWLGDLHHWMDENYLHNCFAHTGEVVSAKVIRNKQTGQSEGYGFVEFYSRA 647
Query: 78 DADYAIKVLN--MIKLYGKPIRVNKAS---QDKKSLDVGANL--FIGNLDPDVDEKLLYD 130
A+ ++ N M+ + R+N A+ +++S D ++L F+G+L DV + +L +
Sbjct: 648 TAEKVLQNYNGTMMPNTDQAFRLNWATFSAGERRSSDATSDLSIFVGDLAIDVTDAMLQE 827
Query: 131 TFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
TF+ + K++ D +TG S+G+GF+ + A+ MNG Y +R + + A
Sbjct: 828 TFAGRYSSIKGAKVVIDSNTGRSKGYGFVRFGDENERTRAMTEMNGVYCSSRPMRIGVAT 1007
Query: 191 KKDTKGERHGTPAERVLAASNPTA 214
K T G + ++ V+ A +A
Sbjct: 1008PKKTYGFQQQYSSQAVVLAGGHSA 1079
Score = 57.4 bits (137), Expect = 1e-08
Identities = 47/224 (20%), Positives = 98/224 (42%), Gaps = 39/224 (17%)
Frame = +3
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFV-QAGPVVNVYVPKDRVTNQHQGYGFVEFR 74
+ S++ D + +VG+L +++ +L E F + + V D T + +GYGFV F
Sbjct: 744 RRSSDATSDLSIFVGDLAIDVTDAMLQETFAGRYSSIKGAKVVIDSNTGRSKGYGFVRFG 923
Query: 75 SEEDADYAIKVLNMIKLYGKPIRVNKASQDKK----------------------SLDVGA 112
E + A+ +N + +P+R+ A+ K ++ G+
Sbjct: 924 DENERTRAMTEMNGVYCSSRPMRIGVATPKKTYGFQQQYSSQAVVLAGGHSANGAVAQGS 1103
Query: 113 N---------LFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDS 163
+ +F+G LD D ++ L F FG +V+ ++ P +G GF+ +
Sbjct: 1104HSEGDINNTTIFVGGLDSDTSDEDLRQPFLQFGEVVS----VKIPV---GKGCGFVQFAD 1262
Query: 164 FEASDSAIEAMNGQYLCNRQITVSYA-------YKKDTKGERHG 200
+ ++ AI+ +NG + + + +S+ ++ D+ G +G
Sbjct: 1263RKNAEEAIQGLNGTVIGKQTVRLSWGRSPGNKHWRSDSNGGHYG 1394
Score = 48.9 bits (115), Expect = 4e-06
Identities = 31/90 (34%), Positives = 49/90 (54%)
Frame = +3
Query: 10 GANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYG 69
GA G HS + T +VG LD S+E L + F+Q G VV+V +P + G G
Sbjct: 1083 GAVAQGSHSEGDINNTTIFVGGLDSDTSDEDLRQPFLQFGEVVSVKIPVGK------GCG 1244
Query: 70 FVEFRSEEDADYAIKVLNMIKLYGKPIRVN 99
FV+F ++A+ AI+ LN + + +R++
Sbjct: 1245 FVQFADRKNAEEAIQGLNGTVIGKQTVRLS 1334
Score = 35.4 bits (80), Expect = 0.043
Identities = 26/66 (39%), Positives = 28/66 (42%), Gaps = 6/66 (9%)
Frame = +3
Query: 261 IQPPPPQAPAFQPMQMPPPGQQVWHQQQQ---GQPMMQQGMPPPMQQFRP---PHSMQMP 314
+Q Q A QP Q QQ W Q MMQQ M Q + P PH P
Sbjct: 234 VQQQQQQWAAAQPHQY----QQQWMAMQYPATAMAMMQQQMMMYPQHYMPYAHPHYPPPP 401
Query: 315 PPPPPQ 320
PPPPPQ
Sbjct: 402 PPPPPQ 419
Score = 27.7 bits (60), Expect = 8.9
Identities = 23/88 (26%), Positives = 29/88 (32%)
Frame = +3
Query: 285 HQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRP 344
+QQ Q Q QQ Q++ M M P M + P M + P
Sbjct: 219 NQQGQVQQQQQQWAAAQPHQYQQQW-MAMQYPATAMAMMQQQMMMYPQHYMPYAHPHYPP 395
Query: 345 PPPPQQQGGHPMGYPQNSMPPPPPPNHH 372
PPPP PPP +HH
Sbjct: 396 PPPP---------------PPPQSSHHH 434
>TC227320 similar to UP|Q93YF1 (Q93YF1) Nucleic acid binding protein, partial
(70%)
Length = 1724
Score = 90.5 bits (223), Expect = 1e-18
Identities = 62/214 (28%), Positives = 108/214 (49%), Gaps = 10/214 (4%)
Frame = +1
Query: 23 QDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYA 82
++ T ++G+L + E L F G + ++ V +++ T +GYGFVEF S A+
Sbjct: 418 ENKTIWIGDLHHWMDENYLHRCFASTGEISSIKVIRNKQTGLSEGYGFVEFYSHATAEKV 597
Query: 83 IKVLNMIKLYG--KPIRVNKAS---QDKKSLDV-GANLFIGNLDPDVDEKLLYDTFSAFG 136
++ I + +P R+N A+ DK S +V ++F+G+L DV + LL++TF++
Sbjct: 598 LQNYAGILMPNAEQPFRLNWATFSTGDKGSDNVPDLSIFVGDLAADVTDSLLHETFASVY 777
Query: 137 VIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKG 196
V K++ D +TG S+G+GF+ + A+ MNG Y +R + + A + + G
Sbjct: 778 PSVKAAKVVFDANTGRSKGYGFVRFGDDNERTQAMTQMNGVYCSSRPMRIGAATPRKSSG 957
Query: 197 ERHGTPAERVLAASNPTAQKSRPH----TLFASG 226
+ G SN TA +S T+F G
Sbjct: 958 HQQG-------GLSNGTANQSEADSTNTTIFVGG 1038
Score = 68.6 bits (166), Expect = 5e-12
Identities = 47/192 (24%), Positives = 89/192 (45%), Gaps = 18/192 (9%)
Frame = +1
Query: 15 GQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVY-VPKDRVTNQHQGYGFVEF 73
G ++ D + +VG+L +++ LL E F P V V D T + +GYGFV F
Sbjct: 673 GDKGSDNVPDLSIFVGDLAADVTDSLLHETFASVYPSVKAAKVVFDANTGRSKGYGFVRF 852
Query: 74 RSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSL-----------------DVGANLFI 116
+ + A+ +N + +P+R+ A+ K S +F+
Sbjct: 853 GDDNERTQAMTQMNGVYCSSRPMRIGAATPRKSSGHQQGGLSNGTANQSEADSTNTTIFV 1032
Query: 117 GNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNG 176
G LDP+V ++ L FS +G IV+ ++ P +G GF+ + + ++ A++ +NG
Sbjct: 1033GGLDPNVSDEDLRQPFSQYGEIVS----VKIP---VGKGCGFVQFANRNNAEEALQKLNG 1191
Query: 177 QYLCNRQITVSY 188
+ + + +S+
Sbjct: 1192TTIGKQTVRLSW 1227
Score = 47.4 bits (111), Expect = 1e-05
Identities = 27/89 (30%), Positives = 49/89 (54%)
Frame = +1
Query: 11 ANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGF 70
+N S + + T +VG LDP +S+E L + F Q G +V+V +P + G GF
Sbjct: 976 SNGTANQSEADSTNTTIFVGGLDPNVSDEDLRQPFSQYGEIVSVKIPVGK------GCGF 1137
Query: 71 VEFRSEEDADYAIKVLNMIKLYGKPIRVN 99
V+F + +A+ A++ LN + + +R++
Sbjct: 1138 VQFANRNNAEEALQKLNGTTIGKQTVRLS 1224
Score = 31.2 bits (69), Expect = 0.81
Identities = 20/54 (37%), Positives = 23/54 (42%), Gaps = 3/54 (5%)
Frame = +1
Query: 322 MQGPPRPLPPPSVMAGQQQVWRPPPPPQQQG---GHPMGYPQNSMPPPPPPNHH 372
+ PP P P+V QQ W P P H M PQ+ PPP P HH
Sbjct: 214 ISAPPPPRQSPAVARPQQ--WVPMQYPAAAAMVMPHHMLPPQHYAPPPYVPYHH 369
>TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, complete
Length = 1937
Score = 87.8 bits (216), Expect = 7e-18
Identities = 62/190 (32%), Positives = 92/190 (47%), Gaps = 19/190 (10%)
Frame = -1
Query: 22 NQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADY 81
++D +VGNL + L ELF AG V V V D+ T + +G+GFV S E+A+
Sbjct: 890 SRDLKLFVGNLPFSVDSARLAELFESAGNVEVVEVIYDKTTGRSRGFGFVTMSSVEEAEA 711
Query: 82 AIKVLNMIKLYGKPIRVNK-------------------ASQDKKSLDVGANLFIGNLDPD 122
A K N +L G+ +RVN S+ D + +GNL
Sbjct: 710 AAKQFNGYELDGRSLRVNSGPPPARNESAPRFRGGSSFGSRGGGPSDSENRVHVGNLAWG 531
Query: 123 VDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNR 182
VD+ L F G V +++ D ++G SRGFGF+++ S + SAI++++G L R
Sbjct: 530 VDDVALESLFREQGKKVLEARVIYDRESGRSRGFGFVTFGSPDEVKSAIQSLDGVDLNGR 351
Query: 183 QITVSYAYKK 192
I VS A K
Sbjct: 350 AIRVSLADSK 321
Score = 65.1 bits (157), Expect = 5e-11
Identities = 39/90 (43%), Positives = 51/90 (56%)
Frame = +2
Query: 107 SLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEA 166
S DV F+G L D+ L FS +G IV KI+ D +TG SRGFGF+++ S ++
Sbjct: 1226 SADVEYRCFVGGLAWATDDHALERAFSQYGEIVET-KIINDRETGRSRGFGFVTFASEQS 1402
Query: 167 SDSAIEAMNGQYLCNRQITVSYAYKKDTKG 196
AI AMNGQ L R ITV+ A + G
Sbjct: 1403 MKDAIGAMNGQNLDGRNITVNEAQSRGGGG 1492
Score = 54.7 bits (130), Expect = 7e-08
Identities = 37/110 (33%), Positives = 46/110 (41%), Gaps = 1/110 (0%)
Frame = -3
Query: 261 IQPP-PPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPP 319
+QPP PP P P +PPP PPP PP ++ PPPPPP
Sbjct: 1695 LQPPSPPSPPRE*PRSLPPP*PPP--------------PPPP*PPP*PPSRLRYPPPPPP 1558
Query: 320 QGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPP 369
++ PP P PP +++ PPPPP PPPPPP
Sbjct: 1557 PPLR-PPYPPPP--------RLYPPPPPP---------------PPPPPP 1480
Score = 50.8 bits (120), Expect = 1e-06
Identities = 29/84 (34%), Positives = 44/84 (51%)
Frame = -1
Query: 104 DKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDS 163
D S LF+GNL VD L + F + G + +++ D TG SRGFGF++ S
Sbjct: 905 DGPSFSRDLKLFVGNLPFSVDSARLAELFESAGNVEV-VEVIYDKTTGRSRGFGFVTMSS 729
Query: 164 FEASDSAIEAMNGQYLCNRQITVS 187
E +++A + NG L R + V+
Sbjct: 728 VEEAEAAAKQFNGYELDGRSLRVN 657
Score = 47.4 bits (111), Expect = 1e-05
Identities = 28/83 (33%), Positives = 41/83 (48%)
Frame = +2
Query: 19 AERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEED 78
A + + +VG L + L F Q G +V + DR T + +G+GFV F SE+
Sbjct: 1223 ASADVEYRCFVGGLAWATDDHALERAFSQYGEIVETKIINDRETGRSRGFGFVTFASEQS 1402
Query: 79 ADYAIKVLNMIKLYGKPIRVNKA 101
AI +N L G+ I VN+A
Sbjct: 1403 MKDAIGAMNGQNLDGRNITVNEA 1471
Score = 46.2 bits (108), Expect = 2e-05
Identities = 31/80 (38%), Positives = 35/80 (43%), Gaps = 1/80 (1%)
Frame = -3
Query: 291 QPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLP-PPSVMAGQQQVWRPPPPPQ 349
+P Q PP P S+ P PPPP PP P P PPS ++ PPPPP
Sbjct: 1707 KP*FLQPPSPPSPPRE*PRSLPPP*PPPPP----PP*PPP*PPS------RLRYPPPPPP 1558
Query: 350 QQGGHPMGYPQNSMPPPPPP 369
P P PPPPPP
Sbjct: 1557 PPLRPPYPPPPRLYPPPPPP 1498
Score = 44.3 bits (103), Expect = 9e-05
Identities = 29/98 (29%), Positives = 50/98 (50%), Gaps = 1/98 (1%)
Frame = -1
Query: 10 GANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGP-VVNVYVPKDRVTNQHQGY 68
G++ + + + +VGNL + + L LF + G V+ V DR + + +G+
Sbjct: 608 GSSFGSRGGGPSDSENRVHVGNLAWGVDDVALESLFREQGKKVLEARVIYDRESGRSRGF 429
Query: 69 GFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKK 106
GFV F S ++ AI+ L+ + L G+ IRV+ A K
Sbjct: 428 GFVTFGSPDEVKSAIQSLDGVDLNGRAIRVSLADSKPK 315
Score = 43.1 bits (100), Expect = 2e-04
Identities = 39/119 (32%), Positives = 46/119 (37%), Gaps = 1/119 (0%)
Frame = -3
Query: 227 PPSLPNAPQA-NGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWH 285
PPS P+ P+ ++P P PP P PPP PPP P+ PPP
Sbjct: 1689 PPSPPSPPRE*PRSLPPP*PPPP------PPP*-----PPP*PPSRLRYPPPPP------ 1561
Query: 286 QQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRP 344
PPP+ RPP PPPP PP P PPP A + RP
Sbjct: 1560 -------------PPPL---RPP----YPPPPRLYPPPPPPPPPPPPRD*ASLTVILRP 1444
Score = 38.5 bits (88), Expect = 0.005
Identities = 24/65 (36%), Positives = 29/65 (43%)
Frame = -3
Query: 313 MPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHH 372
+ PP PP + PR LPPP PPPPP P P + + PPPP
Sbjct: 1695 LQPPSPPSPPRE*PRSLPPP*----------PPPPPPP*---PPP*PPSRLRYPPPPPPP 1555
Query: 373 GMRPP 377
+RPP
Sbjct: 1554 PLRPP 1540
>TC205914 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, partial (9%)
Length = 1513
Score = 87.4 bits (215), Expect = 9e-18
Identities = 54/177 (30%), Positives = 91/177 (50%), Gaps = 6/177 (3%)
Frame = +3
Query: 49 GPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYG--KPIRVNKAS---- 102
G V+++ + ++++T Q +GYGFVEF S A+ ++ N ++ + R+N AS
Sbjct: 39 GEVISIKIIRNKLTGQPEGYGFVEFVSHAAAERVLQTYNGTQMPATDQTFRLNWASFGIG 218
Query: 103 QDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYD 162
+ + ++F+G+L PDV + LL +TF A V K++ DP+T S+G+GF+ +
Sbjct: 219 ERRPDAAPEHSIFVGDLAPDVTDYLLQETFRAHYPSVRGAKVVTDPNTARSKGYGFVKFS 398
Query: 163 SFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERHGTPAERVLAASNPTAQKSRP 219
+ A+ MNG Y R + +S A K T G + PA V P + P
Sbjct: 399 DENERNRAMTEMNGVYCSTRPMRISAATPKKTTG-AYAAPAAPVPKPVYPVPAYTSP 566
Score = 65.1 bits (157), Expect = 5e-11
Identities = 51/208 (24%), Positives = 95/208 (45%), Gaps = 33/208 (15%)
Frame = +3
Query: 14 LGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVN-VYVPKDRVTNQHQGYGFVE 72
+G+ + + + +VG+L P +++ LL E F P V V D T + +GYGFV+
Sbjct: 213 IGERRPDAAPEHSIFVGDLAPDVTDYLLQETFRAHYPSVRGAKVVTDPNTARSKGYGFVK 392
Query: 73 FRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKS------------------------- 107
F E + + A+ +N + +P+R++ A+ K +
Sbjct: 393 FSDENERNRAMTEMNGVYCSTRPMRISAATPKKTTGAYAAPAAPVPKPVYPVPAYTSPVV 572
Query: 108 ------LDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFIS 160
DV +F+GNLD +V E+ L FG IV+ + P +GFGF+
Sbjct: 573 QVQPPDYDVNNTTIFVGNLDLNVSEEELKQNSLQFGEIVS---VKIQP----GKGFGFVQ 731
Query: 161 YDSFEASDSAIEAMNGQYLCNRQITVSY 188
+ + +++ AI+ M G+ + + + +S+
Sbjct: 732 FGTRASAEEAIQKMQGKMIGQQVVRISW 815
Score = 40.8 bits (94), Expect = 0.001
Identities = 26/73 (35%), Positives = 41/73 (55%)
Frame = +3
Query: 24 DATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAI 83
+ T +VGNLD +SEE L + +Q G +V+V + + G+GFV+F + A+ AI
Sbjct: 603 NTTIFVGNLDLNVSEEELKQNSLQFGEIVSVKIQPGK------GFGFVQFGTRASAEEAI 764
Query: 84 KVLNMIKLYGKPI 96
+ K+ GK I
Sbjct: 765 Q-----KMQGKMI 788
Score = 28.5 bits (62), Expect = 5.2
Identities = 23/81 (28%), Positives = 27/81 (32%), Gaps = 11/81 (13%)
Frame = +1
Query: 299 PPPMQQFRPPHSMQMPPPPPPQGMQGPPRPL----PPPSVMAGQQQVWRPPPPPQQQGGH 354
P P RP + + PP P G GPP L P P + PP P
Sbjct: 124 PQPRGFSRPTTAPKCPPLTRPSGSTGPPSALVSAGPMPPLNTPSSSAISPPT*PITFSRK 303
Query: 355 -------PMGYPQNSMPPPPP 368
P P+ S P PP
Sbjct: 304 LFALTTLPFAAPKLSPTPTPP 366
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 87.0 bits (214), Expect = 1e-17
Identities = 61/168 (36%), Positives = 70/168 (41%), Gaps = 2/168 (1%)
Frame = +1
Query: 212 PTAQKSRPHTLFASGPPSLPNAPQANGT--IPAPVPPRPFANGVAPPPIHVIQPPPPQAP 269
P+ P S PP P+ P + T + +P PP P PPP PPPP P
Sbjct: 13 PSPPPPSPPPPVYSPPPPPPSPPPPSPTYCVRSPPPPSP------PPP----SPPPPPPP 162
Query: 270 AFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPL 329
F P PPP Q + P Q PPP PPHS PPP PP + P
Sbjct: 163 VFSP---PPPVQYYY----SSPPPPQHSPPPP-----PPHSPPPPPPSPPPPVYPYLSPP 306
Query: 330 PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPP 377
PPP V + V+ PPPPP P P PPPPPP PP
Sbjct: 307 PPPPVHSPPPPVYSPPPPP------PSPPPCIEPPPPPPPCIEPPPPP 432
Score = 79.7 bits (195), Expect = 2e-15
Identities = 57/175 (32%), Positives = 69/175 (38%), Gaps = 8/175 (4%)
Frame = +1
Query: 212 PTAQKSRPHTLFASGPPSLPNAPQANGTIP---APVPPRPFANGVAPPPIHVIQPPPPQA 268
P+ P S PP P P P +P PP + PPP H PPPP
Sbjct: 70 PSPPPPSPTYCVRSPPPPSPPPPSPPPPPPPVFSPPPPVQYYYSSPPPPQH--SPPPP-P 240
Query: 269 PAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQ-----FRPPHSMQMPPPPPPQGMQ 323
P P P P V+ P PPP+ PP ++ PPPPPP
Sbjct: 241 PHSPPPPPPSPPPPVYPYLSPPPPPPVHSPPPPVYSPPPPPPSPPPCIEPPPPPPPCIEP 420
Query: 324 GPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPPS 378
PP P PPP ++ PPP P HP P + PPPPP ++ PPS
Sbjct: 421 PPPPPSPPPC-----EEHSPPPPSPHPAPYHP---PPSPSPPPPPVQYNSPPPPS 561
Score = 77.4 bits (189), Expect = 1e-14
Identities = 57/178 (32%), Positives = 69/178 (38%), Gaps = 2/178 (1%)
Frame = +1
Query: 202 PAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVI 261
P + +S P Q S P S PP P+ P +P PP P + PPP++
Sbjct: 178 PPVQYYYSSPPPPQHSPPPPPPHSPPPPPPSPPPPVYPYLSPPPPPPVHS--PPPPVYSP 351
Query: 262 QPPPPQAP-AFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMP-PPPPP 319
PPPP P +P PPP + P + PPP P H P PPPPP
Sbjct: 352 PPPPPSPPPCIEPPPPPPPCIEPPPPPPSPPPCEEHSPPPPSPHPAPYHPPPSPSPPPPP 531
Query: 320 QGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPP 377
PP P PPP + PPPP S PPP P + G PP
Sbjct: 532 VQYNSPPPPSPPPPTPVYH---YNSPPPP-------------SFPPPTPV-YEGPLPP 654
Score = 62.4 bits (150), Expect = 3e-10
Identities = 44/138 (31%), Positives = 52/138 (36%)
Frame = +1
Query: 241 PAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPP 300
P+P PP P PPP++ PPPP P PP
Sbjct: 13 PSPPPPSP------PPPVYSPPPPPPSPP--------PPS-------------------- 90
Query: 301 PMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQ 360
P + ++ PPPP P PP P PPP V+ PPPP Q P P
Sbjct: 91 ------PTYCVRSPPPPSPP----PPSPPPPP------PPVFSPPPPVQYYYSSPP--PP 216
Query: 361 NSMPPPPPPNHHGMRPPS 378
PPPPPP+ PPS
Sbjct: 217 QHSPPPPPPHSPPPPPPS 270
Score = 28.9 bits (63), Expect = 4.0
Identities = 19/65 (29%), Positives = 24/65 (36%), Gaps = 5/65 (7%)
Frame = +1
Query: 212 PTAQKSRPHTLFASGPPSLPNAPQA-----NGTIPAPVPPRPFANGVAPPPIHVIQPPPP 266
P+ P + S PP P P + P+ PP P G PP I V PP
Sbjct: 505 PSPSPPPPPVQYNSPPPPSPPPPTPVYHYNSPPPPSFPPPTPVYEGPLPPVIGVSYASPP 684
Query: 267 QAPAF 271
P +
Sbjct: 685 PPPFY 699
>BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvox carteri f.
nagariensis}, partial (25%)
Length = 407
Score = 84.7 bits (208), Expect = 6e-17
Identities = 56/162 (34%), Positives = 61/162 (37%), Gaps = 3/162 (1%)
Frame = -3
Query: 219 PHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAP---AFQPMQ 275
P +L PP P P P P PP P PPP PPPP P + P++
Sbjct: 396 PSSLLPPLPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPPPPPLPPLPXSLSPLK 217
Query: 276 MPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVM 335
PPP P PPP F PP PPPPPP PP P PPP
Sbjct: 216 NPPP---------PPPPPPPPPPPPPPPPFPPPPPPPPPPPPPPPPPLPPPPPPPPP--- 73
Query: 336 AGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPP 377
PPPPP P PPPPPP + PP
Sbjct: 72 --------PPPPP----------PPPPPPPPPPPPRANLSPP 1
Score = 84.3 bits (207), Expect = 8e-17
Identities = 55/155 (35%), Positives = 60/155 (38%)
Frame = -2
Query: 225 SGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVW 284
S PP LP P P PP P + + PPP PPPP P P PPP
Sbjct: 406 SPPPLLPPPSPPPPPPPPPPPPPPPPSPLPPPPPPXPPPPPPPPPPPPPPPSPPPPXXSL 227
Query: 285 HQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRP 344
++ P PPP PP + PPPPPP PP P PPPS P
Sbjct: 226 PPKKPPPPPPPPPPPPPPP---PPPPLPPPPPPPPPPPPPPPPPPPPPS----------P 86
Query: 345 PPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPPSG 379
PPPP PPPPPP PP G
Sbjct: 85 PPPPPPP------------PPPPPPPPPPPPPPPG 17
Score = 80.5 bits (197), Expect = 1e-15
Identities = 54/151 (35%), Positives = 59/151 (38%)
Frame = -1
Query: 227 PPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQ 286
PP P P + + P P PP P PPP+ PPPP P P PPP +
Sbjct: 404 PP--PPPPSSLPSPPPPPPPPP----PPPPPLPPPPPPPPXPPPPPPPPPPPPPPPLSPP 243
Query: 287 QQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPP 346
P + PPP PP PPPPPP PP P PPP PPP
Sbjct: 242 SLXLSPP*KTPPPPP-----PPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPSPPLPPPP 78
Query: 347 PPQQQGGHPMGYPQNSMPPPPPPNHHGMRPP 377
PP P P PPPPPP PP
Sbjct: 77 PP------PPPPPPPPPPPPPPPPPGQTSPP 3
Score = 53.9 bits (128), Expect = 1e-07
Identities = 39/117 (33%), Positives = 42/117 (35%)
Frame = -1
Query: 210 SNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAP 269
S P+ S P PP P P P+P PP P PPP PPPP P
Sbjct: 251 SPPSLXLSPP*KTPPPPPPPPPPPPPPPPPPPSPPPPPP------PPP-----PPPPPPP 105
Query: 270 AFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPP 326
P+ PPP PPP PP PPPPPP G PP
Sbjct: 104 PSPPLPPPPP-------------------PPP----PPPPPPPPPPPPPPPGQTSPP 3
Score = 53.1 bits (126), Expect = 2e-07
Identities = 38/120 (31%), Positives = 39/120 (31%)
Frame = -2
Query: 212 PTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAF 271
P P S PP P P P P PP P PPP PPPP P
Sbjct: 271 PPPPPPSPPPPXXSLPPKKPPPPPPPPPPPPPPPPPPPLPPPPPPPPPPPPPPPPPPPPP 92
Query: 272 QPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPP 331
P PPP PPP PPPPPP P +PLPP
Sbjct: 91 SPPPPPPP-------------------PPP-----------PPPPPPPPPPPPPGKPLPP 2
>TC225739 homologue to UP|Q41124 (Q41124) Chloroplast RNA binding protein
precursor, partial (85%)
Length = 1436
Score = 84.3 bits (207), Expect = 8e-17
Identities = 57/185 (30%), Positives = 93/185 (49%), Gaps = 7/185 (3%)
Frame = +3
Query: 15 GQHSAERNQDATA---YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFV 71
G+ AE++ D++A Y GNL + L L G + V DR T + +G+ FV
Sbjct: 459 GEAVAEQDSDSSATKLYFGNLPYSVDSAKLAGLIQDYGSAELIEVLYDRDTGKSRGFAFV 638
Query: 72 EFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSL----DVGANLFIGNLDPDVDEKL 127
ED + I+ L+ + G+ +RVN +S+ K + LF+GNL V ++
Sbjct: 639 TMSCIEDCNAVIENLDGKEFLGRTLRVNFSSKPKPKEPLYPETEHKLFVGNLSWSVTNEI 818
Query: 128 LYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS 187
L F +G +V +++ D +TG SRG+GF+ Y + ++A+ A+N L R + VS
Sbjct: 819 LTQAFQEYGTVV-GARVLYDGETGRSRGYGFVCYSTKAEMEAALAALNDVELEGRAMRVS 995
Query: 188 YAYKK 192
A K
Sbjct: 996 LAQGK 1010
Score = 45.4 bits (106), Expect = 4e-05
Identities = 29/95 (30%), Positives = 45/95 (46%)
Frame = +3
Query: 101 ASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFIS 160
A QD S L+ GNL VD L +G +++ D DTG SRGF F++
Sbjct: 471 AEQDSDSS--ATKLYFGNLPYSVDSAKLAGLIQDYGSAELI-EVLYDRDTGKSRGFAFVT 641
Query: 161 YDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTK 195
E ++ IE ++G+ R + V+++ K K
Sbjct: 642 MSCIEDCNAVIENLDGKEFLGRTLRVNFSSKPKPK 746
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,691,535
Number of Sequences: 63676
Number of extensions: 812362
Number of successful extensions: 39223
Number of sequences better than 10.0: 1963
Number of HSP's better than 10.0 without gapping: 8206
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 18109
length of query: 379
length of database: 12,639,632
effective HSP length: 99
effective length of query: 280
effective length of database: 6,335,708
effective search space: 1773998240
effective search space used: 1773998240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC139882.5


