

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139709.6 + phase: 0
(138 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
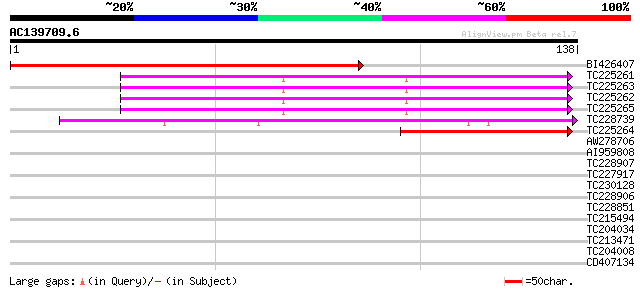
Score E
Sequences producing significant alignments: (bits) Value
BI426407 homologue to SP|Q9BBQ3|RR11_ Chloroplast 30S ribosomal ... 149 4e-37
TC225261 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal prote... 63 4e-11
TC225263 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal prote... 63 4e-11
TC225262 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal prote... 63 4e-11
TC225265 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal prote... 63 4e-11
TC228739 weakly similar to UP|P93408 (P93408) Mitochondrial ribo... 49 6e-07
TC225264 homologue to UP|Q6PDV6 (Q6PDV6) Rps14 protein, partial ... 43 4e-05
AW278706 similar to SP|O22584|RS14 40S ribosomal protein S14. [Y... 38 0.002
AI959808 28 1.4
TC228907 similar to GB|AAO39954.1|28372944|BT003726 At5g63130 {A... 28 1.8
TC227917 UP|Q9M4R7 (Q9M4R7) Beta-ketoacyl-acyl carrier protein s... 28 1.8
TC230128 27 2.4
TC228906 similar to UP|Q6ID88 (Q6ID88) At3g26510, partial (48%) 27 4.1
TC228851 similar to GB|AAT06465.1|46931322|BT012646 At1g44542 {A... 26 6.9
TC215494 similar to UP|Q9SBR6 (Q9SBR6) Enod93 protein, partial (... 26 6.9
TC204034 similar to UP|Q9XQB1 (Q9XQB1) LHCII type III chlorophyl... 26 6.9
TC213471 25 9.0
TC204008 homologue to UP|Q9XQB1 (Q9XQB1) LHCII type III chloroph... 25 9.0
CD407134 similar to GP|4689380|gb|A LHCII type III chlorophyll a... 25 9.0
>BI426407 homologue to SP|Q9BBQ3|RR11_ Chloroplast 30S ribosomal protein S11.
{Lotus japonicus}, partial (62%)
Length = 406
Score = 149 bits (376), Expect = 4e-37
Identities = 76/86 (88%), Positives = 78/86 (90%)
Frame = +2
Query: 1 MAKSIPKIGSRKNGRIGSRKHPRKIPKGVIYVQASFNNTIVTVTDVRGRVISWSSAGSCG 60
MAK I K GSRKN RIGSRKH KIPKGVI+VQASFNNTIVTVTDVRGRVISWSSAG+CG
Sbjct: 149 MAKPISKKGSRKNVRIGSRKHTLKIPKGVIHVQASFNNTIVTVTDVRGRVISWSSAGTCG 328
Query: 61 FKGTRRGTPFAAQTAAANAIRTVVDQ 86
FKGTRRGTPFAAQTAA NAIRTV DQ
Sbjct: 329 FKGTRRGTPFAAQTAAGNAIRTVSDQ 406
>TC225261 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein S14,
complete
Length = 849
Score = 63.2 bits (152), Expect = 4e-11
Identities = 43/119 (36%), Positives = 59/119 (49%), Gaps = 9/119 (7%)
Frame = +2
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 230 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVAARCKEL 409
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 410 GITALHIKLRATGGNKTKTPGPGAQSALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 586
>TC225263 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein S14,
complete
Length = 810
Score = 63.2 bits (152), Expect = 4e-11
Identities = 43/119 (36%), Positives = 59/119 (49%), Gaps = 9/119 (7%)
Frame = +1
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 160 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVAARCKEL 339
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 340 GITALHIKLRATGGNKTKTPGPGAQSALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 516
>TC225262 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein S14,
complete
Length = 800
Score = 63.2 bits (152), Expect = 4e-11
Identities = 43/119 (36%), Positives = 59/119 (49%), Gaps = 9/119 (7%)
Frame = +3
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 153 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVAARCKEL 332
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 333 GITALHIKLRATGGNKTKTPGPGAQSALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 509
>TC225265 homologue to UP|RS14_LUPLU (O22584) 40S ribosomal protein S14,
complete
Length = 1024
Score = 63.2 bits (152), Expect = 4e-11
Identities = 43/119 (36%), Positives = 59/119 (49%), Gaps = 9/119 (7%)
Frame = +1
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 253 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVAARCKEL 432
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 433 GITALHIKLRATGGNKTKTPGPGAQSALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 609
>TC228739 weakly similar to UP|P93408 (P93408) Mitochondrial ribosomal
protein S11 (Nuclear encoded), partial (39%)
Length = 1290
Score = 49.3 bits (116), Expect = 6e-07
Identities = 42/139 (30%), Positives = 62/139 (44%), Gaps = 13/139 (9%)
Frame = +3
Query: 13 NGRIGSRKHPRKIPKGVIYVQASF--NNTIVTVTDVRGRVISWSSAGSC-GFKGTRRGTP 69
N R GS H + + +V NNT VTVTD +G + SAG K ++ +
Sbjct: 612 NFRKGSPYHQYQYEQDADFVHIKMLRNNTFVTVTDSKGNIKLSGSAGPLKDLKSGQKLSR 791
Query: 70 FAAQTAAANAIRTVVDQGMQRAVVIIKGPGLGRDAALRAIA-RSGIL---------LRFI 119
+AA+ A R G++ V+ + G R ++ + G + +I
Sbjct: 792 YAAEATAEVVGRKARGLGLKSVVMKVNGFTHFRRKRNAIMSWKEGFTAGSRGDRNPIVYI 971
Query: 120 RDVTPIPHNGCRAPKKRRV 138
D T PHNGCR PKKRR+
Sbjct: 972 EDTTRKPHNGCRLPKKRRI 1028
>TC225264 homologue to UP|Q6PDV6 (Q6PDV6) Rps14 protein, partial (60%)
Length = 603
Score = 43.1 bits (100), Expect = 4e-05
Identities = 22/42 (52%), Positives = 27/42 (63%)
Frame = +1
Query: 96 KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 133 KTPGPGAQSALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 258
>AW278706 similar to SP|O22584|RS14 40S ribosomal protein S14. [Yellow
lupine] {Lupinus luteus}, complete
Length = 448
Score = 37.7 bits (86), Expect = 0.002
Identities = 31/107 (28%), Positives = 49/107 (44%), Gaps = 9/107 (8%)
Frame = +2
Query: 34 ASFNNTIVTVTDVRGRVISWS-SAGSCGFKGTRRGTPFAAQTAAANAIRTVVDQGMQRAV 92
A FN++ + VTD+ GR +AG +P+AA A + + G+
Sbjct: 98 AFFNDSFIHVTDLSGRETHVRITAGMKA*ADRYESSPYAAMLAPHDVAA*CKELGVTALH 277
Query: 93 VIIKG--------PGLGRDAALRAIARSGILLRFIRDVTPIPHNGCR 131
+ ++ PG G +AL A+A S + + I DVTPIP + R
Sbjct: 278 ITLRATGGNKTMTPGAGAQSALPALALS*MKIVRIDDVTPIPSDATR 418
>AI959808
Length = 410
Score = 28.1 bits (61), Expect = 1.4
Identities = 12/34 (35%), Positives = 22/34 (64%)
Frame = +1
Query: 73 QTAAANAIRTVVDQGMQRAVVIIKGPGLGRDAAL 106
+++ A+ + VD+ ++R ++ GPGLGRD L
Sbjct: 25 KSSIASKVLAEVDKWLERFDCLVVGPGLGRDPFL 126
>TC228907 similar to GB|AAO39954.1|28372944|BT003726 At5g63130 {Arabidopsis
thaliana;} , partial (53%)
Length = 537
Score = 27.7 bits (60), Expect = 1.8
Identities = 15/33 (45%), Positives = 22/33 (66%)
Frame = +3
Query: 106 LRAIARSGILLRFIRDVTPIPHNGCRAPKKRRV 138
LRAIA +G LR +R+V P+P R P++ R+
Sbjct: 291 LRAIAEAGGALRRLREVPPLPIT-FRRPRRSRL 386
>TC227917 UP|Q9M4R7 (Q9M4R7) Beta-ketoacyl-acyl carrier protein synthase III,
complete
Length = 1744
Score = 27.7 bits (60), Expect = 1.8
Identities = 11/32 (34%), Positives = 15/32 (46%)
Frame = +3
Query: 36 FNNTIVTVTDVRGRVISWSSAGSCGFKGTRRG 67
FNN +V D R + W+ G+C G G
Sbjct: 693 FNNVLVIGADALSRYVDWTDRGTCILFGDAAG 788
>TC230128
Length = 1104
Score = 27.3 bits (59), Expect = 2.4
Identities = 17/53 (32%), Positives = 25/53 (47%)
Frame = +3
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTRRGTPFAAQTAAANAI 80
G IY+ S+N+ I + RV + + G GFK GT AQ + + I
Sbjct: 24 GEIYIADSYNHKIKKLDPTSKRVSTIAGTGKAGFKD---GTAVKAQLSEPSGI 173
>TC228906 similar to UP|Q6ID88 (Q6ID88) At3g26510, partial (48%)
Length = 927
Score = 26.6 bits (57), Expect = 4.1
Identities = 14/33 (42%), Positives = 21/33 (63%)
Frame = +1
Query: 106 LRAIARSGILLRFIRDVTPIPHNGCRAPKKRRV 138
LRAIA +G LR +R+ P+P R P++ R+
Sbjct: 268 LRAIAETGGALRRLREAPPLPIT-LRRPRRSRL 363
>TC228851 similar to GB|AAT06465.1|46931322|BT012646 At1g44542 {Arabidopsis
thaliana;} , partial (75%)
Length = 959
Score = 25.8 bits (55), Expect = 6.9
Identities = 16/40 (40%), Positives = 20/40 (50%)
Frame = -1
Query: 63 GTRRGTPFAAQTAAANAIRTVVDQGMQRAVVIIKGPGLGR 102
GTR +PFAAQ AA A + D R +V + GR
Sbjct: 122 GTRLSSPFAAQITAAKA-QASTDNNAPRFIVRFRHNQSGR 6
>TC215494 similar to UP|Q9SBR6 (Q9SBR6) Enod93 protein, partial (85%)
Length = 761
Score = 25.8 bits (55), Expect = 6.9
Identities = 10/24 (41%), Positives = 14/24 (57%)
Frame = -2
Query: 51 ISWSSAGSCGFKGTRRGTPFAAQT 74
I+W+S SC + +RG PF T
Sbjct: 169 IAWTSKSSCPLRERQRGRPFCWAT 98
>TC204034 similar to UP|Q9XQB1 (Q9XQB1) LHCII type III chlorophyll a/b
binding protein, partial (47%)
Length = 934
Score = 25.8 bits (55), Expect = 6.9
Identities = 16/43 (37%), Positives = 22/43 (50%), Gaps = 2/43 (4%)
Frame = +2
Query: 47 RGRVISWSSAGSCGFKGTRRGTPFAAQ--TAAANAIRTVVDQG 87
RG + ++ + GT + TPF Q A ANA+R VV G
Sbjct: 425 RGLEVMAATLMAATSSGTLKATPFLGQGKGANANALRDVVSMG 553
>TC213471
Length = 422
Score = 25.4 bits (54), Expect = 9.0
Identities = 12/17 (70%), Positives = 12/17 (70%)
Frame = -2
Query: 63 GTRRGTPFAAQTAAANA 79
GTR PFAAQ AAA A
Sbjct: 109 GTRLSLPFAAQIAAAKA 59
>TC204008 homologue to UP|Q9XQB1 (Q9XQB1) LHCII type III chlorophyll a/b
binding protein, complete
Length = 1220
Score = 25.4 bits (54), Expect = 9.0
Identities = 14/27 (51%), Positives = 16/27 (58%), Gaps = 2/27 (7%)
Frame = +1
Query: 63 GTRRGTPFAAQ--TAAANAIRTVVDQG 87
GT + TPF Q A ANA+R VV G
Sbjct: 175 GTLKATPFLGQGKGANANALRDVVSMG 255
>CD407134 similar to GP|4689380|gb|A LHCII type III chlorophyll a/b binding
protein {Vigna radiata}, partial (21%)
Length = 636
Score = 25.4 bits (54), Expect = 9.0
Identities = 14/27 (51%), Positives = 16/27 (58%), Gaps = 2/27 (7%)
Frame = +2
Query: 63 GTRRGTPFAAQ--TAAANAIRTVVDQG 87
GT + TPF Q A ANA+R VV G
Sbjct: 500 GTLKATPFLGQGKGANANALRDVVSMG 580
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.138 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,507,688
Number of Sequences: 63676
Number of extensions: 59208
Number of successful extensions: 290
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 284
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 284
length of query: 138
length of database: 12,639,632
effective HSP length: 87
effective length of query: 51
effective length of database: 7,099,820
effective search space: 362090820
effective search space used: 362090820
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC139709.6


