

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139709.19 + phase: 0
(353 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
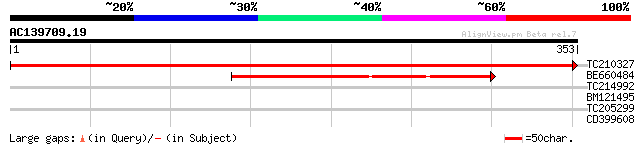
Score E
Sequences producing significant alignments: (bits) Value
TC210327 UP|Q6LDZ6 (Q6LDZ6) Photosystem II thylakoid membrane pr... 714 0.0
BE660484 homologue to GP|11022864|gb photosystem II D2 protein {... 133 1e-31
TC214992 homologue to UP|FD62_SOYBN (P48631) Omega-6 fatty acid ... 30 1.7
BM121495 similar to SP|P48631|FD62 Omega-6 fatty acid desaturase... 30 1.7
TC205299 similar to UP|Q940Z9 (Q940Z9) AT4g02580/T10P11_14, part... 30 2.2
CD399608 28 6.3
>TC210327 UP|Q6LDZ6 (Q6LDZ6) Photosystem II thylakoid membrane protein,
complete
Length = 1114
Score = 714 bits (1843), Expect = 0.0
Identities = 348/353 (98%), Positives = 352/353 (99%)
Frame = +1
Query: 1 MTAILERRDSENLWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI 60
MTAILERR+SE+LW RFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI
Sbjct: 19 MTAILERRESESLWGRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI 198
Query: 61 DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL 120
DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL
Sbjct: 199 DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL 378
Query: 121 LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF 180
LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF
Sbjct: 379 LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF 558
Query: 181 NFMIVFQAEHNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFG 240
NFMIVFQAEHNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFG
Sbjct: 559 NFMIVFQAEHNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFG 738
Query: 241 QEEETYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGF 300
QEEETYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGF
Sbjct: 739 QEEETYNIVAAHGYFGRLIFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGF 918
Query: 301 NFNQSVVDSQGRVINTWADIINRANLGMEVMHERNAHNFPLDLAAVEAPSING 353
NFNQSVVDSQGRVINTWADIINRANLGMEVMHERNAHNFPLDLAA++APSING
Sbjct: 919 NFNQSVVDSQGRVINTWADIINRANLGMEVMHERNAHNFPLDLAAIDAPSING 1077
>BE660484 homologue to GP|11022864|gb photosystem II D2 protein {Acorus
calamus}, partial (66%)
Length = 723
Score = 133 bits (334), Expect = 1e-31
Identities = 64/164 (39%), Positives = 103/164 (62%)
Frame = +3
Query: 139 MRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTFNFMIVFQAEHNILMHPFH 198
+RP+ A+A+S P+A +VFLIYP+GQ + G++ F F++ FQ HN ++PFH
Sbjct: 3 LRPYNAIAFSGPIAVFVSVFLIYPLGQSGWFFAPSFGVAAIFRFILFFQGFHNWTLNPFH 182
Query: 199 MLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFGQEEETYNIVAAHGYFGRL 258
M+GVAGV G +L A+HG+ V ++L E + + + Q EETY++V A+ ++ ++
Sbjct: 183 MMGVAGVLGAALLCAIHGATVENTLF-EDGDGANTFRAFNPTQAEETYSMVTANRFWSQI 359
Query: 259 IFQYASFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGFNF 302
+F+N R LHFF+ PV G+W +ALG+ +A NL ++F
Sbjct: 360 F--GVAFSNKRWLHFFMLFVPVTGLWMSALGVVGLALNLRAYDF 485
>TC214992 homologue to UP|FD62_SOYBN (P48631) Omega-6 fatty acid desaturase,
endoplasmic reticulum isozyme 2 , complete
Length = 1556
Score = 30.0 bits (66), Expect = 1.7
Identities = 11/25 (44%), Positives = 18/25 (72%)
Frame = +2
Query: 177 SGTFNFMIVFQAEHNILMHPFHMLG 201
SGT N + + QAE ++L+ P H++G
Sbjct: 581 SGTLNTLTILQAESSLLLSPSHLVG 655
>BM121495 similar to SP|P48631|FD62 Omega-6 fatty acid desaturase
endoplasmic reticulum isozyme 2 (EC 1.14.19.-).
[Soybean], partial (62%)
Length = 852
Score = 30.0 bits (66), Expect = 1.7
Identities = 11/25 (44%), Positives = 18/25 (72%)
Frame = +2
Query: 177 SGTFNFMIVFQAEHNILMHPFHMLG 201
SGT N + + QAE ++L+ P H++G
Sbjct: 242 SGTLNTLTILQAESSLLLSPSHLVG 316
>TC205299 similar to UP|Q940Z9 (Q940Z9) AT4g02580/T10P11_14, partial (98%)
Length = 1232
Score = 29.6 bits (65), Expect = 2.2
Identities = 13/43 (30%), Positives = 20/43 (46%)
Frame = +1
Query: 277 AWPVVGIWFTALGISTMAFNLNGFNFNQSVVDSQGRVINTWAD 319
AW + W + +STM + N N +D+ RV +W D
Sbjct: 1075 AWHALPTWVCRMDLSTMVWENNHVQCNIHFLDNGQRVFRSWHD 1203
>CD399608
Length = 668
Score = 28.1 bits (61), Expect = 6.3
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +1
Query: 279 PVVGIWFTALGISTMAFNLNGFNFNQSVVDS 309
P +G+W++ L S++ + G F+ S+VDS
Sbjct: 553 PXIGLWYSNLR*SSVLSTILGVKFSGSIVDS 645
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.139 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,876,359
Number of Sequences: 63676
Number of extensions: 241232
Number of successful extensions: 1515
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1508
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1514
length of query: 353
length of database: 12,639,632
effective HSP length: 98
effective length of query: 255
effective length of database: 6,399,384
effective search space: 1631842920
effective search space used: 1631842920
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139709.19


