

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130810.2 - phase: 0 /pseudo
(326 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
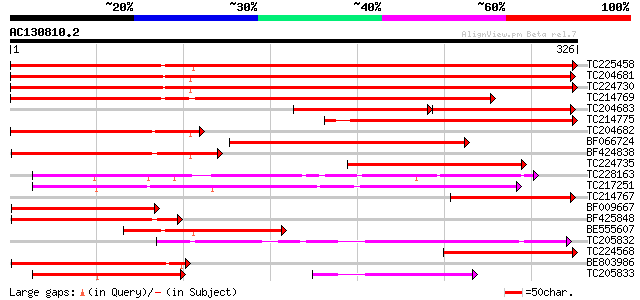
Score E
Sequences producing significant alignments: (bits) Value
TC225458 UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37, complete 633 0.0
TC204681 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein... 629 0.0
TC224730 similar to UP|O24074 (O24074) DnaJ-like protein MsJ1, p... 520 e-148
TC214769 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein... 493 e-140
TC204683 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein... 162 7e-82
TC214775 UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37, partia... 260 7e-70
TC204682 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein... 215 2e-56
BF066724 204 3e-53
BF424838 191 3e-49
TC224735 similar to UP|O24074 (O24074) DnaJ-like protein MsJ1, p... 171 5e-43
TC228163 similar to UP|Q8LEU4 (Q8LEU4) DnaJ protein-like, partia... 158 3e-39
TC217251 weakly similar to UP|DJB1_HUMAN (P25685) DnaJ homolog s... 155 2e-38
TC214767 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein... 146 1e-35
BF009667 similar to GP|5802244|gb|A seed maturation protein PM37... 143 1e-34
BF425848 similar to GP|5802244|gb|A seed maturation protein PM37... 119 1e-27
BE555607 similar to GP|5802244|gb|A seed maturation protein PM37... 118 3e-27
TC205832 similar to GB|AAM78044.1|21928031|AY125534 At3g62600/F2... 116 2e-26
TC224568 weakly similar to UP|Q9ZWK3 (Q9ZWK3) DnaJ homolog, part... 115 2e-26
BE803986 similar to PIR|T43929|T439 DnaJ protein homolog [import... 112 2e-25
TC205833 similar to GB|AAM78044.1|21928031|AY125534 At3g62600/F2... 96 3e-20
>TC225458 UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37, complete
Length = 1797
Score = 633 bits (1632), Expect = 0.0
Identities = 312/330 (94%), Positives = 317/330 (95%), Gaps = 4/330 (1%)
Frame = +2
Query: 1 MFGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 60
MFGRAPKKSD+TRYYEILGVSK ASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV
Sbjct: 116 MFGRAPKKSDNTRYYEILGVSKNASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 295
Query: 61 LSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGG----GGSSRGRRQRRG 116
LSDPEKREIYD YGEDALKEGMGGGGG HDPFDIFSSFFGGG GGSSRGRRQRRG
Sbjct: 296 LSDPEKREIYDQYGEDALKEGMGGGGG--HDPFDIFSSFFGGGSPFGSGGSSRGRRQRRG 469
Query: 117 EDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRH 176
EDVVHPLKVSLEDLYLGTSKKLSLSRNV+CSKCSGKGSKSGASMKCAGCQGTGMKVSIRH
Sbjct: 470 EDVVHPLKVSLEDLYLGTSKKLSLSRNVICSKCSGKGSKSGASMKCAGCQGTGMKVSIRH 649
Query: 177 LGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITF 236
LGPSMIQQMQH CNECKGTGETIND+DRCPQCKGEKVVQEKKVLEV VEKGMQN QKITF
Sbjct: 650 LGPSMIQQMQHACNECKGTGETINDRDRCPQCKGEKVVQEKKVLEVIVEKGMQNGQKITF 829
Query: 237 PGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQ 296
PGEADEAPDT+TGDIVFVLQQKEHPKFKRK+EDLFVEHTLSLTEALCGFQFVLTHLD RQ
Sbjct: 830 PGEADEAPDTITGDIVFVLQQKEHPKFKRKAEDLFVEHTLSLTEALCGFQFVLTHLDSRQ 1009
Query: 297 LLIKSNPGEVVKPDSYKAINDEGMPMYQRP 326
LLIKSNPGEVVKPDSYKAINDEGMPMYQRP
Sbjct: 1010LLIKSNPGEVVKPDSYKAINDEGMPMYQRP 1099
>TC204681 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
complete
Length = 1542
Score = 629 bits (1621), Expect = 0.0
Identities = 311/328 (94%), Positives = 315/328 (95%), Gaps = 3/328 (0%)
Frame = +3
Query: 1 MFGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 60
MFGRAPKKSDSTRYYEILGVSK AS DDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV
Sbjct: 111 MFGRAPKKSDSTRYYEILGVSKNASPDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 290
Query: 61 LSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGG---GGGGSSRGRRQRRGE 117
LSDPEKREIYDTYGEDALKEGMGGGGGG HDPFDIFSSFFGG G GGSSRGRRQRRGE
Sbjct: 291 LSDPEKREIYDTYGEDALKEGMGGGGGG-HDPFDIFSSFFGGSPFGSGGSSRGRRQRRGE 467
Query: 118 DVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHL 177
DVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKC+GKGSKSGASM CAGCQGTGMKVSIRHL
Sbjct: 468 DVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCNGKGSKSGASMTCAGCQGTGMKVSIRHL 647
Query: 178 GPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFP 237
GPSMIQQMQHPCNECKGTGETIND+DRC QCKGEKVVQEKKVLEV VEKGMQN QKITFP
Sbjct: 648 GPSMIQQMQHPCNECKGTGETINDRDRCQQCKGEKVVQEKKVLEVVVEKGMQNGQKITFP 827
Query: 238 GEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQL 297
GEADEAPDTVTGDIVFVLQQKEHPKFKRK++DLFVEHTLSLTEALCGFQFVL HLDGRQL
Sbjct: 828 GEADEAPDTVTGDIVFVLQQKEHPKFKRKADDLFVEHTLSLTEALCGFQFVLAHLDGRQL 1007
Query: 298 LIKSNPGEVVKPDSYKAINDEGMPMYQR 325
LIKSNPGEVVKPDSYKAINDEGMP YQR
Sbjct: 1008LIKSNPGEVVKPDSYKAINDEGMPNYQR 1091
>TC224730 similar to UP|O24074 (O24074) DnaJ-like protein MsJ1, partial (96%)
Length = 1913
Score = 520 bits (1338), Expect = e-148
Identities = 244/330 (73%), Positives = 287/330 (86%), Gaps = 4/330 (1%)
Frame = +1
Query: 1 MFGRA-PKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYE 59
MFGR P++SD+++YY+ILG+SK AS+D++KKAY+KAA+KNHPDKGGDPEKFKEL QAYE
Sbjct: 358 MFGRGGPRRSDNSKYYDILGISKNASEDEIKKAYRKAAMKNHPDKGGDPEKFKELGQAYE 537
Query: 60 VLSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGG---GGGGSSRGRRQRRG 116
VLSDPEK+E+YD YGEDALKEGMGGGG H+PFDIF SFFGG GGGGSSRGRRQ+ G
Sbjct: 538 VLSDPEKKELYDQYGEDALKEGMGGGGSF-HNPFDIFESFFGGASFGGGGSSRGRRQKHG 714
Query: 117 EDVVHPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRH 176
EDVVH LKVSLED+Y GT+KKLSLSRN+LC KC GKGSKSG + +C GC+GTGMK++ R
Sbjct: 715 EDVVHSLKVSLEDVYNGTTKKLSLSRNILCPKCKGKGSKSGTAGRCFGCKGTGMKITRRQ 894
Query: 177 LGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITF 236
+G MIQQMQH C +C+G+GE IN++D+CP CKG KV QEKKVLEVHVEKGMQ QKI F
Sbjct: 895 IGLGMIQQMQHVCPDCRGSGEVINERDKCPLCKGNKVSQEKKVLEVHVEKGMQQGQKIVF 1074
Query: 237 PGEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQ 296
G+ADEAPDT+TGDIVFVLQ K+HPKF+R+ +DL+++H LSLTEALCGFQF + HLDGRQ
Sbjct: 1075EGQADEAPDTITGDIVFVLQVKDHPKFRREQDDLYIDHNLSLTEALCGFQFAVKHLDGRQ 1254
Query: 297 LLIKSNPGEVVKPDSYKAINDEGMPMYQRP 326
LLIKSNPGEV+KP YKAINDEGMP + RP
Sbjct: 1255LLIKSNPGEVIKPGQYKAINDEGMPQHNRP 1344
>TC214769 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (64%)
Length = 943
Score = 493 bits (1268), Expect = e-140
Identities = 243/279 (87%), Positives = 255/279 (91%)
Frame = +2
Query: 1 MFGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 60
MFGRAPKKSD+TRYYEILGVSK ASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV
Sbjct: 122 MFGRAPKKSDNTRYYEILGVSKNASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 301
Query: 61 LSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDVV 120
LSDPEKREIYD YGEDALKEGMGGGGG HDPFDIFSSFFGGG SSRGRRQRRGEDVV
Sbjct: 302 LSDPEKREIYDQYGEDALKEGMGGGGG--HDPFDIFSSFFGGG---SSRGRRQRRGEDVV 466
Query: 121 HPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHLGPS 180
HPLKVSLEDLYLGTSKKLSLSRNV+CSKC+GKGSKSGASMKCAGCQGTGMKVSI+HLGPS
Sbjct: 467 HPLKVSLEDLYLGTSKKLSLSRNVICSKCTGKGSKSGASMKCAGCQGTGMKVSIKHLGPS 646
Query: 181 MIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEA 240
MIQQMQH CNECKGTGETIND+DRCPQCKGEKVVQEKKVLEV V GM N KITFP EA
Sbjct: 647 MIQQMQHACNECKGTGETINDRDRCPQCKGEKVVQEKKVLEVIVANGMHNGHKITFPDEA 826
Query: 241 DEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLT 279
D+ P T+TGDIV+VLQQKEH KF R+++DL +H LT
Sbjct: 827 DDTPYTITGDIVYVLQQKEHRKFTREADDLCAQHISVLT 943
>TC204683 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (60%)
Length = 1229
Score = 162 bits (410), Expect(2) = 7e-82
Identities = 76/80 (95%), Positives = 77/80 (96%)
Frame = +1
Query: 164 GCQGTGMKVSIRHLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVH 223
GCQGTGMKVSIRHLGPSMIQQMQHPCNECKGTGETIND+DRC QCKGEKVVQEKKVLEV
Sbjct: 1 GCQGTGMKVSIRHLGPSMIQQMQHPCNECKGTGETINDRDRCQQCKGEKVVQEKKVLEVV 180
Query: 224 VEKGMQNSQKITFPGEADEA 243
VEKGMQN QKITFPGEADEA
Sbjct: 181 VEKGMQNGQKITFPGEADEA 240
Score = 159 bits (403), Expect(2) = 7e-82
Identities = 76/82 (92%), Positives = 80/82 (96%)
Frame = +3
Query: 244 PDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNP 303
PDTVTGDIVFVLQQKEHPKFKRK++DLFVEHTLSLTEALCGFQFVLTHLD RQLLIKSNP
Sbjct: 315 PDTVTGDIVFVLQQKEHPKFKRKADDLFVEHTLSLTEALCGFQFVLTHLDSRQLLIKSNP 494
Query: 304 GEVVKPDSYKAINDEGMPMYQR 325
GEVVKP+S+KAINDEGMP YQR
Sbjct: 495 GEVVKPESFKAINDEGMPNYQR 560
>TC214775 UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37, partial (42%)
Length = 530
Score = 260 bits (664), Expect = 7e-70
Identities = 129/145 (88%), Positives = 133/145 (90%)
Frame = +3
Query: 182 IQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEAD 241
+Q MQ P GETIND+DRCPQCKGEKVVQEKKVLEV VEKGMQN QKITFPGEAD
Sbjct: 15 LQ*MQSP-------GETINDRDRCPQCKGEKVVQEKKVLEVIVEKGMQNGQKITFPGEAD 173
Query: 242 EAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKS 301
EAPDT+TGDIVFVLQQKEHPKFKRK+EDLFVEHTLSLTEALCGFQFVLTHLD RQLLIKS
Sbjct: 174 EAPDTITGDIVFVLQQKEHPKFKRKAEDLFVEHTLSLTEALCGFQFVLTHLDSRQLLIKS 353
Query: 302 NPGEVVKPDSYKAINDEGMPMYQRP 326
NPGEVVKPDSYKAINDEGMPMYQRP
Sbjct: 354 NPGEVVKPDSYKAINDEGMPMYQRP 428
>TC204682 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (27%)
Length = 447
Score = 215 bits (548), Expect = 2e-56
Identities = 108/115 (93%), Positives = 108/115 (93%), Gaps = 3/115 (2%)
Frame = +2
Query: 1 MFGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 60
MFGRAPKKSDSTRYYEILGVSK AS DDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV
Sbjct: 104 MFGRAPKKSDSTRYYEILGVSKNASPDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEV 283
Query: 61 LSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGG---GGGGSSRGRR 112
LSDPEKREIYDTYGEDALKEGM GGGGGGHDPFDIFSSFFGG G GGSSRGRR
Sbjct: 284 LSDPEKREIYDTYGEDALKEGM-GGGGGGHDPFDIFSSFFGGSPFGSGGSSRGRR 445
>BF066724
Length = 415
Score = 204 bits (520), Expect = 3e-53
Identities = 101/138 (73%), Positives = 111/138 (80%)
Frame = +2
Query: 127 LEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHLGPSMIQQMQ 186
LEDLYLGTS KLSLSRNV+CSKCSGKGSKSGASMKCAGCQGTG+KVSIRHLGPSMIQQMQ
Sbjct: 2 LEDLYLGTSTKLSLSRNVICSKCSGKGSKSGASMKCAGCQGTGLKVSIRHLGPSMIQQMQ 181
Query: 187 HPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEAPDT 246
CNEC+GTGETI D DRCPQCKGE VVQEKKVLEV V+KGM IT E+DE P+
Sbjct: 182 LACNECEGTGETITD*DRCPQCKGENVVQEKKVLEVIVKKGMSTGATITCACESDETPEH 361
Query: 247 VTGDIVFVLQQKEHPKFK 264
+ + VL + P +K
Sbjct: 362 GLDNCIIVLDIQSFPDYK 415
>BF424838
Length = 370
Score = 191 bits (486), Expect = 3e-49
Identities = 96/125 (76%), Positives = 103/125 (81%), Gaps = 4/125 (3%)
Frame = +2
Query: 2 FGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEVL 61
FGRAP KSD+TRYYEILGVS ASQDDLKKAY+ AA+ NHPDKGGDPEKF+ELAQAYE L
Sbjct: 2 FGRAPTKSDNTRYYEILGVSNNASQDDLKKAYQIAALNNHPDKGGDPEKFQELAQAYEAL 181
Query: 62 SDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGG----GGGGSSRGRRQRRGE 117
SDP KREIYD +G+DAL EGM GGG GHDPFDIFSSFFGG G GS RG R RGE
Sbjct: 182 SDPVKREIYDQHGDDALTEGM--GGGRGHDPFDIFSSFFGGGSPFGSRGSRRGPRHMRGE 355
Query: 118 DVVHP 122
DV+HP
Sbjct: 356 DVLHP 370
>TC224735 similar to UP|O24074 (O24074) DnaJ-like protein MsJ1, partial (24%)
Length = 1154
Score = 171 bits (432), Expect = 5e-43
Identities = 77/103 (74%), Positives = 90/103 (86%)
Frame = +1
Query: 195 TGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEAPDTVTGDIVFV 254
TGE IN++D+CP CKG KV QEKKVLEVHVEKGMQ QKI F G+ADEAPDT+TGDIVFV
Sbjct: 844 TGEVINERDKCPLCKGNKVSQEKKVLEVHVEKGMQQGQKIVFEGQADEAPDTITGDIVFV 1023
Query: 255 LQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQL 297
LQ K+HPKF+R+ +DL+++H LSLTEALCGFQF + HLDGRQL
Sbjct: 1024LQVKDHPKFRREQDDLYIDHNLSLTEALCGFQFAVKHLDGRQL 1152
>TC228163 similar to UP|Q8LEU4 (Q8LEU4) DnaJ protein-like, partial (65%)
Length = 1660
Score = 158 bits (400), Expect = 3e-39
Identities = 107/307 (34%), Positives = 163/307 (52%), Gaps = 16/307 (5%)
Frame = +2
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGD----PEKFKELAQAYEVLSDPEKREI 69
YY+ILGVSK AS ++KKAY A K HPD D +KF+E++ AYEVL D E+R+
Sbjct: 401 YYDILGVSKNASSSEIKKAYYGLAKKLHPDTNKDDPQAEKKFQEVSIAYEVLKDEERRQQ 580
Query: 70 YDTYGEDAL--KEGMGGGGGGGHDPF-------DIFSSFFGGGGGGSSRGRRQRRGEDVV 120
YD G DA ++ G GG GG +PF D SFF G GEDV
Sbjct: 581 YDQLGHDAYVNQQSTGFGGEGGFNPFEQIFRDHDFVKSFFHENIG----------GEDVK 730
Query: 121 HPLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGASMK-CAGCQGTGMKVSIRHLGP 179
+++S + G +K L+ +VLC+ C G G G + C C+G+ +V G
Sbjct: 731 TFIELSFMEAVQGCTKTLTFQTDVLCNACGGSGVPPGTRPETCKPCKGS--RVLFVQAG- 901
Query: 180 SMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQ--KITFP 237
I +M+ C CKGTG+ ++ D C C+G K+V+ K +++ + G+ +++ K+
Sbjct: 902 --IFRMESTCGTCKGTGKIVS--DYCKSCRGAKIVKGMKSIKLDIMPGIDSNETIKVYRS 1069
Query: 238 GEADEAPDTVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQL 297
G AD D GD+ ++ +E P F+R+ D+ V+ LS+T+A+ G + L G +
Sbjct: 1070GGADPDGDQ-PGDLYVTIKVREDPVFRREGSDIHVDAVLSITQAILGGTIQVPTLTG-DV 1243
Query: 298 LIKSNPG 304
++K PG
Sbjct: 1244VLKVRPG 1264
>TC217251 weakly similar to UP|DJB1_HUMAN (P25685) DnaJ homolog subfamily B
member 1 (Heat shock 40 kDa protein 1) (Heat shock
protein 40) (HSP40) (DnaJ protein homolog 1) (HDJ-1),
partial (21%)
Length = 1669
Score = 155 bits (393), Expect = 2e-38
Identities = 99/289 (34%), Positives = 147/289 (50%), Gaps = 8/289 (2%)
Frame = +2
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDP---EKFKELAQAYEVLSDPEKREIY 70
YY +LGVS+ AS+ ++K AY+K A HPD +P +KFKE++ AYEVLSD EKR IY
Sbjct: 365 YYSVLGVSRNASKSEIKSAYRKLARNYHPDVNKEPGAEQKFKEISNAYEVLSDDEKRSIY 544
Query: 71 DTYGEDALKEGMGGGGGGGHDPFDIFSSFFGG-GGGGSSRGRRQRR--GEDVVHPLKVSL 127
D +GE LK G G G +PFD+F S F G G SRG GED + L ++
Sbjct: 545 DRFGEAGLK-GSAMGMGDFSNPFDLFESLFEGMNRGAGSRGSWNGAIDGEDEYYSLVLNF 721
Query: 128 EDLYLGTSKKLSLSRNVLCSKCSGKGSKSGAS-MKCAGCQGTGMKVSIRHLGPSMIQQMQ 186
++ G K++ +SR C C+G G+K G + +C+ C G G VS P I Q
Sbjct: 722 KEAVFGIEKEIEISRLESCGTCNGSGAKPGTTPSRCSTCGGQGRVVSSTRT-PLGIFQQS 898
Query: 187 HPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEA-PD 245
C+ C GTGE C C G+ +++ K + + V G+ + ++ E +
Sbjct: 899 MTCSSCNGTGEI---STPCNTCSGDGRLRKSKRISLKVPAGVDSGSRLRVRNEGNAGRRG 1069
Query: 246 TVTGDIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDG 294
GD+ V++ P KR ++ +S +A+ G + +DG
Sbjct: 1070GSPGDLFVVIEVIPDPVLKRDDTNILYTCKVSYIDAILGTTIKVPTVDG 1216
>TC214767 homologue to UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37,
partial (39%)
Length = 713
Score = 146 bits (368), Expect = 1e-35
Identities = 70/72 (97%), Positives = 71/72 (98%)
Frame = +2
Query: 254 VLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYK 313
VLQQKEHPKFKRK+EDLFVEH LSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYK
Sbjct: 2 VLQQKEHPKFKRKAEDLFVEHILSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYK 181
Query: 314 AINDEGMPMYQR 325
AINDEGMPMYQR
Sbjct: 182 AINDEGMPMYQR 217
>BF009667 similar to GP|5802244|gb|A seed maturation protein PM37 {Glycine
max}, partial (20%)
Length = 256
Score = 143 bits (360), Expect = 1e-34
Identities = 68/85 (80%), Positives = 75/85 (88%)
Frame = +2
Query: 2 FGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEVL 61
FGRAPKKSDSTRYYEIL VSK A+ DDLKKAY AA+ NHPDKGGDP+KF +LAQAY+VL
Sbjct: 2 FGRAPKKSDSTRYYEILWVSKNAAPDDLKKAYMIAAVNNHPDKGGDPDKFTQLAQAYDVL 181
Query: 62 SDPEKREIYDTYGEDALKEGMGGGG 86
S+PEKREIYDTYGE AL EG+G GG
Sbjct: 182SEPEKREIYDTYGEHALTEGLGVGG 256
>BF425848 similar to GP|5802244|gb|A seed maturation protein PM37 {Glycine
max}, partial (20%)
Length = 289
Score = 119 bits (299), Expect = 1e-27
Identities = 67/98 (68%), Positives = 69/98 (70%)
Frame = +2
Query: 2 FGRAPKKSDSTRYYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPEKFKELAQAYEVL 61
FGRAP K D T YYEIL V A QDDLKKAY KAAI NHPD GGDP KFKELAQAYEVL
Sbjct: 2 FGRAPNKIDYTMYYEILCVFNNAPQDDLKKAYIKAAIYNHPD*GGDPYKFKELAQAYEVL 181
Query: 62 SDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSF 99
DP EIYD +GE AL + M G DP DIFSSF
Sbjct: 182YDPVNLEIYDHFGEFALNDRM--VGRCSLDPRDIFSSF 289
>BE555607 similar to GP|5802244|gb|A seed maturation protein PM37 {Glycine
max}, partial (23%)
Length = 289
Score = 118 bits (296), Expect = 3e-27
Identities = 64/98 (65%), Positives = 71/98 (72%), Gaps = 4/98 (4%)
Frame = +2
Query: 66 KREIYDTYGEDALKEGMGGGGGGGHDPFDIFSSFFGGGG----GGSSRGRRQRRGEDVVH 121
KR+IYD +GEDAL EGMGGGGG HDPFDIF FFGGG G SS+ R+QR ED H
Sbjct: 2 KRDIYDQHGEDALNEGMGGGGG--HDPFDIF*FFFGGGSPFGSGASSQCRKQRLVEDAAH 175
Query: 122 PLKVSLEDLYLGTSKKLSLSRNVLCSKCSGKGSKSGAS 159
PL+ +EDLYL TSK L LS V+CSKCS GS SGAS
Sbjct: 176 PLQDFVEDLYLATSKNLYLSIIVVCSKCSVNGSNSGAS 289
>TC205832 similar to GB|AAM78044.1|21928031|AY125534 At3g62600/F26K9_30
{Arabidopsis thaliana;} , partial (68%)
Length = 1171
Score = 116 bits (290), Expect = 2e-26
Identities = 76/239 (31%), Positives = 118/239 (48%)
Frame = +1
Query: 85 GGGGGHDPFDIFSSFFGGGGGGSSRGRRQRRGEDVVHPLKVSLEDLYLGTSKKLSLSRNV 144
G GGG + DIFS+FFGGG + +G+D+V L +LEDLY+G + K+ +NV
Sbjct: 7 GRGGGMNFQDIFSTFFGGGP--MEEEEKIVKGDDLVVDLDATLEDLYMGGTLKVWREKNV 180
Query: 145 LCSKCSGKGSKSGASMKCAGCQGTGMKVSIRHLGPSMIQQMQHPCNECKGTGETINDKDR 204
L K + C+ +V + +GP M QQM
Sbjct: 181 L---------KPAPGKRRCNCRN---EVYHKQIGPGMFQQMTEQV--------------- 279
Query: 205 CPQCKGEKVVQEKKVLEVHVEKGMQNSQKITFPGEADEAPDTVTGDIVFVLQQKEHPKFK 264
C QC K V+E + V +EKGMQ+ Q++ F + + D +GD+ F ++ H F+
Sbjct: 280 CEQCPNVKYVREGYFITVDIEKGMQDGQEVLFYEDGEPIIDGESGDLRFRIRTAPHDVFR 459
Query: 265 RKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKPDSYKAINDEGMPMY 323
R+ DL T++L +AL GF+ + HLD + L+ + E+ KP + EGMP++
Sbjct: 460 REGNDLHSTVTITLVQALVGFEKTIKHLD--EHLVDISTKEITKPKQVRKFKGEGMPLH 630
>TC224568 weakly similar to UP|Q9ZWK3 (Q9ZWK3) DnaJ homolog, partial (38%)
Length = 784
Score = 115 bits (289), Expect = 2e-26
Identities = 51/77 (66%), Positives = 61/77 (78%)
Frame = +1
Query: 250 DIVFVLQQKEHPKFKRKSEDLFVEHTLSLTEALCGFQFVLTHLDGRQLLIKSNPGEVVKP 309
D FVLQ K+HP+F+R + F++ LSL EALCGFQF + HLDGRQLLIKSNPGEV+KP
Sbjct: 1 DXXFVLQVKDHPRFRRXXDXXFIDQNLSLXEALCGFQFAVKHLDGRQLLIKSNPGEVIKP 180
Query: 310 DSYKAINDEGMPMYQRP 326
YKA+NDEGMP + RP
Sbjct: 181 GQYKALNDEGMPQHNRP 231
>BE803986 similar to PIR|T43929|T439 DnaJ protein homolog [imported] - Salix
gilgiana, partial (15%)
Length = 328
Score = 112 bits (280), Expect = 2e-25
Identities = 61/110 (55%), Positives = 76/110 (68%), Gaps = 7/110 (6%)
Frame = +2
Query: 2 FGRA-PKKSDSTRYYEILGVSKTASQDDLKKA-YKKAAIKNHPDKGGDPEKFKELAQAYE 59
FGR P++SD +YY+ILG+S+ AS+D ++ YK + HP KGGDPE KEL QAYE
Sbjct: 2 FGRGGPRRSDIFKYYDILGISRNASEDRNQEGLYKGGY*RTHPYKGGDPE*LKELGQAYE 181
Query: 60 VLSDPEKREIYDTYGEDALKEGMGGGGGGGHDPFDIFS-----SFFGGGG 104
LSDP+K E+YD YGEDALK+GMGGG + P DIF + FGGGG
Sbjct: 182 ALSDPDKNELYDRYGEDALKQGMGGGCSLRY-PLDIFELLVGRACFGGGG 328
>TC205833 similar to GB|AAM78044.1|21928031|AY125534 At3g62600/F26K9_30
{Arabidopsis thaliana;} , partial (55%)
Length = 866
Score = 95.5 bits (236), Expect = 3e-20
Identities = 51/93 (54%), Positives = 64/93 (67%), Gaps = 5/93 (5%)
Frame = +1
Query: 14 YYEILGVSKTASQDDLKKAYKKAAIKNHPDKGGDPE----KFKELAQAYEVLSDPEKREI 69
YY+IL +SK AS + +K+AY+K A+K HPDK E KF E++ AYEVLSD EKR I
Sbjct: 283 YYDILQLSKGASDEQIKRAYRKLALKYHPDKNPGNEEANKKFAEISNAYEVLSDSEKRNI 462
Query: 70 YDTYGEDALKE-GMGGGGGGGHDPFDIFSSFFG 101
YD YGE+ LK+ GG GGG + DIFS+F G
Sbjct: 463 YDRYGEEGLKQHAASGGRGGGMNFQDIFSTFGG 561
Score = 50.8 bits (120), Expect = 8e-07
Identities = 28/95 (29%), Positives = 44/95 (45%)
Frame = +3
Query: 175 RHLGPSMIQQMQHPCNECKGTGETINDKDRCPQCKGEKVVQEKKVLEVHVEKGMQNSQKI 234
+ +GP M QQM C QC K V+E + V +EKGMQ+ Q++
Sbjct: 627 KQIGPGMFQQMTEQV---------------CEQCPNVKYVREGYFITVDIEKGMQDGQEV 761
Query: 235 TFPGEADEAPDTVTGDIVFVLQQKEHPKFKRKSED 269
F + + D +GD+ F ++ H F+R+ D
Sbjct: 762 LFYEDGEPIIDGESGDLRFRIRTAPHDVFRREGND 866
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.135 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,527,769
Number of Sequences: 63676
Number of extensions: 199107
Number of successful extensions: 3957
Number of sequences better than 10.0: 334
Number of HSP's better than 10.0 without gapping: 2132
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2961
length of query: 326
length of database: 12,639,632
effective HSP length: 98
effective length of query: 228
effective length of database: 6,399,384
effective search space: 1459059552
effective search space used: 1459059552
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC130810.2


