

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.7 + phase: 0
(123 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
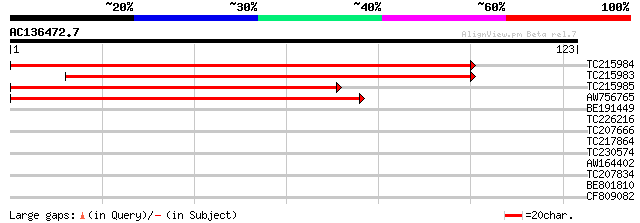
Score E
Sequences producing significant alignments: (bits) Value
TC215984 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase (Dihy... 197 1e-51
TC215983 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase (Dihy... 168 7e-43
TC215985 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase (Dihy... 134 8e-33
AW756765 weakly similar to GP|9759387|dbj dihydropyrimidinase {A... 80 2e-16
BE191449 38 0.001
TC226216 homologue to UP|Q6S4R9 (Q6S4R9) Allantoinase , partial... 31 0.12
TC207666 similar to UP|Q9ZSP8 (Q9ZSP8) Latex-abundant protein, p... 26 4.0
TC217864 similar to GB|AAP12861.1|30017255|BT006212 At2g40095 {A... 26 4.0
TC230574 26 5.2
AW164402 26 5.2
TC207834 similar to UP|Q8H2C2 (Q8H2C2) Serine/threonine kinase, ... 25 6.8
BE801810 similar to GP|15450451|gb| AT4g02580/T10P11_14 {Arabido... 25 8.9
CF809082 25 8.9
>TC215984 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase
(Dihydropyrimidine amidohydrolase) , partial (90%)
Length = 1922
Score = 197 bits (501), Expect = 1e-51
Identities = 94/101 (93%), Positives = 98/101 (96%)
Frame = +2
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MSIDAMEE+AKARKSGQRVIGEPV SGLALD+SWLWHPDF+TAAKYVMSPPIRK+GHDKA
Sbjct: 854 MSIDAMEEIAKARKSGQRVIGEPVASGLALDESWLWHPDFETAAKYVMSPPIRKRGHDKA 1033
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
LQAALSTGVLQLVGTDHC FNSTQKA GIDDFRKIPNGVNG
Sbjct: 1034LQAALSTGVLQLVGTDHCAFNSTQKALGIDDFRKIPNGVNG 1156
>TC215983 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase
(Dihydropyrimidine amidohydrolase) , partial (43%)
Length = 938
Score = 168 bits (425), Expect = 7e-43
Identities = 78/89 (87%), Positives = 84/89 (93%)
Frame = +1
Query: 13 RKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAALSTGVLQL 72
RK+GQRVIGEP+ GLALD+SWLWHPDF+ AAKYVMSPPIRK+GHDKALQAALSTGVLQL
Sbjct: 1 RKAGQRVIGEPIAYGLALDESWLWHPDFEIAAKYVMSPPIRKRGHDKALQAALSTGVLQL 180
Query: 73 VGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
VGTDHC FNSTQKA GIDDFRK+PNGVNG
Sbjct: 181 VGTDHCAFNSTQKARGIDDFRKMPNGVNG 267
>TC215985 similar to UP|Q9FMP3 (Q9FMP3) Dihydropyrimidinase
(Dihydropyrimidine amidohydrolase) , partial (31%)
Length = 821
Score = 134 bits (338), Expect = 8e-33
Identities = 63/72 (87%), Positives = 70/72 (96%)
Frame = +3
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MSIDAMEE+AKARK+GQRVIGEP+ SGLALD+SWLWHPDF+ AAKYVMSPPIRK+GHDKA
Sbjct: 168 MSIDAMEEIAKARKAGQRVIGEPIASGLALDESWLWHPDFEIAAKYVMSPPIRKRGHDKA 347
Query: 61 LQAALSTGVLQL 72
LQAALSTGVLQ+
Sbjct: 348 LQAALSTGVLQV 383
>AW756765 weakly similar to GP|9759387|dbj dihydropyrimidinase {Arabidopsis
thaliana}, partial (20%)
Length = 421
Score = 80.1 bits (196), Expect = 2e-16
Identities = 37/77 (48%), Positives = 57/77 (73%)
Frame = +2
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MSID M+++ KA+K Q+VIG+P+ S L L+ LWHP+F+ +Y++S P+RK+ H ++
Sbjct: 176 MSIDTMKKITKAQK*RQKVIGKPITSRLTLNKC*LWHPNFNITTEYIISRPMRKRRHYES 355
Query: 61 LQAALSTGVLQLVGTDH 77
LQA L+T VL+L+ T H
Sbjct: 356 LQATLATRVLRLIKTHH 406
>BE191449
Length = 538
Score = 38.1 bits (87), Expect = 0.001
Identities = 21/54 (38%), Positives = 31/54 (56%)
Frame = +2
Query: 4 DAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGH 57
+A+ +++ ARKS ++I PV S L S L H F+T YVM+PP + H
Sbjct: 371 NALTQISHARKSLLKLIIYPVASHYTLYQSKLIHAYFNTTTNYVMTPPHMRLKH 532
>TC226216 homologue to UP|Q6S4R9 (Q6S4R9) Allantoinase , partial (61%)
Length = 1367
Score = 31.2 bits (69), Expect = 0.12
Identities = 27/102 (26%), Positives = 44/102 (42%)
Frame = +2
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
+++ + +A+ G + E LA + P+ DT ++ SPPIR + L A
Sbjct: 419 SLDLIKEAKSHGDSISVETCPHYLAFTSEEI--PNGDT--RFKCSPPIRDAYNKDKLWEA 586
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNL 106
+ G + L+ TDH K DF K G++ NL
Sbjct: 587 VLEGHIDLLSTDHSPTVPELKLMEEGDFLKAWGGISSLQFNL 712
>TC207666 similar to UP|Q9ZSP8 (Q9ZSP8) Latex-abundant protein, partial (41%)
Length = 747
Score = 26.2 bits (56), Expect = 4.0
Identities = 16/48 (33%), Positives = 20/48 (41%)
Frame = +3
Query: 52 IRKQGHDKALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGV 99
+ GH L A TG G D C+ S DDFR+ +GV
Sbjct: 252 VHYSGHGTRLPA--ETGEDDDTGYDECIVPSDMNLITDDDFREFVDGV 389
>TC217864 similar to GB|AAP12861.1|30017255|BT006212 At2g40095 {Arabidopsis
thaliana;} , partial (59%)
Length = 1220
Score = 26.2 bits (56), Expect = 4.0
Identities = 14/44 (31%), Positives = 24/44 (53%)
Frame = -1
Query: 54 KQGHDKALQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPN 97
K+G D+ AA+ TG++ + D C+ TQ + +R +PN
Sbjct: 137 KEGADQG--AAILTGLVHSLSDDKCLGLYTQTPLWVMYYRGVPN 12
>TC230574
Length = 848
Score = 25.8 bits (55), Expect = 5.2
Identities = 12/36 (33%), Positives = 16/36 (44%)
Frame = +3
Query: 10 AKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAK 45
A RK G ++G P G + W+ DFD K
Sbjct: 372 AIVRKKGLGMVGRPCYHGSSSSMEWVKRQDFDELRK 479
>AW164402
Length = 427
Score = 25.8 bits (55), Expect = 5.2
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +3
Query: 38 PDFDTAAKYVMSPPIRKQGHDKALQAALSTGV 69
PD+ T+ +SPP R+ HD Q AL V
Sbjct: 3 PDYGTSDLQDISPPRRRGRHDSPSQDALHGSV 98
>TC207834 similar to UP|Q8H2C2 (Q8H2C2) Serine/threonine kinase, partial
(42%)
Length = 1169
Score = 25.4 bits (54), Expect = 6.8
Identities = 8/21 (38%), Positives = 12/21 (57%)
Frame = -2
Query: 34 WLWHPDFDTAAKYVMSPPIRK 54
W+W DF + Y+ P+RK
Sbjct: 130 WVWIQDFRQQSAYIRCKPVRK 68
>BE801810 similar to GP|15450451|gb| AT4g02580/T10P11_14 {Arabidopsis
thaliana}, partial (25%)
Length = 283
Score = 25.0 bits (53), Expect = 8.9
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +2
Query: 33 SWLWHPDFDTAAKYVMSPPIRK 54
SWL + D AK V PPIR+
Sbjct: 137 SWLPVSEMDAVAKIVKVPPIRR 202
>CF809082
Length = 730
Score = 25.0 bits (53), Expect = 8.9
Identities = 10/16 (62%), Positives = 12/16 (74%)
Frame = -2
Query: 40 FDTAAKYVMSPPIRKQ 55
FDTA K V+ PP RK+
Sbjct: 387 FDTAIKVVLQPPRRKE 340
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,011,299
Number of Sequences: 63676
Number of extensions: 57458
Number of successful extensions: 270
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 270
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 270
length of query: 123
length of database: 12,639,632
effective HSP length: 85
effective length of query: 38
effective length of database: 7,227,172
effective search space: 274632536
effective search space used: 274632536
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC136472.7


