

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
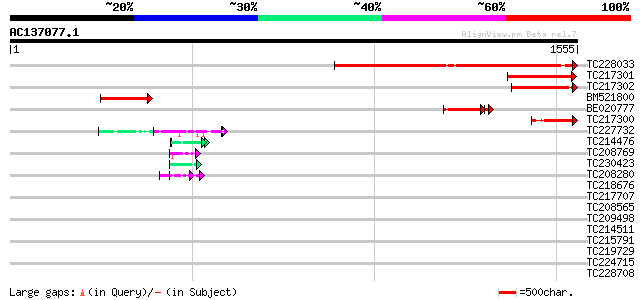
Query= AC137077.1 - phase: 0
(1555 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC228033 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (... 1021 0.0
TC217301 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (5%) 276 7e-74
TC217302 263 4e-70
BM521800 233 5e-61
BE020777 187 1e-53
TC217300 147 4e-35
TC227732 similar to UP|Q6P623 (Q6P623) Fibrillarin, partial (16%) 50 1e-05
TC214476 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 47 5e-05
TC208769 similar to PIR|C85356|C85356 glycine-rich protein [impo... 45 2e-04
TC230423 similar to UP|Q41187 (Q41187) Glycine-rich protein (Fra... 44 4e-04
TC208280 similar to PIR|C85356|C85356 glycine-rich protein [impo... 44 6e-04
TC218676 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, part... 42 0.002
TC217707 similar to UP|Q941H8 (Q941H8) RNA-binding protein precu... 42 0.002
TC208565 similar to GB|AAM97131.1|22531255|AY136466 ribonucleopr... 42 0.002
TC209498 similar to UP|Q8L8Z3 (Q8L8Z3) Structural protein-like p... 41 0.004
TC214511 weakly similar to PIR|C85356|C85356 glycine-rich protei... 41 0.004
TC215791 similar to GB|CAE76446.1|38567152|BX842633 related to p... 40 0.006
TC219729 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 40 0.010
TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%) 39 0.018
TC228708 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wal... 38 0.030
>TC228033 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (25%)
Length = 2651
Score = 1021 bits (2640), Expect = 0.0
Identities = 523/667 (78%), Positives = 571/667 (85%), Gaps = 3/667 (0%)
Frame = +2
Query: 892 EGQLWGPALVLASQLGEQFYVDTVRQMALRQLVAGSPLRTLCLLIAGRPNDVFPTEETSI 951
+GQLWGPALVLASQLGEQFYVDTV+QMALRQLV+GSPLRTLCLLIAG+ ++F T+ TS
Sbjct: 5 QGQLWGPALVLASQLGEQFYVDTVKQMALRQLVSGSPLRTLCLLIAGQQAEIFSTD-TSN 181
Query: 952 SGHPGAVGMPQQSEQAGSNDMLEDWEENLAVITANRTKGDELVMMHLGDCLWKEKREITA 1011
SGHPGA M QQS Q GSN ML+DWEENLAVITANRTK DELV++HLGDCLWKE+ EITA
Sbjct: 182 SGHPGASNMAQQSPQVGSNGMLDDWEENLAVITANRTKDDELVIIHLGDCLWKERSEITA 361
Query: 1012 AHICYLIAEVNFSSYSDATRLCLIGADHWTRPRTYASPEAIQRTELYEYSKLLGNSQFVL 1071
AHICYL+AE NF SYSD+ RLCLIGADHW PRTYASPEAIQRTELYEYSK++GNSQF L
Sbjct: 362 AHICYLVAEANFESYSDSARLCLIGADHWKCPRTYASPEAIQRTELYEYSKVVGNSQFTL 541
Query: 1072 HSFQPYKLIYAHMLAEVGKVSDSLKYCQAVLKSLKTGRAPEVETWKQLVLALEERIRTHQ 1131
H FQPYKLIYA +LAEVGKVSDSLKYCQA+LKSLKTGRAPEVE+WKQL L+LEERIR HQ
Sbjct: 542 HPFQPYKLIYAFLLAEVGKVSDSLKYCQALLKSLKTGRAPEVESWKQLALSLEERIRIHQ 721
Query: 1132 QGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPTSSQATVHGSEQHYQHMAPRVST 1191
QGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAP+SS TVHGSE+ YQ+MAPRVS+
Sbjct: 722 QGGYAANLAPAKLVGKLLNFFDSTAHRVVGGLPPPAPSSSAGTVHGSEKQYQNMAPRVSS 901
Query: 1192 SQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSEPDIGRSPRQESPSPDAQGKVQ 1251
SQSTM SL PSAS+EPISEWTADNNRM KPNRSVS+PD GR+PRQE+ SPDAQ K Q
Sbjct: 902 SQSTM---SLAPSASMEPISEWTADNNRM-GKPNRSVSDPDFGRTPRQETTSPDAQEKPQ 1069
Query: 1252 VSGGASRFSRFGFGSQLLQKTVGLVL--RSGKQAKLGEKNKFYYDEKLKRWVEEGAEVPA 1309
SGG SRFSRFGFGSQLLQKTVGLVL RSG+QAKLG+KNKFYYDEKLKRWVEEGAEVPA
Sbjct: 1070 ASGGTSRFSRFGFGSQLLQKTVGLVLKPRSGRQAKLGDKNKFYYDEKLKRWVEEGAEVPA 1249
Query: 1310 EEAALPPPPPTTAAFQNGSADYNLKSALKTEGLTPNEFSSTRTSSPELSPGMPPIPPSSN 1369
EEAA PPPTTAAFQNGS +YNL+SALKTE P E SS RTSS ELSPGMP IPPS+N
Sbjct: 1250 EEAAALTPPPTTAAFQNGSTEYNLRSALKTESSPPIEGSSIRTSSLELSPGMPLIPPSAN 1429
Query: 1370 QFSARSRLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKPALPANAKFFIPAPVPSSSEQN 1429
QFSAR RLGVRSRYVDTFNQ GG+SANLFQSPSV SVKPA+ ANAKFFIP+ PSS+EQ
Sbjct: 1430 QFSARGRLGVRSRYVDTFNQGGGTSANLFQSPSVPSVKPAVAANAKFFIPSAAPSSNEQT 1609
Query: 1430 MEAIAESNLEDSAANENPSTSSTND-WSYHPPKHAQTMTMQRFPSAGNISKQGQTDGNES 1488
MEAI ES EDSA NE+PSTS+TN+ WSY PK + T+QRFPS GNIS Q T+G+ S
Sbjct: 1610 MEAIVESKQEDSATNEDPSTSATNEWWSYQSPKQVSSTTIQRFPSLGNISNQRATEGSNS 1789
Query: 1489 HFSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFMPDPSSLMQGPTRSGSFGE 1548
H HSRRT+SWSGSFNDSF+PPKM GMP S FMPD SLM+ +S S+ E
Sbjct: 1790 HLPHSRRTSSWSGSFNDSFTPPKM----------GMPSSRFMPD-ESLMRTHVKSSSYAE 1936
Query: 1549 DLQEVEL 1555
DLQEVEL
Sbjct: 1937 DLQEVEL 1957
>TC217301 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (5%)
Length = 1110
Score = 276 bits (705), Expect = 7e-74
Identities = 136/189 (71%), Positives = 149/189 (77%)
Frame = +2
Query: 1366 PSSNQFSARSRLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKPALPANAKFFIPAPVPSS 1425
PSSNQFSAR RLGVRSRYVDTFNQ GG+SANLFQSPSV SVKP L ANAKFF+P P PSS
Sbjct: 8 PSSNQFSARGRLGVRSRYVDTFNQGGGTSANLFQSPSVPSVKPVLAANAKFFVPTPAPSS 187
Query: 1426 SEQNMEAIAESNLEDSAANENPSTSSTNDWSYHPPKHAQTMTMQRFPSAGNISKQGQTDG 1485
+E+ +EAI ES ED+A NE PS S+TN+WSY PKH + T+QRFPS GNIS Q DG
Sbjct: 188 NERTIEAIVESKQEDNATNEYPSISTTNEWSYQSPKHVSSTTIQRFPSMGNISNQVAADG 367
Query: 1486 NESHFSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFMPDPSSLMQGPTRSGS 1545
N SH HSRRTASWSGSFNDSF+P KMG IKP G ALGMPPS F PD SLM P +S S
Sbjct: 368 NNSHLPHSRRTASWSGSFNDSFTPQKMGNIKPLGEALGMPPSRFSPD-ESLMHKPVKSSS 544
Query: 1546 FGEDLQEVE 1554
+GEDL EVE
Sbjct: 545 YGEDLHEVE 571
>TC217302
Length = 590
Score = 263 bits (672), Expect = 4e-70
Identities = 130/180 (72%), Positives = 144/180 (79%)
Frame = +1
Query: 1376 RLGVRSRYVDTFNQNGGSSANLFQSPSVQSVKPALPANAKFFIPAPVPSSSEQNMEAIAE 1435
RLGVRSRYVDTFNQ GG+SANLFQSPSV SVKPAL ANAKFF+P P PSS+EQ M+AIAE
Sbjct: 1 RLGVRSRYVDTFNQGGGTSANLFQSPSVPSVKPALAANAKFFVPTPAPSSNEQAMDAIAE 180
Query: 1436 SNLEDSAANENPSTSSTNDWSYHPPKHAQTMTMQRFPSAGNISKQGQTDGNESHFSHSRR 1495
EDSA NE PSTS+TNDWSY PKH + +QRFPS GNISKQG T+G+ SH HSRR
Sbjct: 181 GKQEDSATNEYPSTSATNDWSYRSPKHVSSTAIQRFPSMGNISKQGATEGSNSHLPHSRR 360
Query: 1496 TASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFMPDPSSLMQGPTRSGSFGEDLQEVEL 1555
TASWSGSFNDSF+P KMG +KP G ALGMP S + PD SS M P +S S+GEDL EVEL
Sbjct: 361 TASWSGSFNDSFTPQKMGNMKPLGEALGMPLSRYSPDESS-MHKPVKSSSYGEDLHEVEL 537
>BM521800
Length = 428
Score = 233 bits (594), Expect = 5e-61
Identities = 110/142 (77%), Positives = 123/142 (86%)
Frame = +1
Query: 249 GDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSSSQNWEDLYPGWKYDHITGQWYQIED 308
GDGLN S N+VQYQEG Y ASS H NGQDLSSSQ WEDLYPGWKYDH TGQWYQI+
Sbjct: 4 GDGLNASANHVQYQEGE-TYVASSEEHPNGQDLSSSQYWEDLYPGWKYDHNTGQWYQIDG 180
Query: 309 YNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQSVAGTLAETGTTESVSSWNQVSQ 368
Y T+T+QQ+SEANTA D +AASDGKTEISY+QQ AQSVAGTLAE+GTT++VSSW+QVS+
Sbjct: 181 YIVTSTTQQSSEANTAADLSAASDGKTEISYMQQTAQSVAGTLAESGTTKNVSSWSQVSE 360
Query: 369 GNNGYLEHMVFDPQYPDWYYDT 390
GNNGY EHM+FDPQYP WYYDT
Sbjct: 361 GNNGYPEHMIFDPQYPGWYYDT 426
>BE020777
Length = 413
Score = 187 bits (475), Expect(2) = 1e-53
Identities = 97/118 (82%), Positives = 105/118 (88%), Gaps = 2/118 (1%)
Frame = +3
Query: 1189 VSTSQSTMAMSSLVPSASLEPISEWTADNNRMAAKPNRSVSEPDIGRSPRQESPSPDAQG 1248
VS+SQSTM SL PSAS+EPIS+WTADNN+MA KPNRS+SEPDIGR+PRQE+ SPD QG
Sbjct: 6 VSSSQSTM---SLAPSASMEPISDWTADNNKMA-KPNRSISEPDIGRTPRQETTSPDIQG 173
Query: 1249 KVQVSGGASRFSRFGFGSQLLQKTVGLVL--RSGKQAKLGEKNKFYYDEKLKRWVEEG 1304
K Q SGG SRFSRFGFGSQLLQKTVGLVL RSG+QAKLGEKNKFYYDEKLKRWV EG
Sbjct: 174 KAQASGGTSRFSRFGFGSQLLQKTVGLVLKPRSGRQAKLGEKNKFYYDEKLKRWVXEG 347
Score = 42.7 bits (99), Expect(2) = 1e-53
Identities = 19/23 (82%), Positives = 21/23 (90%)
Frame = +1
Query: 1303 EGAEVPAEEAALPPPPPTTAAFQ 1325
+GAE+PAEEAA PPPPTTAAFQ
Sbjct: 343 KGAELPAEEAAALPPPPTTAAFQ 411
>TC217300
Length = 793
Score = 147 bits (371), Expect = 4e-35
Identities = 81/126 (64%), Positives = 89/126 (70%)
Frame = +2
Query: 1430 MEAIAESNLEDSAANENPSTSSTNDWSYHPPKHAQTMTMQRFPSAGNISKQGQTDGNESH 1489
MEAIAES EDSA NE SY K + T+QRFPS GNIS QG TDGN SH
Sbjct: 2 MEAIAESKQEDSAXNE---------CSYQSXK--SSTTIQRFPSLGNISNQGATDGNNSH 148
Query: 1490 FSHSRRTASWSGSFNDSFSPPKMGEIKPSGAALGMPPSAFMPDPSSLMQGPTRSGSFGED 1549
HSRRTASWSGSFNDSF+P KMG IKP G +LGMPPS F+PD SLM+ +S S+GED
Sbjct: 149 LPHSRRTASWSGSFNDSFTPRKMGNIKPLGESLGMPPSRFLPD-ESLMRTHVKSSSYGED 325
Query: 1550 LQEVEL 1555
LQEVEL
Sbjct: 326 LQEVEL 343
>TC227732 similar to UP|Q6P623 (Q6P623) Fibrillarin, partial (16%)
Length = 1693
Score = 49.7 bits (117), Expect = 1e-05
Identities = 81/376 (21%), Positives = 132/376 (34%), Gaps = 23/376 (6%)
Frame = +3
Query: 243 NHTGKQGDGLNTSVNYVQYQEGGGAYDASSNLHNNGQDLSS-SQNWEDLYPGWKYDHITG 301
N G Q L+ N+ G A D S+ N+G S+ + NW G ++ +
Sbjct: 105 NEPGNQDSNLDKKSNWNSGNSGNLASDPKSSNWNSGSGNSNENSNW-----GTNVNNKSS 269
Query: 302 QWYQIEDYNATTTSQQTSEANTAVDWAAASDGKTEISYLQQAAQSVAGTLAETGTTESVS 361
E+ N++ +S + N + S+ + S Q + + + + S
Sbjct: 270 WGTGNENKNSSWSSGHSDPGNQDANQGKKSNWNSGNSGNQPSDPN-------SNWNSNKS 428
Query: 362 SWNQVSQGNNGYLEHMVFDPQYPDWYYDTIAQEWRSLATYNSSVQSSVHGLQNGHTS-TS 420
SW S GN + +W + N+ + S G N +TS S
Sbjct: 429 SW---SAGNEN---------KKSNWSSGDPGNTDSNWGNKNNCISGS--GDANQNTSWRS 566
Query: 421 TSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAG----NYDGSHGVGSQA 476
SS+N N+ + ++ N S GVG G + G G + G + G G
Sbjct: 567 NSSWNTANASSDDGNEGSNENSDGVGGGNWRGGYRGRGGSDRGGFRGRGFRGRGERGGFG 746
Query: 477 VNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG--------VNQAGNYGGSQGVGS 528
G G G + G +GG + G W G +Q+G + + G
Sbjct: 747 GRGEHGGFGGRGERGGFGGRGRSDREGFGGRWGSEGGRGGRGRGRNDQSGGWNNRRDSGE 926
Query: 529 Q--------AVNGSWSGSHGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVN 580
A NG W S+G Q ++ + + S GN H NQ
Sbjct: 927 DGPSDWKKGADNGGWKNSNG---SQAWNQDTGDKD----RQSWSQGNADKEHPSWNQGSG 1085
Query: 581 TSSSFGSVA-LNNKGS 595
+ S+ S + N+ GS
Sbjct: 1086RNQSWSSASGANDNGS 1133
Score = 43.5 bits (101), Expect = 7e-04
Identities = 53/215 (24%), Positives = 88/215 (40%), Gaps = 12/215 (5%)
Frame = +3
Query: 395 WRSLATYNSSVQSSVHGLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSW 454
W + +NS +S QN H S+ + + +S + S + S + S + +W
Sbjct: 27 WGQKSNWNSGSRSGNEN-QNSHWSSGRNEPGNQDSNLDKKSNWNSGNSGNLASDPKSSNW 203
Query: 455 SGSHGVSQAG-------NYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGS 507
+ G S N S G G++ N SWS H + GN +QG S +G+
Sbjct: 204 NSGSGNSNENSNWGTNVNNKSSWGTGNENKNSSWSSGH--SDPGNQDANQGKKSNWNSGN 377
Query: 508 WSGSHGVNQAGNYG---GSQGVGSQAVNGSWSGSHGVNHQQGFDMYATEASTKIGNNTAS 564
SG+ + N+ S G++ +WS N T+++ NN S
Sbjct: 378 -SGNQPSDPNSNWNSNKSSWSAGNENKKSNWSSGDPGN---------TDSNWGNKNNCIS 527
Query: 565 -SGNQQVHHSY-GNQQVNTSSSFGSVALNNKGSFE 597
SG+ + S+ N NT+++ S N+GS E
Sbjct: 528 GSGDANQNTSWRSNSSWNTANA--SSDDGNEGSNE 626
>TC214476 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (14%)
Length = 642
Score = 47.4 bits (111), Expect = 5e-05
Identities = 38/103 (36%), Positives = 42/103 (39%), Gaps = 4/103 (3%)
Frame = -3
Query: 442 SQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGS 501
S+G G A G G G G+ G GS GS GS G G GSQG GS
Sbjct: 640 SEGGGGSAGGGGSKGGGGYGGGGSQGGG---GSAGGGGS*GGSGGSAGGGEGSGSQGGGS 470
Query: 502 ----QAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG 540
GS SG G G+ GGS+G GS G G G
Sbjct: 469 GGGGSQGGGSGSGGGGYGGGGS-GGSEGGGSSGGGGGGGGGGG 344
Score = 45.1 bits (105), Expect = 2e-04
Identities = 36/103 (34%), Positives = 41/103 (38%)
Frame = -3
Query: 444 GVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQA 503
G GS GS GS G + G GS G GS GS G G + G YGG GS+
Sbjct: 562 GGGSAGGGGS*GGSGGSAGGGEGSGSQGGGSGG-GGSQGGGSG-SGGGGYGGGGSGGSE- 392
Query: 504 VNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNHQQG 546
G SG G G GG G G G +H +G
Sbjct: 391 -GGGSSGGGGGGGGGGGGGGYGGGGDKGGGIGGDKGDYHHGKG 266
Score = 35.8 bits (81), Expect = 0.15
Identities = 35/96 (36%), Positives = 39/96 (40%), Gaps = 5/96 (5%)
Frame = -3
Query: 435 SQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNY- 493
SQ G S G GSQ GS SG G G+ GS G GS G G G G Y
Sbjct: 487 SQGGG--SGGGGSQG-GGSGSGGGGYGGGGS-GGSEGGGSSGGGGGGGGGGG---GGGYG 329
Query: 494 -GGSQGVGSQAVNGSW---SGSHGVNQAGNYGGSQG 525
GG +G G G + G G G+ GG G
Sbjct: 328 GGGDKGGGIGGDKGDYHHGKGGKGDKDGGDKGGGGG 221
>TC208769 similar to PIR|C85356|C85356 glycine-rich protein [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(53%)
Length = 679
Score = 45.1 bits (105), Expect = 2e-04
Identities = 38/98 (38%), Positives = 52/98 (52%), Gaps = 12/98 (12%)
Frame = +1
Query: 438 GNHVSQGV------------GSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSH 485
G V GV G+ A +GS SGS S +G+ S G GS A GS +GS+
Sbjct: 82 GIGVGAGVGLGLGGGGGAGSGAGAGSGSGSGSSSYSSSGSSSSSSGSGSGA--GSEAGSY 255
Query: 486 GVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGS 523
+ AG+ GS GS+A GS++GSH +G+YGG+
Sbjct: 256 AGSHAGS--GSSRAGSEA--GSYAGSHA--GSGSYGGN 351
Score = 39.3 bits (90), Expect = 0.014
Identities = 25/76 (32%), Positives = 37/76 (47%)
Frame = +1
Query: 468 GSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVG 527
G G G+ A +GS SGS + +G+ S G SGS ++AG+Y GS
Sbjct: 127 GGAGSGAGAGSGSGSGSSSYSSSGSSSSSSG----------SGSGAGSEAGSYAGSHAGS 276
Query: 528 SQAVNGSWSGSHGVNH 543
+ GS +GS+ +H
Sbjct: 277 GSSRAGSEAGSYAGSH 324
Score = 37.7 bits (86), Expect = 0.040
Identities = 28/70 (40%), Positives = 35/70 (50%), Gaps = 1/70 (1%)
Frame = +1
Query: 411 GLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSH 470
G +G S+S SS +S S AG+ GS A +GS S S+AG+Y GSH
Sbjct: 154 GSGSGSGSSSYSSSGSSSSSSGSGSGAGSEAGSYAGSHAGSGS---SRAGSEAGSYAGSH 324
Query: 471 -GVGSQAVNG 479
G GS NG
Sbjct: 325 AGSGSYGGNG 354
Score = 30.8 bits (68), Expect = 4.8
Identities = 25/84 (29%), Positives = 34/84 (39%), Gaps = 8/84 (9%)
Frame = +1
Query: 486 GVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQG--------VGSQAVNGSWSG 537
GV GG G G+ A +GS SGS + +G+ S G GS A + + SG
Sbjct: 100 GVGLGLGGGGGAGSGAGAGSGSGSGSSSYSSSGSSSSSSGSGSGAGSEAGSYAGSHAGSG 279
Query: 538 SHGVNHQQGFDMYATEASTKIGNN 561
S + G + S G N
Sbjct: 280 SSRAGSEAGSYAGSHAGSGSYGGN 351
>TC230423 similar to UP|Q41187 (Q41187) Glycine-rich protein (Fragment),
partial (40%)
Length = 522
Score = 44.3 bits (103), Expect = 4e-04
Identities = 28/89 (31%), Positives = 35/89 (38%), Gaps = 1/89 (1%)
Frame = +2
Query: 438 GNHVSQGVGSQA-VNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGS 496
G V G+G G + G HG G G G G A G + G G G +GG
Sbjct: 5 GGGVGGGIGGGGGAGGGFGGGHGGGVGGGIGGGAGGGGGA-GGGFGGGAGGGAGGGFGGG 181
Query: 497 QGVGSQAVNGSWSGSHGVNQAGNYGGSQG 525
G G+ G + G G G +GG G
Sbjct: 182 AGGGA---GGGFGGGAGGGAGGGFGGGGG 259
Score = 40.4 bits (93), Expect = 0.006
Identities = 28/86 (32%), Positives = 34/86 (38%)
Frame = +2
Query: 462 QAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYG 521
+ G G G G A G + G HG G GG G G A G + G G G +G
Sbjct: 2 KGGGVGGGIGGGGGA-GGGFGGGHGGGVGGGIGGGAGGGGGA-GGGFGGGAGGGAGGGFG 175
Query: 522 GSQGVGSQAVNGSWSGSHGVNHQQGF 547
G G G+ G + G G GF
Sbjct: 176 GGAGGGA---GGGFGGGAGGGAGGGF 244
Score = 38.9 bits (89), Expect = 0.018
Identities = 28/91 (30%), Positives = 36/91 (38%)
Frame = +2
Query: 450 VNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWS 509
V G G G G + G HG G G +G G G +GG G G+ G +
Sbjct: 14 VGGGIGGGGGAG--GGFGGGHGGGVGGGIGGGAGGGG-GAGGGFGGGAGGGA---GGGFG 175
Query: 510 GSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG 540
G G G +GG G G+ G + G G
Sbjct: 176 GGAGGGAGGGFGGGAGGGA---GGGFGGGGG 259
>TC208280 similar to PIR|C85356|C85356 glycine-rich protein [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(52%)
Length = 762
Score = 43.9 bits (102), Expect = 6e-04
Identities = 38/97 (39%), Positives = 54/97 (55%), Gaps = 1/97 (1%)
Frame = +3
Query: 438 GNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQ 497
G V G+G A +G+ +GS S + +Y S GS + +GS SG+ ++AG+Y GS
Sbjct: 201 GAGVGLGLGG-AGSGAGAGSGSGSGSSSYSSS---GSSSSSGSGSGAG--SEAGSYAGSH 362
Query: 498 GVGSQAVNGSWSGSHGVNQAGNYGGSQ-GVGSQAVNG 533
GS A +GS S ++AG+Y GS G GS NG
Sbjct: 363 -AGSSAGSGS---SRAGSEAGSYAGSHAGSGSYGGNG 461
Score = 43.9 bits (102), Expect = 6e-04
Identities = 36/97 (37%), Positives = 47/97 (48%), Gaps = 1/97 (1%)
Frame = +3
Query: 411 GLQNGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSH 470
GL G + + + S S YS +G+ S G SGS S+AG+Y GSH
Sbjct: 213 GLGLGGAGSGAGAGSGSGSGSSSYSSSGSSSSSG----------SGSGAGSEAGSYAGSH 362
Query: 471 GVGSQAVNGSWSGSHGVNQAGNYGGSQ-GVGSQAVNG 506
GS A +GS S ++AG+Y GS G GS NG
Sbjct: 363 -AGSSAGSGS---SRAGSEAGSYAGSHAGSGSYGGNG 461
Score = 35.8 bits (81), Expect = 0.15
Identities = 31/87 (35%), Positives = 39/87 (44%), Gaps = 8/87 (9%)
Frame = +3
Query: 468 GSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQ--- 524
G G GS A GS SGS G S S + + S SGS ++AG+Y GS
Sbjct: 219 GLGGAGSGAGAGSGSGS---------GSSSYSSSGSSSSSGSGSGAGSEAGSYAGSHAGS 371
Query: 525 --GVGSQAVN---GSWSGSHGVNHQQG 546
G GS GS++GSH + G
Sbjct: 372 SAGSGSSRAGSEAGSYAGSHAGSGSYG 452
>TC218676 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, partial (5%)
Length = 879
Score = 42.4 bits (98), Expect = 0.002
Identities = 37/116 (31%), Positives = 48/116 (40%), Gaps = 15/116 (12%)
Frame = +3
Query: 438 GNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGS----WSGSHGVNQ---- 489
G+ V G+G +GS SG G+Y + G+GS G + +H +N
Sbjct: 12 GDGVGAGMGIXXNSGSGSGPGQYGGPGHYGPTVGMGSYGGFGGQPPMGNNNHPLNSSNPV 191
Query: 490 ------AGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYG-GSQGVGSQAVNGSWSGS 538
AG YGG G G G GS G AG G G GVG + G+ GS
Sbjct: 192 PGSMAGAGGYGGGMG-GPYGGYGGGPGSTGFAGAGGMGAGGGGVGGGGLGGAGGGS 356
Score = 33.9 bits (76), Expect = 0.57
Identities = 35/123 (28%), Positives = 45/123 (36%), Gaps = 15/123 (12%)
Frame = +3
Query: 434 YSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNY 493
Y G G + +N S ++ AG Y G G G G GS G AG
Sbjct: 123 YGGFGGQPPMGNNNHPLNSSNPVPGSMAGAGGYGGGMG-GPYGGYGGGPGSTGFAGAGGM 299
Query: 494 G-GSQGVGSQAVNGSWSGS--------------HGVNQAGNYGGSQGVGSQAVNGSWSGS 538
G G GVG + G+ GS G +G+YG S G Q + SG+
Sbjct: 300 GAGGGGVGGGGLGGAGGGSMYRLPGSGGMPAGGGGYPDSGHYGLSASSGYQNQHHPPSGA 479
Query: 539 HGV 541
V
Sbjct: 480 SPV 488
>TC217707 similar to UP|Q941H8 (Q941H8) RNA-binding protein precursor,
partial (35%)
Length = 1008
Score = 42.0 bits (97), Expect = 0.002
Identities = 37/100 (37%), Positives = 47/100 (47%), Gaps = 3/100 (3%)
Frame = +1
Query: 446 GSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYG-GSQGVGSQAV 504
G GS GS G ++ GNY G + SGS G N+ GNYG G+ V S
Sbjct: 421 GDGGYRGS-GGSDGYNRGGNYGGGYN----------SGSDGYNRGGNYGSGNYNVTSSYS 567
Query: 505 NGSWSGSH--GVNQAGNYGGSQGVGSQAVNGSWSGSHGVN 542
+G+ S+ G N AGNY ++ G V GS SG N
Sbjct: 568 DGNAETSYTSGAN-AGNYQFNENSG--GVFGSASGEFSSN 678
Score = 35.0 bits (79), Expect = 0.26
Identities = 35/106 (33%), Positives = 45/106 (42%), Gaps = 3/106 (2%)
Frame = +1
Query: 414 NGHTSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNY-DGSHGV 472
N T S F D Y +G G G SGS G ++ GNY G++ V
Sbjct: 385 NYATERSRPGFGGDGG----YRGSGGSDGYNRGGNYGGGYNSGSDGYNRGGNYGSGNYNV 552
Query: 473 GSQAVNGSWSGSH--GVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQ 516
S +G+ S+ G N AGNY ++ G V GS SG NQ
Sbjct: 553 TSSYSDGNAETSYTSGAN-AGNYQFNENSG--GVFGSASGEFSSNQ 681
>TC208565 similar to GB|AAM97131.1|22531255|AY136466 ribonucleoprotein-like
{Arabidopsis thaliana;} , partial (34%)
Length = 936
Score = 42.0 bits (97), Expect = 0.002
Identities = 28/107 (26%), Positives = 41/107 (38%)
Frame = +2
Query: 432 SEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAG 491
S Y G+ G G + G + AG Y G +G G G +G G
Sbjct: 305 SAYGGFGSEFGGYGGYAGAMGPYRGDPSLGYAGRYGGGYGRGYDL------GGYGGPSDG 466
Query: 492 NYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGS 538
YG GVG + ++ S+ + G YGG G G ++G+
Sbjct: 467 -YGAYGGVGGGSSGSAYGSSYDASLGGGYGGPAGGSFYGTRGGYAGA 604
>TC209498 similar to UP|Q8L8Z3 (Q8L8Z3) Structural protein-like protein,
partial (51%)
Length = 795
Score = 41.2 bits (95), Expect = 0.004
Identities = 30/92 (32%), Positives = 39/92 (41%), Gaps = 4/92 (4%)
Frame = +2
Query: 446 GSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNG----SWSGSHGVNQAGNYGGSQGVGS 501
G + G+ +G+ G N+D S G GS G S SG A YG G G+
Sbjct: 224 GGAPLAGTPTGATGSGHGANWDYSWGWGSGPGTGYGYGSGSGRSPTGFARGYGYGFGSGT 403
Query: 502 QAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNG 533
+ +G GS G AG YG G G+ G
Sbjct: 404 GSGSGYGYGSGGGAHAGGYGSGSGSGNSGGGG 499
Score = 35.4 bits (80), Expect = 0.20
Identities = 29/108 (26%), Positives = 45/108 (40%)
Frame = +2
Query: 479 GSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGS 538
G+ HG N ++G G G+ GS SG A YG G G+ + +G GS
Sbjct: 254 GATGSGHGANWDYSWGWGSGPGTGYGYGSGSGRSPTGFARGYGYGFGSGTGSGSGYGYGS 433
Query: 539 HGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVNTSSSFG 586
G H G+ + + GN+ + + N ++ +SSS G
Sbjct: 434 GGGAHAGGYG-----SGSGSGNSGGGGYDDKNGGGDDNNRLPSSSSRG 562
>TC214511 weakly similar to PIR|C85356|C85356 glycine-rich protein [imported]
- Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(88%)
Length = 1085
Score = 41.2 bits (95), Expect = 0.004
Identities = 30/107 (28%), Positives = 47/107 (43%), Gaps = 3/107 (2%)
Frame = +2
Query: 444 GVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGV---NQAGNYGGSQGVG 500
G+G + +G+ +G+ S +G+ S S A + S SGS G ++AG+Y GS+
Sbjct: 512 GIGGGSGSGAGAGAGSGSGSGSGSSSSSASSSASSSSSSGSGGSGAGSEAGSYAGSR--- 682
Query: 501 SQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNHQQGF 547
+ SGS G GG G G + G H +G+
Sbjct: 683 ------AGSGSGRSQGRGGGGGGGGGGGGGGGSGYGHGEGYGHGEGY 805
Score = 35.0 bits (79), Expect = 0.26
Identities = 28/101 (27%), Positives = 40/101 (38%), Gaps = 10/101 (9%)
Frame = +2
Query: 437 AGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNY--- 493
+G+ G GS + +GS S S S + + S G G GS +GS+ ++AG+
Sbjct: 527 SGSGAGAGAGSGSGSGSGSSSSSASSSASSSSSSGSGGSGA-GSEAGSYAGSRAGSGSGR 703
Query: 494 -------GGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVG 527
GG G G G G YG +G G
Sbjct: 704 SQGRGGGGGGGG------GGGGGGGSGYGHGEGYGHGEGYG 808
Score = 31.2 bits (69), Expect = 3.7
Identities = 23/84 (27%), Positives = 31/84 (36%)
Frame = +2
Query: 417 TSTSTSSFNDDNSLYSEYSQAGNHVSQGVGSQAVNGSWSGSHGVSQAGNYDGSHGVGSQA 476
+ +S+SS + S S G+ GS A + + SGS G G G G
Sbjct: 575 SGSSSSSASSSASSSSSSGSGGSGAGSEAGSYAGSRAGSGS------GRSQGRGGGGGGG 736
Query: 477 VNGSWSGSHGVNQAGNYGGSQGVG 500
G G G YG +G G
Sbjct: 737 GGGGGGGGSGYGHGEGYGHGEGYG 808
>TC215791 similar to GB|CAE76446.1|38567152|BX842633 related to peroxisomal
membrane protein pex13 {Neurospora crassa;} , partial
(9%)
Length = 1440
Score = 40.4 bits (93), Expect = 0.006
Identities = 27/103 (26%), Positives = 40/103 (38%), Gaps = 3/103 (2%)
Frame = +3
Query: 481 WSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQG---VGSQAVNGSWSG 537
W ++G + G YG + S +G + S+G G YG S G G+ G + G
Sbjct: 339 WEQNYGNSTYGGYGSTMNYNSGYGSGMYGSSYGGLGGGMYGSSYGGGMYGNSMYRGGYGG 518
Query: 538 SHGVNHQQGFDMYATEASTKIGNNTASSGNQQVHHSYGNQQVN 580
+G + G MY + +G G YG Q N
Sbjct: 519 MYGSSGMYGGGMYNSGLGGPMGGYGMGGG------PYGEQDPN 629
Score = 39.3 bits (90), Expect = 0.014
Identities = 43/172 (25%), Positives = 66/172 (38%), Gaps = 17/172 (9%)
Frame = +3
Query: 383 YPDWYYDTIAQEWRSLATYNSSVQSSVH--GLQNGHTSTSTSSFNDDNSLY--SEYSQAG 438
Y W + TI E ++ NS G +G S + + + S ++ G
Sbjct: 84 YIIWCFSTIPMEPKTQPQANSPPPKPWEQAGSSSGPAPFKPPSAGNTSDVVEASGTAKPG 263
Query: 439 NHVSQGVGSQAVNGS----------WSGSHGVSQAGNYDGSHGVGSQAVNGSWSGSHGVN 488
VS S AVN + W ++G S G Y + S +G + S+G
Sbjct: 264 EIVSSSDRSAAVNRNTLGRPVPTRPWEQNYGNSTYGGYGSTMNYNSGYGSGMYGSSYGGL 443
Query: 489 QAGNYGGSQG---VGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSG 537
G YG S G G+ G + G +G +G YGG G+ + + G G
Sbjct: 444 GGGMYGSSYGGGMYGNSMYRGGYGGMYG--SSGMYGG--GMYNSGLGGPMGG 587
>TC219729 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (7%)
Length = 785
Score = 39.7 bits (91), Expect = 0.010
Identities = 22/80 (27%), Positives = 32/80 (39%), Gaps = 2/80 (2%)
Frame = -2
Query: 458 HGVSQAGNYDGSHGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAV--NGSWSGSHGVN 515
HG G + + G G + +G G H + G GG+ G G + G G H +
Sbjct: 364 HGGFGGGGHSTTTGGGGGSGSGFGGGGHLITGGGGDGGNGGGGDGGIGGGGDGGGGHSIG 185
Query: 516 QAGNYGGSQGVGSQAVNGSW 535
G GG S + G+W
Sbjct: 184 LGGGGGGGSLHSSHIMRGTW 125
Score = 31.2 bits (69), Expect = 3.7
Identities = 16/56 (28%), Positives = 23/56 (40%)
Frame = -2
Query: 485 HGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGVGSQAVNGSWSGSHG 540
HG G + + G G + +G G H + G GG+ G G + G G G
Sbjct: 364 HGGFGGGGHSTTTGGGGGSGSGFGGGGHLITGGGGDGGNGGGGDGGIGGGGDGGGG 197
>TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%)
Length = 1270
Score = 38.9 bits (89), Expect = 0.018
Identities = 21/61 (34%), Positives = 29/61 (47%)
Frame = -2
Query: 39 GNDEGNDSDDANAFANLSISDVDAGAFDNSVVGESGVEVKEELGVVKSDVGLDGGGDDDG 98
G DEG + + L DVD A + V + G E + V DVG DG GD++G
Sbjct: 879 GGDEGQEEGGDDGAEELG-DDVDDAAEEGDVAADEGAEGDGGVDVAAGDVGADGDGDEEG 703
Query: 99 K 99
+
Sbjct: 702 E 700
>TC228708 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall structural
protein 1.8 precursor (GRP 1.8), partial (15%)
Length = 494
Score = 38.1 bits (87), Expect = 0.030
Identities = 23/61 (37%), Positives = 28/61 (45%), Gaps = 1/61 (1%)
Frame = +3
Query: 468 GSHGVGSQAVNGSWSGS-HGVNQAGNYGGSQGVGSQAVNGSWSGSHGVNQAGNYGGSQGV 526
G HG G GS GS G G +GG G G+ A G G+ G + G +GG G
Sbjct: 15 GEHGXGYGGAGGSGGGSGGGYGDGGAHGGGYGGGAGAGGGGGYGAGGAH-GGGFGGGGGS 191
Query: 527 G 527
G
Sbjct: 192 G 194
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.128 0.370
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 64,085,628
Number of Sequences: 63676
Number of extensions: 918965
Number of successful extensions: 6149
Number of sequences better than 10.0: 143
Number of HSP's better than 10.0 without gapping: 5296
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5974
length of query: 1555
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1445
effective length of database: 5,635,272
effective search space: 8142968040
effective search space used: 8142968040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 65 (29.6 bits)
Medicago: description of AC137077.1


