

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139344.4 - phase: 0 /pseudo
(826 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
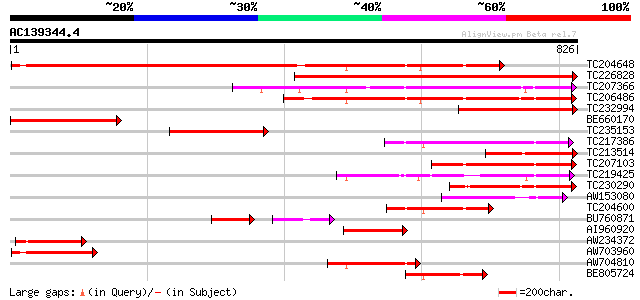
Score E
Sequences producing significant alignments: (bits) Value
TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, com... 836 0.0
TC226828 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 680 0.0
TC207366 similar to UP|Q94BZ3 (Q94BZ3) At2g32810/F24L7.5, partia... 424 e-118
TC206486 similar to UP|O65761 (O65761) Beta-galactosidase (Frag... 366 e-101
TC232994 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 313 2e-85
BE660170 299 4e-81
TC235153 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 281 8e-76
TC217386 similar to GB|AAF70823.1|7939621|AF154422 beta-galactos... 253 2e-67
TC213514 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , par... 233 2e-61
TC207103 homologue to UP|O65761 (O65761) Beta-galactosidase (Fr... 215 7e-56
TC219425 212 5e-55
TC230290 weakly similar to UP|Q93X58 (Q93X58) Beta-galactosidase... 173 3e-43
AW153080 weakly similar to GP|14517399|gb| At2g32810/F24L7.5 {Ar... 154 2e-37
TC204600 homologue to UP|Q9M5J3 (Q9M5J3) Beta galactosidase, par... 149 4e-36
BU760871 homologue to GP|3204134|emb beta-galactosidase {Cicer a... 107 2e-35
AI960920 similar to GP|14970841|emb beta-galactosidase {Fragaria... 146 4e-35
AW234372 homologue to GP|7682680|gb| beta galactosidase {Vigna r... 142 5e-34
AW703960 similar to GP|7682677|gb| beta galactosidase {Vigna rad... 140 2e-33
AW704810 similar to GP|7682677|gb| beta galactosidase {Vigna rad... 125 8e-29
BE805724 120 2e-27
>TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, complete
Length = 2817
Score = 836 bits (2159), Expect = 0.0
Identities = 405/728 (55%), Positives = 505/728 (68%), Gaps = 9/728 (1%)
Frame = +3
Query: 3 GTQIVFVLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQ 62
G ++ + LW GV ++VTYDH+A+V+DGKRR+L+SGSIHYPRSTPQMWPDLIQ
Sbjct: 93 GVVLMMLCLWVCGV------TASVTYDHKAIVVDGKRRILISGSIHYPRSTPQMWPDLIQ 254
Query: 63 KSKDGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEW 122
K+KDGG+DVI+TYVFWN HEP GQY FE R DLV FVK AGLYVHLRIGPY+CAEW
Sbjct: 255 KAKDGGLDVIQTYVFWNGHEPSPGQYYFEDRFDLVKFVKLAQQAGLYVHLRIGPYICAEW 434
Query: 123 NYGGFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENE 182
N GGFP+WL ++ GI FRT+NEPFKA M++FTAKIV +MK+ L+ SQGGPIILSQIENE
Sbjct: 435 NLGGFPVWLKYVPGIAFRTDNEPFKAAMQKFTAKIVSLMKENRLFQSQGGPIILSQIENE 614
Query: 183 YGNIDTHDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNS 242
YG ++ K+Y WAA MA LDTGVPW+MC+Q +APDP+I+TCN FYC+ F PN
Sbjct: 615 YGPVEWEIGAPGKAYTKWAAQMAVGLDTGVPWVMCKQEDAPDPVIDTCNGFYCENFKPNK 794
Query: 243 DNKPKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTT 302
+ KPKMWTENW+GW+ FGGAVP RP EDLAF+VARF Q GG+F NYYMYHGGTNFGRT+
Sbjct: 795 NTKPKMWTENWTGWYTDFGGAVPRRPAEDLAFSVARFIQNGGSFVNYYMYHGGTNFGRTS 974
Query: 303 GGPFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETA 362
GG FI+TSYDYDAP+DEYG +PK+ HL+ LHKAIK E AL+A+DP + S G NLE
Sbjct: 975 GGLFIATSYDYDAPLDEYGLENEPKYEHLRALHKAIKQSEPALVATDPKVQSLGYNLEAH 1154
Query: 363 VYKTGAVCSAFLANIG-MSDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMIS 421
V+ C+AF+AN S A F Y LP WS+SILPDCK VV NTAKV +
Sbjct: 1155VFSAPGACAAFIANYDTKSYAKAKFGNGQYDLPPWSISILPDCKTVVYNTAKVGYGWL-- 1328
Query: 422 SFATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLS 481
+K+ ++S+ + S+ EP S D+ L EQ+N T D SDYLWY
Sbjct: 1329--------KKMTPVNSAFAWQSYNEEPASSSQADSIAAYALWEQVNVTRDSSDYLWYMTD 1484
Query: 482 IVYEDNA-----GDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGK 536
+ N G P+L + S GH LH F+NG+LAG+ G GN K+ + L G
Sbjct: 1485VNVNANEGFLKNGQSPLLTVMSAGHVLHVFINGQLAGTVWGGLGNPKLTFSDNVKLRAGN 1664
Query: 537 NTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVG 596
N + LLS+ VGL N G ++T AG+ GPV LKGL G+ DL+ Q+W+Y+VGL+GE +
Sbjct: 1665NKLSLLSVAVGLPNVGVHFETWNAGVLGPVTLKGLNEGTR-DLSRQKWSYKVGLKGESLS 1841
Query: 597 L---SSGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIG 653
L S + +W S + QPLTWYKT F AP+G++P+A+D MGKGE WVNG+SIG
Sbjct: 1842LHTESGSSSVEWIQGSLVAKKQPLTWYKTTFSAPAGNDPLALDLGSMGKGEVWVNGRSIG 2021
Query: 654 RYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEES 713
R+WP YI+ G ++CNY G Y+ +KC NCG+PSQ YHVPR+WL N+ V+FEE
Sbjct: 2022RHWPGYIA--HGSCNACNYAGYYTDTKCRTNCGQPSQRWYHVPRSWLSSGGNSLVVFEEW 2195
Query: 714 GGDPTKIS 721
GGDP I+
Sbjct: 2196GGDPNGIA 2219
>TC226828 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (47%)
Length = 1488
Score = 680 bits (1755), Expect = 0.0
Identities = 319/412 (77%), Positives = 362/412 (87%)
Frame = +1
Query: 415 NTASMISSFATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSD 474
N+AS ISSF TESLKE + S ++SS+GWSWISEPVGIS D+F ++GLLEQINTTAD+SD
Sbjct: 4 NSASAISSFTTESLKEDIGSSEASSTGWSWISEPVGISKADSFPQTGLLEQINTTADKSD 183
Query: 475 YLWYSLSIVYEDNAGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVT 534
YLWYSLSI Y+ +AG Q VLHIESLGHALHAF+NGKLAGS+ G+SG K VDIP+TLV
Sbjct: 184 YLWYSLSIDYKGDAGSQTVLHIESLGHALHAFINGKLAGSQTGNSGKYKFTVDIPVTLVA 363
Query: 535 GKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEF 594
GKNTIDLLSLTVGLQNYGAF+DT GAGITGPVILKGL NG+++DL+ Q+WTYQVGL+GE
Sbjct: 364 GKNTIDLLSLTVGLQNYGAFFDTWGAGITGPVILKGLANGNTLDLSYQKWTYQVGLKGED 543
Query: 595 VGLSSGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGR 654
+GLSSG+ GQWNSQS P NQPL WYKT F APSGS+PVAIDFTGMGKGEAWVNGQSIGR
Sbjct: 544 LGLSSGSSGQWNSQSTFPKNQPLIWYKTTFAAPSGSDPVAIDFTGMGKGEAWVNGQSIGR 723
Query: 655 YWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESG 714
YWPTY++ ++GCTDSCNYRG YSASKC +NCGKPSQTLYHVPR+WLKP N VLFEE G
Sbjct: 724 YWPTYVASDAGCTDSCNYRGPYSASKCRRNCGKPSQTLYHVPRSWLKPSGNILVLFEEKG 903
Query: 715 GDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESERKVGPVLSLECPYPNQAISSIK 774
GDPT+ISF TKQ ES+C+HV++SHPPPVD WNS+ ES RKVGPVLSL CP+ NQ ISSIK
Sbjct: 904 GDPTQISFVTKQTESLCAHVSDSHPPPVDLWNSDTESGRKVGPVLSLTCPHDNQVISSIK 1083
Query: 775 FASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFGNPCR 826
FAS+GTP GTCGN+ HG CSSN+ALSIVQKACIGSSSC++GVS TFGNPCR
Sbjct: 1084FASYGTPLGTCGNFYHGRCSSNKALSIVQKACIGSSSCSVGVSSETFGNPCR 1239
>TC207366 similar to UP|Q94BZ3 (Q94BZ3) At2g32810/F24L7.5, partial (82%)
Length = 2012
Score = 424 bits (1089), Expect = e-118
Identities = 237/544 (43%), Positives = 316/544 (57%), Gaps = 43/544 (7%)
Frame = +2
Query: 325 QPKWGHLKDLHKAIKLCEEALIASD-PTITSPGPNLETAVYK--------------TGAV 369
+PKWGHLKDLH A+KLCE AL+A+D PT GP E VY+ + ++
Sbjct: 2 EPKWGHLKDLHAALKLCEPALVATDSPTYIKLGPKQEAHVYQANVHLEGLNLSMFESSSI 181
Query: 370 CSAFLANIG-MSDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMI-------- 420
CSAFLANI +ATVTF G Y +P WSVS+LPDC+N V NTAKV + +
Sbjct: 182 CSAFLANIDEWKEATVTFRGQRYTIPPWSVSVLPDCRNTVFNTAKVRAQTSVKLVESYLP 361
Query: 421 ---SSFATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLW 477
+ F + L+ + D S S W EP+ I + +FT G+ E +N T D+SDYLW
Sbjct: 362 TVSNIFPAQQLRHQNDFYYISKS-WMTTKEPLNIWSKSSFTVEGIWEHLNVTKDQSDYLW 538
Query: 478 YSLSIVYEDNA-------GDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPI 530
YS + D+ P L I+ + L F+NG+L G+ G + V +
Sbjct: 539 YSTRVYVSDSDILFWEENDVHPKLTIDGVRDILRVFINGQLIGNVVGHW----IKVVQTL 706
Query: 531 TLVTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGL 590
+ G N + LL+ TVGLQNYGAF + GAGI G + + G +NG +DL+ WTYQVGL
Sbjct: 707 QFLPGYNDLTLLTQTVGLQNYGAFLEKDGAGIRGKIKITGFENGD-IDLSKSLWTYQVGL 883
Query: 591 QGEFVGLSS--GNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVN 648
QGEF+ S +W + TWYKT F P G +PVA+DF MGKG+AWVN
Sbjct: 884 QGEFLKFYSEENENSEWVELTPDAIPSTFTWYKTYFDVPGGIDPVALDFKSMGKGQAWVN 1063
Query: 649 GQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFV 708
GQ IGRYW T +SP SGC C+YRG Y++ KC NCGKP+QTLYHVPR+WLK +N V
Sbjct: 1064GQHIGRYW-TRVSPKSGCQQVCDYRGAYNSDKCSTNCGKPTQTLYHVPRSWLKATNNLLV 1240
Query: 709 LFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAE------SERKVGPVLSLE 762
+ EE+GG+P +IS +C+ V+ES+ PP+ NA+ S + P L L
Sbjct: 1241ILEETGGNPFEISVKLHSSRIICAQVSESNYPPLQKL-VNADLIGEEVSANNMIPELHLH 1417
Query: 763 CPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFG 822
C ISS+ FASFGTP G+C N++ G+C + ++SIV +AC G SC+I +S + FG
Sbjct: 1418C-QQGHTISSVAFASFGTPGGSCQNFSRGNCHAPSSMSIVSEACQGKRSCSIKISDSAFG 1594
Query: 823 -NPC 825
+PC
Sbjct: 1595VDPC 1606
>TC206486 similar to UP|O65761 (O65761) Beta-galactosidase (Fragment) ,
partial (61%)
Length = 1763
Score = 366 bits (940), Expect = e-101
Identities = 194/437 (44%), Positives = 265/437 (60%), Gaps = 11/437 (2%)
Frame = +2
Query: 400 ILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSGWSWISEPVGISTPD--AF 457
ILPDCKN V NTA+V + S + G+SW+S +T D +F
Sbjct: 2 ILPDCKNTVYNTARVGSQSAQMKMTRVPIH----------GGFSWLSFNEETTTTDDSSF 151
Query: 458 TKSGLLEQINTTADRSDYLWYSLSIVYEDNA-----GDQPVLHIESLGHALHAFVNGKLA 512
T +GLLEQ+NTT D SDYLWYS +V + N G PVL + S GHALH F+NG+L+
Sbjct: 152 TMTGLLEQLNTTRDLSDYLWYSTDVVLDPNERFLRNGKDPVLTVFSAGHALHVFINGQLS 331
Query: 513 GSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLK 572
G+ GS K+ + + L G N I LLS+ VGL N G ++T AG+ GP+ L GL
Sbjct: 332 GTAYGSLEFPKLTFNEGVKLRAGVNKISLLSVAVGLPNVGPHFETWNAGVLGPISLSGLN 511
Query: 573 NGSSVDLTSQQWTYQVGLQGEFVGL---SSGNVGQWNSQSNLPANQPLTWYKTNFVAPSG 629
G DL+ Q+W+Y+VGL+GE + L S + +W S + QPLTWYKT F AP+G
Sbjct: 512 EGRR-DLSWQKWSYKVGLKGETLSLHSLSGSSSVEWIQGSLVSQRQPLTWYKTTFDAPAG 688
Query: 630 SNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPS 689
+ P+A+D MGKG+ W+NGQ++GRYWP Y + SG D C+Y GTY+ +KC NCG+ S
Sbjct: 689 TAPLALDMDSMGKGQVWLNGQNLGRYWPAYKA--SGTCDYCDYAGTYNENKCRSNCGEAS 862
Query: 690 QTLYHVPRAWLKPDSNTFVLFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNA 749
Q YHVP++WLKP N V+FEE GGDP I + I+SVC+ + E P + ++
Sbjct: 863 QRWYHVPQSWLKPTGNLLVVFEELGGDPNGIFLVRRDIDSVCADIYEWQPNLI-SYQMQT 1039
Query: 750 ESERKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGS 809
+ V P + L C P Q ISSIKFASFGTP G+CGN++ GSC ++++ ++ C+G
Sbjct: 1040SGKAPVRPKVHLSCS-PGQKISSIKFASFGTPAGSCGNFHEGSCHAHKSYDAFERNCVGQ 1216
Query: 810 SSCNIGVSINTF-GNPC 825
+ C + VS F G+PC
Sbjct: 1217NWCTVTVSPENFGGDPC 1267
>TC232994 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (22%)
Length = 735
Score = 313 bits (803), Expect = 2e-85
Identities = 140/173 (80%), Positives = 154/173 (88%)
Frame = +1
Query: 654 RYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEES 713
RYWPTY SP GCTDSCNYRG Y ASKCLKNCGKPSQTLYHVPR+WL+PD NT VLFEES
Sbjct: 1 RYWPTYASPKGGCTDSCNYRGAYDASKCLKNCGKPSQTLYHVPRSWLRPDRNTLVLFEES 180
Query: 714 GGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESERKVGPVLSLECPYPNQAISSI 773
GG+P +ISF TKQI SVCSHV+ESHPPPVD+WNSN ES RKV PV+SLECPYPNQ +SSI
Sbjct: 181 GGNPKQISFATKQIGSVCSHVSESHPPPVDSWNSNTESGRKVVPVVSLECPYPNQVVSSI 360
Query: 774 KFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFGNPCR 826
KFASFGTP GTCGN+ HG CSSN+ALSIVQKACIGSSSC I +S+NTFG+PC+
Sbjct: 361 KFASFGTPLGTCGNFKHGLCSSNKALSIVQKACIGSSSCRIELSVNTFGDPCK 519
>BE660170
Length = 512
Score = 299 bits (765), Expect = 4e-81
Identities = 137/163 (84%), Positives = 152/163 (93%)
Frame = +1
Query: 1 MRGTQIVFVLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDL 60
MR +QI+ VLLWF +Y P+SF +NVTYDHRALVIDGKRRVL+SGSIHYPRSTP+MWPDL
Sbjct: 19 MRTSQILLVLLWFFCIYAPSSFGANVTYDHRALVIDGKRRVLVSGSIHYPRSTPEMWPDL 198
Query: 61 IQKSKDGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCA 120
IQKSKDGG+DVIETYVFWNLHEPVRGQYNFEGRGDLV FVK VAAAGLYVHLRIGPY CA
Sbjct: 199 IQKSKDGGLDVIETYVFWNLHEPVRGQYNFEGRGDLVKFVKVVAAAGLYVHLRIGPYACA 378
Query: 121 EWNYGGFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQ 163
EWNYGGFPLWLHFI GI+FRT+N+PF+AEMK+FTAKIVD+M +
Sbjct: 379 EWNYGGFPLWLHFIPGIQFRTDNKPFEAEMKQFTAKIVDLMNK 507
>TC235153 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (18%)
Length = 445
Score = 281 bits (719), Expect = 8e-76
Identities = 127/145 (87%), Positives = 137/145 (93%)
Frame = +3
Query: 233 FYCDQFTPNSDNKPKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMY 292
FY D+FTPNS+ KPKMWTENWSGWFL FGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMY
Sbjct: 3 FYGDEFTPNSNTKPKMWTENWSGWFLVFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMY 182
Query: 293 HGGTNFGRTTGGPFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTI 352
HGGTNF R +GGPFI+TSYDYDAPIDEYG IRQPKWGHLK++HKAIKLCEEALIA+DPTI
Sbjct: 183 HGGTNFDRASGGPFIATSYDYDAPIDEYGIIRQPKWGHLKEVHKAIKLCEEALIATDPTI 362
Query: 353 TSPGPNLETAVYKTGAVCSAFLANI 377
TS GPNLE AVYKTG+VC+AFLAN+
Sbjct: 363 TSLGPNLEAAVYKTGSVCAAFLANV 437
>TC217386 similar to GB|AAF70823.1|7939621|AF154422 beta-galactosidase
{Lycopersicon esculentum;} , partial (21%)
Length = 1267
Score = 253 bits (647), Expect = 2e-67
Identities = 130/281 (46%), Positives = 168/281 (59%), Gaps = 6/281 (2%)
Frame = +1
Query: 547 GLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGLSSGN---VG 603
GLQ G FYD +GAG+T V +KGLKNG+ +DL+S WTY++G+QGE++ L GN
Sbjct: 67 GLQTAGPFYDFIGAGLTS-VKIKGLKNGT-IDLSSYAWTYKIGVQGEYLRLYQGNGLNKV 240
Query: 604 QWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPN 663
W S S QPLTWYK AP G PV +D MGKG AW+NG+ IGRYWP
Sbjct: 241 NWTSTSEPQKMQPLTWYKAIVDAPPGDEPVGLDMLHMGKGLAWLNGEEIGRYWPRKSEFK 420
Query: 664 S-GCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKISF 722
S C C+YRG ++ KC CG+P+Q YHVPR+W KP N VLFEE GGDP KI F
Sbjct: 421 SEDCVKECDYRGKFNPDKCDTGCGEPTQRWYHVPRSWFKPSGNILVLFEEKGGDPEKIKF 600
Query: 723 GTKQIESVCSHVTESHPPP--VDTWNSNAESERKVGPVLSLECPYPNQAISSIKFASFGT 780
+++ C+ V E +P V +S + + P L CP N IS++KFASFGT
Sbjct: 601 VRRKVSGACALVAEDYPSVALVSQGEDKIQSNKNI-PFAHLTCP-SNTRISAVKFASFGT 774
Query: 781 PRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTF 821
P GTCG+Y G C + +IV+KAC+ + C I ++ F
Sbjct: 775 PSGTCGSYLKGDCHDPNSSTIVEKACLNKNDCVIKLTEENF 897
>TC213514 similar to UP|Q93X57 (Q93X57) Beta-galactosidase , partial (17%)
Length = 661
Score = 233 bits (595), Expect = 2e-61
Identities = 107/134 (79%), Positives = 119/134 (87%)
Frame = +2
Query: 693 YHVPRAWLKPDSNTFVLFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESE 752
YH+PR+WL+PDSNT VLFEESGGDPT+ISF TKQI S+CSHV+ESHPPPVD WNS +
Sbjct: 80 YHIPRSWLQPDSNTLVLFEESGGDPTQISFATKQIGSMCSHVSESHPPPVDLWNS--DKG 253
Query: 753 RKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSC 812
RKVGPVLSLECPYPNQ ISSIKFASFGTP GTCGN+ HG C SN+AL IVQKACIGSSSC
Sbjct: 254 RKVGPVLSLECPYPNQLISSIKFASFGTPYGTCGNFKHGRCRSNKALFIVQKACIGSSSC 433
Query: 813 NIGVSINTFGNPCR 826
IG+SINTFG+PC+
Sbjct: 434 RIGISINTFGDPCK 475
>TC207103 homologue to UP|O65761 (O65761) Beta-galactosidase (Fragment) ,
partial (36%)
Length = 1230
Score = 215 bits (547), Expect = 7e-56
Identities = 107/213 (50%), Positives = 133/213 (62%), Gaps = 2/213 (0%)
Frame = +2
Query: 615 QPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRG 674
QPLTWYKT F A +G P+A+D MGKG+ W+NGQS+GRYWP Y + SG CNY G
Sbjct: 98 QPLTWYKTTFDAQAGVAPLALDMGSMGKGQVWINGQSLGRYWPAYKA--SGSCGYCNYAG 271
Query: 675 TYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKISFGTKQIESVCSHV 734
TY+ KC NCG+ SQ YHVP +WLKP N V+FEE GGDP I + I+SVC+ +
Sbjct: 272 TYNEKKCGSNCGQASQRWYHVPHSWLKPTGNLLVVFEELGGDPNGIFLVRRDIDSVCADI 451
Query: 735 TESHPPPVD-TWNSNAESERKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYNHGSC 793
E P V ++ + V P L C P Q ISSIKFASFGTP G+CGNY GSC
Sbjct: 452 YEWQPNLVSYDMQASGKVRSPVRPKAHLSCG-PGQKISSIKFASFGTPVGSCGNYREGSC 628
Query: 794 SSNRALSIVQKACIGSSSCNIGVSINTF-GNPC 825
++++ QK C+G S C + VS F G+PC
Sbjct: 629 HAHKSYDAFQKNCVGQSWCTVTVSPEIFGGDPC 727
>TC219425
Length = 1170
Score = 212 bits (540), Expect = 5e-55
Identities = 130/360 (36%), Positives = 190/360 (52%), Gaps = 13/360 (3%)
Frame = +3
Query: 477 WYSLS--IVYEDNA---GDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPIT 531
WY+ S + ED + G PVL + SLGH++ AFVNG + G+ G+ P+
Sbjct: 6 WYTTSFELSQEDMSMKPGVLPVLRVMSLGHSMVAFVNGDIVGTAHGTHEEKSFEFQTPVL 185
Query: 532 LVTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQ 591
L G N I LLS TVGL + GA+ + AG IL GL G+ +DLT W ++VGL+
Sbjct: 186 LRVGTNYISLLSSTVGLPDSGAYMEHRYAGPKSINIL-GLNRGT-LDLTRNGWGHRVGLK 359
Query: 592 GE----FVGLSSGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWV 647
GE F S +V +W +P + L+WY+T F P G+ PVAI +GM KG WV
Sbjct: 360 GEGKKVFSEEGSTSV-KWKPLGAVP--RALSWYRTRFGTPEGTGPVAIRMSGMAKGMVWV 530
Query: 648 NGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTF 707
NG +IGRYW +Y+SP GKP+Q+ YH+PR++L P N
Sbjct: 531 NGNNIGRYWMSYLSP----------------------LGKPTQSEYHIPRSFLNPQDNLL 644
Query: 708 VLFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAES----ERKVGPVLSLEC 763
V+FEE P ++ +++CS V E P V++W S + + VG S+ C
Sbjct: 645 VIFEEEARVPAQVEILNVNRDTICSVVGERDPANVNSWVSRRGNFHPVVKSVGAAASMAC 824
Query: 764 PYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFGN 823
+ I +++FASFG P G CG++ GSC++ + IV++ C+G +C + + F N
Sbjct: 825 A-TGKRIVAVEFASFGNPSGYCGDFAMGSCNAAASKQIVERECLGQEACTLALDRAVFNN 1001
>TC230290 weakly similar to UP|Q93X58 (Q93X58) Beta-galactosidase , partial
(23%)
Length = 585
Score = 173 bits (438), Expect = 3e-43
Identities = 86/186 (46%), Positives = 115/186 (61%), Gaps = 1/186 (0%)
Frame = +1
Query: 641 GKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWL 700
GKG+ W+NG+SIGR+WP YI+ S C D C+Y GTY+ KC NCG+PSQ YH+PR+WL
Sbjct: 1 GKGQVWINGRSIGRHWPGYIARGS-CGD-CDYAGTYTDKKCRTNCGEPSQRWYHIPRSWL 174
Query: 701 KPDSNTFVLFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESERKVGPVLS 760
P N V+FEE GGDPT I+ + SVC+ + + P + +S + + P
Sbjct: 175 NPSGNYLVVFEEWGGDPTGITLVKRTTASVCADIYQGQPTLKN--RQMLDSGKVIRPKAH 348
Query: 761 LECPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINT 820
L CP P + IS IKFAS+G P GTCGN+ GSC ++++ QK CIG S + V
Sbjct: 349 LWCP-PGKNISKIKFASYGLPHGTCGNFREGSCHAHKSYDAPQKNCIGKQSSLVTVPPEV 525
Query: 821 F-GNPC 825
F G+PC
Sbjct: 526 FGGDPC 543
>AW153080 weakly similar to GP|14517399|gb| At2g32810/F24L7.5 {Arabidopsis
thaliana}, partial (16%)
Length = 493
Score = 154 bits (388), Expect = 2e-37
Identities = 80/184 (43%), Positives = 107/184 (57%), Gaps = 1/184 (0%)
Frame = +2
Query: 630 SNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTY-SASKCLKNCGKP 688
++PV +D G+GKGEAWVNG SIGRYW ++I+ +GC+D+C+YRG Y A KC NCG P
Sbjct: 2 NDPVVVDLQGLGKGEAWVNGLSIGRYWTSWITATNGCSDTCDYRG*YVPAQKCNTNCGDP 181
Query: 689 SQTLYHVPRAWLKPDSNTFVLFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSN 748
SQ HVPR++LK D NT VLFEE GG+P +SF T ++C+ V E
Sbjct: 182 SQMWVHVPRSFLKNDKNTLVLFEEIGGNP*NVSFRTVITGTICARVQEC----------- 328
Query: 749 AESERKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIG 808
+L L C N IS I+F+SFG P CG + + + V+ AC+G
Sbjct: 329 --------ALLELSCQGGN-TISQIQFSSFGNPTRNCGLFTKCAWEATDGQPAVEAACVG 481
Query: 809 SSSC 812
+SC
Sbjct: 482 MNSC 493
>TC204600 homologue to UP|Q9M5J3 (Q9M5J3) Beta galactosidase, partial (21%)
Length = 567
Score = 149 bits (377), Expect = 4e-36
Identities = 77/160 (48%), Positives = 99/160 (61%), Gaps = 4/160 (2%)
Frame = +2
Query: 550 NYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGLSSGN---VGQWN 606
N G ++T AGITG V+L GL +G DLT Q+W+YQ+GL+GE + L + N W
Sbjct: 2 NVGFHFET*KAGITG-VLLNGLDHGQK-DLTWQKWSYQIGLRGEAMNLVAPNGVSSVDWE 175
Query: 607 SQSNLPANQP-LTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSG 665
S +Q L W+K F AP G P+A+D + MGKG+ W+NGQSIGRYW Y G
Sbjct: 176 KDSLAVRSQSQLKWHKAYFNAPEGVEPLALDLSSMGKGQVWINGQSIGRYWMVYA---KG 346
Query: 666 CTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSN 705
SCNY GTY +KC CG+P+Q YHVPR+WL+P N
Sbjct: 347 SCSSCNYAGTYRPAKCQLGCGQPTQRWYHVPRSWLRPTKN 466
>BU760871 homologue to GP|3204134|emb beta-galactosidase {Cicer arietinum},
partial (21%)
Length = 445
Score = 107 bits (268), Expect(2) = 2e-35
Identities = 47/62 (75%), Positives = 53/62 (84%)
Frame = +2
Query: 295 GTNFGRTTGGPFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITS 354
GTNFGRT GGPFI+TSYDYDAP+DEYG RQPKWGHLKDLH+AIKLCE AL++ D T+
Sbjct: 2 GTNFGRTAGGPFIATSYDYDAPLDEYGLARQPKWGHLKDLHRAIKLCEPALVSGDSTVQR 181
Query: 355 PG 356
G
Sbjct: 182 LG 187
Score = 60.8 bits (146), Expect(2) = 2e-35
Identities = 33/90 (36%), Positives = 44/90 (48%)
Frame = +1
Query: 384 VTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSGWS 443
V F Y+LP WS+SILP+CK+ V NTA+V + S + W
Sbjct: 199 VAFGNQHYNLPPWSISILPNCKHTVYNTARVGSQSTTMKMTRVPI--------HGGLSWK 354
Query: 444 WISEPVGISTPDAFTKSGLLEQINTTADRS 473
+E + +FT +GLLEQIN T D S
Sbjct: 355 AFNEETTTTDDSSFTVTGLLEQINATRDLS 444
>AI960920 similar to GP|14970841|emb beta-galactosidase {Fragaria x
ananassa}, partial (10%)
Length = 285
Score = 146 bits (368), Expect = 4e-35
Identities = 74/93 (79%), Positives = 81/93 (86%)
Frame = +2
Query: 487 NAGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTV 546
+AG Q LHI+SLGHALHAF+NGKLAGS G+ A V VDIPITLV+GKNTIDLLSLTV
Sbjct: 5 DAGAQTFLHIKSLGHALHAFINGKLAGSGTGNHEKANVEVDIPITLVSGKNTIDLLSLTV 184
Query: 547 GLQNYGAFYDTVGAGITGPVILKGLKNGSSVDL 579
GLQNYGAF+DT GAGITGPVILK LKNGS+VDL
Sbjct: 185 GLQNYGAFFDTWGAGITGPVILKCLKNGSNVDL 283
>AW234372 homologue to GP|7682680|gb| beta galactosidase {Vigna radiata},
partial (15%)
Length = 406
Score = 142 bits (359), Expect = 5e-34
Identities = 67/105 (63%), Positives = 85/105 (80%), Gaps = 2/105 (1%)
Frame = +1
Query: 9 VLLWFLGVYVPASF--CSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKD 66
+LL F ++V + CS VTYD +A++I+G+RR+L+SGSIHYPRSTP+MW DLI+K+KD
Sbjct: 94 LLLVFSVLFVGSELIHCS-VTYDRKAIIINGQRRILISGSIHYPRSTPEMWEDLIRKAKD 270
Query: 67 GGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVH 111
GG+DVI+TYVFWN+HEP G YNFEGR DLV F+K V GLYVH
Sbjct: 271 GGLDVIDTYVFWNVHEPSPGNYNFEGRNDLVRFIKTVQRVGLYVH 405
>AW703960 similar to GP|7682677|gb| beta galactosidase {Vigna radiata},
partial (16%)
Length = 370
Score = 140 bits (354), Expect = 2e-33
Identities = 67/125 (53%), Positives = 88/125 (69%)
Frame = +2
Query: 3 GTQIVFVLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQ 62
G ++ + LW GV ++VTYDH A+V+DG RR+L+SGSIH+P TPQMWP LIQ
Sbjct: 14 GVVLMMLCLWVCGV------TASVTYDHIAIVVDGNRRILISGSIHFPSCTPQMWPYLIQ 175
Query: 63 KSKDGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEW 122
++KDGG+D I+TYVFWN+HEP + FE R DLV + K+ A LYV L IGP++C+ W
Sbjct: 176 EAKDGGLDAIQTYVFWNVHEPSPRRCYFEDRVDLV*YEKSAQ*ARLYVLLSIGPFICS*W 355
Query: 123 NYGGF 127
N GF
Sbjct: 356 NLWGF 370
>AW704810 similar to GP|7682677|gb| beta galactosidase {Vigna radiata},
partial (19%)
Length = 425
Score = 125 bits (314), Expect = 8e-29
Identities = 63/140 (45%), Positives = 88/140 (62%), Gaps = 5/140 (3%)
Frame = +3
Query: 464 EQINTTADRSDYLWYSLSIVYEDNAG-----DQPVLHIESLGHALHAFVNGKLAGSKAGS 518
EQIN T D +DYLWY + + N G PVL + S GHALH F+NG+L+G+ G
Sbjct: 9 EQINVTRDSTDYLWYMTDVNIDANEGFIKNGQSPVLTVMSAGHALHVFINGQLSGTVYGG 188
Query: 519 SGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVD 578
+ K+ + L G N I LLS+TVGL N G +++T AG+ GPV LKGL G+ D
Sbjct: 189 LDSHKLTFSDSVKLRVGNNKISLLSITVGLPNVGPYFETWNAGVLGPVTLKGLNEGTR-D 365
Query: 579 LTSQQWTYQVGLQGEFVGLS 598
L+ Q+W+Y++GL+GE + L+
Sbjct: 366 LSKQKWSYKIGLKGEALNLN 425
>BE805724
Length = 370
Score = 120 bits (302), Expect = 2e-27
Identities = 60/123 (48%), Positives = 77/123 (61%), Gaps = 4/123 (3%)
Frame = +1
Query: 577 VDLTSQQWTYQVGLQGEFVGLSSGN----VGQWNSQSNLPANQPLTWYKTNFVAPSGSNP 632
+DL+ Q+WTYQVGL+GE + L+S N V S NQPLTW+KT F AP G P
Sbjct: 10 LDLSWQKWTYQVGLKGEAMNLASPNGISSVEWMQSALVSEKNQPLTWHKTYFDAPDGDEP 189
Query: 633 VAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTL 692
+A+D GMGKG+ W+NG SIGRYW +P +G + C+Y GT+ KC G P+Q
Sbjct: 190 LALDMEGMGKGQIWINGLSIGRYW---TAPAAGICNGCSYAGTFRPPKCQVGXGXPTQQW 360
Query: 693 YHV 695
YHV
Sbjct: 361 YHV 369
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,748,532
Number of Sequences: 63676
Number of extensions: 674009
Number of successful extensions: 3317
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 3246
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3267
length of query: 826
length of database: 12,639,632
effective HSP length: 105
effective length of query: 721
effective length of database: 5,953,652
effective search space: 4292583092
effective search space used: 4292583092
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC139344.4


