

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
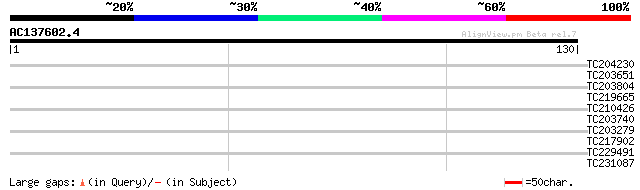
Query= AC137602.4 - phase: 0
(130 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204230 similar to UP|Q9M6E0 (Q9M6E0) DNA-binding protein 4 (Fr... 31 0.14
TC203651 UP|TBA1_PEA (P46259) Tubulin alpha-1 chain, complete 29 0.54
TC203804 homologue to UP|Q6VAF9 (Q6VAF9) Alpha-tubulin 4, complete 29 0.54
TC219665 27 2.1
TC210426 similar to UP|Q9FK53 (Q9FK53) Similarity to GAR1 protei... 27 2.1
TC203740 homologue to UP|TBA2_MAIZE (P14641) Tubulin alpha-2 cha... 27 2.7
TC203279 homologue to UP|TBA1_ANEPH (P33623) Tubulin alpha-1 cha... 26 4.6
TC217902 similar to UP|Q8S490 (Q8S490) Transcription factor RAU1... 26 6.0
TC229491 similar to GB|AAM10311.1|20147191|AY091763 AT3g15940/MV... 25 7.9
TC231087 similar to UP|Q39203 (Q39203) Receptor-like protein kin... 25 7.9
>TC204230 similar to UP|Q9M6E0 (Q9M6E0) DNA-binding protein 4 (Fragment),
partial (24%)
Length = 717
Score = 31.2 bits (69), Expect = 0.14
Identities = 17/65 (26%), Positives = 31/65 (47%)
Frame = -2
Query: 50 TLISFVAEFVNYVMHPVSMMKRCPMKQNFWMMRKRLSTRDCKNIIKEVVTIKTPIERMEN 109
T I+F+ E NY++H + +K F + + TR C + KE K + R+
Sbjct: 686 TDITFI*EKYNYILHNLFCTAFAEVKVKFKKKKSQTKTRTCSSNFKEGFKDKIKLGRIIQ 507
Query: 110 NIKQI 114
+I+ +
Sbjct: 506 DIRNV 492
>TC203651 UP|TBA1_PEA (P46259) Tubulin alpha-1 chain, complete
Length = 1725
Score = 29.3 bits (64), Expect = 0.54
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 2/91 (2%)
Frame = +3
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 855 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAYHEQLS---VAEITNSAFEPSSMMAKCD 1019
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V TIKT
Sbjct: 1020PRHGKYMACCLMYRGDVVPKDVNAAVATIKT 1112
>TC203804 homologue to UP|Q6VAF9 (Q6VAF9) Alpha-tubulin 4, complete
Length = 1692
Score = 29.3 bits (64), Expect = 0.54
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 2/91 (2%)
Frame = +1
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 886 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAYHEQLS---VAEITNSAFEPSSMMAKCD 1050
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V TIKT
Sbjct: 1051PRHGKYMACCLMYRGDVVPKDVNAAVATIKT 1143
>TC219665
Length = 1277
Score = 27.3 bits (59), Expect = 2.1
Identities = 11/25 (44%), Positives = 18/25 (72%)
Frame = +3
Query: 9 GSILWITERQTSLGLIDEIFGQVKN 33
GSI+++ + SLG ID+++ VKN
Sbjct: 123 GSIMYLVNGKLSLGSIDKLYTSVKN 197
>TC210426 similar to UP|Q9FK53 (Q9FK53) Similarity to GAR1 protein, partial
(50%)
Length = 434
Score = 27.3 bits (59), Expect = 2.1
Identities = 8/24 (33%), Positives = 16/24 (66%)
Frame = +3
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVR 39
+ T +G +DEIFG + Y++++
Sbjct: 45 KNMTQIGKVDEIFGPINEAYFSIK 116
>TC203740 homologue to UP|TBA2_MAIZE (P14641) Tubulin alpha-2 chain (Alpha-2
tubulin), complete
Length = 1600
Score = 26.9 bits (58), Expect = 2.7
Identities = 27/91 (29%), Positives = 39/91 (42%), Gaps = 2/91 (2%)
Frame = +1
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 814 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAYHEQLS---VAEITNSAFEPSSMMAKCD 978
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V IKT
Sbjct: 979 PRHGKYMACCLMYRGDVVPKDVNAAVAIIKT 1071
>TC203279 homologue to UP|TBA1_ANEPH (P33623) Tubulin alpha-1 chain, partial
(39%)
Length = 773
Score = 26.2 bits (56), Expect = 4.6
Identities = 17/50 (34%), Positives = 23/50 (46%), Gaps = 2/50 (4%)
Frame = +3
Query: 55 VAEFVNYVMHPVSMMKRCPMKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
VAE N P SMM +C + +M + D K++ V TIKT
Sbjct: 144 VAEITNSAFEPSSMMAKCDPRHGKYMACCLMYRGDVVPKDVNAAVATIKT 293
>TC217902 similar to UP|Q8S490 (Q8S490) Transcription factor RAU1 (Fragment),
partial (69%)
Length = 1451
Score = 25.8 bits (55), Expect = 6.0
Identities = 14/42 (33%), Positives = 25/42 (59%), Gaps = 7/42 (16%)
Frame = +1
Query: 52 ISFVAEFVNY--VMHPVSMMKRCPMK-----QNFWMMRKRLS 86
++++A +NY V+H + +R MK Q FW +RK++S
Sbjct: 676 VTYLASPMNYGKVLHSMPQKQRMKMKSCFPLQIFWNLRKQIS 801
>TC229491 similar to GB|AAM10311.1|20147191|AY091763 AT3g15940/MVC8_7
{Arabidopsis thaliana;} , partial (45%)
Length = 1307
Score = 25.4 bits (54), Expect = 7.9
Identities = 13/51 (25%), Positives = 20/51 (38%)
Frame = +3
Query: 45 GIREGTLISFVAEFVNYVMHPVSMMKRCPMKQNFWMMRKRLSTRDCKNIIK 95
G G V V ++HPV + QN W + K S R +++
Sbjct: 891 GTDAGGTQEIVEHNVTGLLHPVGHPGNLVLAQNLWFLLKNQSARKQMGVVR 1043
>TC231087 similar to UP|Q39203 (Q39203) Receptor-like protein kinase
precursor, partial (30%)
Length = 826
Score = 25.4 bits (54), Expect = 7.9
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -2
Query: 103 PIERMENNIKQIPLKDGSIPTMP 125
P+ R +I Q+PLK P+MP
Sbjct: 72 PLHRHNRSITQLPLKHSPKPSMP 4
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,482,970
Number of Sequences: 63676
Number of extensions: 58428
Number of successful extensions: 277
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 277
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 277
length of query: 130
length of database: 12,639,632
effective HSP length: 86
effective length of query: 44
effective length of database: 7,163,496
effective search space: 315193824
effective search space used: 315193824
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC137602.4


