

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126784.7 - phase: 0
(339 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
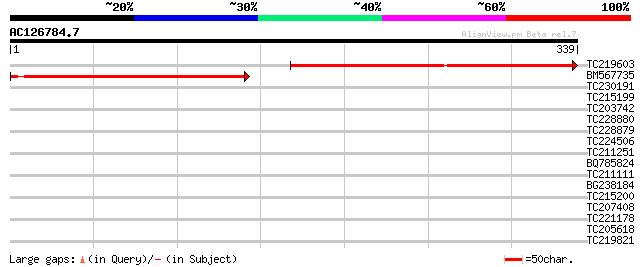
Score E
Sequences producing significant alignments: (bits) Value
TC219603 320 8e-88
BM567735 191 3e-49
TC230191 weakly similar to GB|AAF44683.1|7271905|AF239765 nuclea... 31 0.70
TC215199 similar to UP|Q84PC7 (Q84PC7) PP2A regulatory subunit-l... 31 0.70
TC203742 similar to UP|Q8H6S5 (Q8H6S5) CTV.2, partial (15%) 31 0.92
TC228880 similar to UP|Q6NPN9 (Q6NPN9) At1g76260, partial (43%) 30 1.2
TC228879 similar to UP|Q6NPN9 (Q6NPN9) At1g76260, partial (30%) 30 1.2
TC224506 weakly similar to UP|GCP2_ARATH (Q9M1S8) Probable gluta... 30 1.6
TC211251 weakly similar to UP|Q6TAF9 (Q6TAF9) Blight resistance ... 30 1.6
BQ785824 29 2.7
TC211111 homologue to UP|EBN2_EBV (P12978) EBNA-2 nuclear protei... 29 2.7
BG238184 29 2.7
TC215200 similar to UP|Q9XHD0 (Q9XHD0) PP2A regulatory subunit, ... 28 4.5
TC207408 homologue to UP|Q6T2Z2 (Q6T2Z2) Cyclin-dependent kinase... 28 5.9
TC221178 similar to UP|PERX_NICSY (Q02200) Lignin forming anioni... 28 7.7
TC205618 similar to GB|AAL60196.1|18139887|AF441079 O-linked N-a... 28 7.7
TC219821 weakly similar to UP|PJA1_MOUSE (O55176) Ubiquitin prot... 28 7.7
>TC219603
Length = 1210
Score = 320 bits (819), Expect = 8e-88
Identities = 154/171 (90%), Positives = 157/171 (91%)
Frame = +3
Query: 169 TSVKWASPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHT 228
T+V WASP+EFATGGYGFGLHWWDQRKPGGPVSQFKGNWDK L SGIVHSIDIHPSRKHT
Sbjct: 3 TAVSWASPMEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKKLTSGIVHSIDIHPSRKHT 182
Query: 229 CLAAGSLGTVLAWDLRMQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTS 288
CLA GSLGTV AWDLR QQQPIILSG GAG G GNTAVQSISESEVWEVQYDRCIK NTS
Sbjct: 183 CLAGGSLGTVFAWDLRWQQQPIILSGAGAGAG-GNTAVQSISESEVWEVQYDRCIKPNTS 359
Query: 289 STRILPAMICSEDGILGVIEQGEEPIELLAEPCAINSFDIDRHNPLVSGCS 339
STRILP+MICSEDGILGVIEQGEEPIELLAEPCAINSFDIDRHNP CS
Sbjct: 360 STRILPSMICSEDGILGVIEQGEEPIELLAEPCAINSFDIDRHNPSDVICS 512
>BM567735
Length = 428
Score = 191 bits (486), Expect = 3e-49
Identities = 96/143 (67%), Positives = 114/143 (79%)
Frame = +1
Query: 1 MAVTANSEIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSF 60
MA TA I RFPQ+KY+DAVRWLP+ SAF+RFAVLA DFD + SSIEIHS PNPLS
Sbjct: 7 MAATA---IQRFPQHKYVDAVRWLPLQSAFDRFAVLALFDFDFDASSIEIHSLNPNPLSL 177
Query: 61 EFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHL 120
E SSW+SPS +SS+K+S FLQK +I +STSSGSLHFLF+DS+ A L SEVS+PE LH
Sbjct: 178 EPHSSWSSPSRVSSLKASHFLQKPLIVASTSSGSLHFLFSDSSVATLESEVSIPEGALHS 357
Query: 121 EGKCCIDLMDGGVECVTVGDDGR 143
C+DL+DGGVECVTVG+DG+
Sbjct: 358 VPASCVDLLDGGVECVTVGEDGK 426
>TC230191 weakly similar to GB|AAF44683.1|7271905|AF239765 nuclear protein
Ytm1p {Mus musculus;} , partial (6%)
Length = 1106
Score = 31.2 bits (69), Expect = 0.70
Identities = 22/67 (32%), Positives = 29/67 (42%), Gaps = 2/67 (2%)
Frame = +2
Query: 180 ATGGYGFGLHWWDQRKPG--GPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGT 237
A GG + WD RKPG PV QF ++ + + H L+A G
Sbjct: 308 AAGGSDPVIRIWDPRKPGTSAPVFQFSS------HTSWISACKWHDQSWFHLLSASYDGK 469
Query: 238 VLAWDLR 244
V+ WDLR
Sbjct: 470 VMLWDLR 490
>TC215199 similar to UP|Q84PC7 (Q84PC7) PP2A regulatory subunit-like protein,
partial (16%)
Length = 782
Score = 31.2 bits (69), Expect = 0.70
Identities = 21/53 (39%), Positives = 27/53 (50%)
Frame = -1
Query: 68 SPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHL 120
+P +SS +SSQ L S ASS+SS S F F L S +S+ HL
Sbjct: 257 APRGLSSFQSSQALAFSCAASSSSSSSPGFDFLSLCQDELASSMSLTFLSCHL 99
>TC203742 similar to UP|Q8H6S5 (Q8H6S5) CTV.2, partial (15%)
Length = 903
Score = 30.8 bits (68), Expect = 0.92
Identities = 25/100 (25%), Positives = 40/100 (40%), Gaps = 6/100 (6%)
Frame = +3
Query: 39 SDFDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASS------TSS 92
+ F LS +H F+P+ E + W P PI + +S S+ ASS + S
Sbjct: 393 NQFAVGLSDGGVHVFEPH----ESEGKWGVPPPIENGSTSNMAATSVGASSDEAQR*SMS 560
Query: 93 GSLHFLFADSTDARLVSEVSVPENELHLEGKCCIDLMDGG 132
FL + V + +N + L G C + + G
Sbjct: 561 AKDKFLLHTWLAGHFKN*VCI*DNAIVLSGLCAVTYQNNG 680
>TC228880 similar to UP|Q6NPN9 (Q6NPN9) At1g76260, partial (43%)
Length = 761
Score = 30.4 bits (67), Expect = 1.2
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = +3
Query: 216 VHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQPI 250
V S+D HP ++H + A + WDLR + PI
Sbjct: 99 VCSVDYHPQKQHILVTAEHESGIHIWDLRKPKVPI 203
Score = 30.4 bits (67), Expect = 1.2
Identities = 17/61 (27%), Positives = 28/61 (45%)
Frame = +3
Query: 181 TGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLA 240
T + G+H WD RKP P+ + G ++ ++ +P L+AG+ TV
Sbjct: 144 TAEHESGIHIWDLRKPKVPIQELPG------HTHWTWTVKCNPEYDGMILSAGTDSTVNL 305
Query: 241 W 241
W
Sbjct: 306 W 308
>TC228879 similar to UP|Q6NPN9 (Q6NPN9) At1g76260, partial (30%)
Length = 742
Score = 30.4 bits (67), Expect = 1.2
Identities = 17/61 (27%), Positives = 28/61 (45%)
Frame = +3
Query: 181 TGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLA 240
T + G+H WD RKP P+ + G ++ ++ +P L+AG+ TV
Sbjct: 87 TAEHESGIHIWDLRKPKVPIQELPG------HTHWTWTVKCNPEYDGMILSAGTDSTVNL 248
Query: 241 W 241
W
Sbjct: 249 W 251
Score = 28.1 bits (61), Expect = 5.9
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = +3
Query: 216 VHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQPI 250
V S+D HP ++ + A + WDLR + PI
Sbjct: 42 VXSVDFHPQKQXMLVTAEHESGIHIWDLRKPKVPI 146
>TC224506 weakly similar to UP|GCP2_ARATH (Q9M1S8) Probable glutamate
carboxypeptidase II , partial (15%)
Length = 515
Score = 30.0 bits (66), Expect = 1.6
Identities = 17/46 (36%), Positives = 24/46 (51%)
Frame = -2
Query: 67 TSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVS 112
TS P+S+ S F + I+ S S HF F DS+ A L +E +
Sbjct: 313 TSKKPLSASISLSFNARIFISIERSEASCHFNFFDSSLASLAAEAN 176
>TC211251 weakly similar to UP|Q6TAF9 (Q6TAF9) Blight resistance protein SH10
(Fragment), partial (8%)
Length = 1062
Score = 30.0 bits (66), Expect = 1.6
Identities = 20/65 (30%), Positives = 32/65 (48%), Gaps = 3/65 (4%)
Frame = -3
Query: 56 NPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASST---SSGSLHFLFADSTDARLVSEVS 112
N SF F+ + SP P+++ S F+ + + SSG L TD+ L + S
Sbjct: 631 NVTSFNFKLKF*SPWPLNNSNLSHFITCKVSNAGKCLLSSGKDFSLLQHQTDSSLRTGNS 452
Query: 113 VPENE 117
+PEN+
Sbjct: 451 IPENK 437
>BQ785824
Length = 426
Score = 29.3 bits (64), Expect = 2.7
Identities = 19/65 (29%), Positives = 27/65 (41%)
Frame = +1
Query: 27 LSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSII 86
+S+F +A D +L+ HS L F F SW P P S + S L
Sbjct: 64 VSSFLLAVAVAVVD*PHSLTHSLTHSLATTTLDFNFFHSWLRPPPPPSARGSSLLACQSR 243
Query: 87 ASSTS 91
A+ T+
Sbjct: 244 ATPTA 258
>TC211111 homologue to UP|EBN2_EBV (P12978) EBNA-2 nuclear protein, partial
(5%)
Length = 769
Score = 29.3 bits (64), Expect = 2.7
Identities = 18/51 (35%), Positives = 30/51 (58%), Gaps = 1/51 (1%)
Frame = +2
Query: 25 PVLSAFNRFAVLATSDFDSNLSSIEI-HSFKPNPLSFEFQSSWTSPSPISS 74
P+LS+F +++ + S + N +S I H+ PN L ++ S+ T P P SS
Sbjct: 305 PLLSSFLTYSLASFSPYSLNPTSPFIFHNTSPNLLCPQYASASTFPPPSSS 457
>BG238184
Length = 465
Score = 29.3 bits (64), Expect = 2.7
Identities = 14/39 (35%), Positives = 21/39 (52%)
Frame = +2
Query: 58 LSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLH 96
LSF+ Q+ W P S++S +++STS SLH
Sbjct: 35 LSFQSQTPWRFGDPSRSLRSFSRSSSMALSASTSPASLH 151
>TC215200 similar to UP|Q9XHD0 (Q9XHD0) PP2A regulatory subunit, partial (93%)
Length = 1530
Score = 28.5 bits (62), Expect = 4.5
Identities = 26/71 (36%), Positives = 33/71 (45%), Gaps = 2/71 (2%)
Frame = -1
Query: 52 SFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSL--HFLFADSTDARLVS 109
S P +SF F + P +SS +SSQ L S ASS+SS S F F L S
Sbjct: 1347 SVYPQGVSFLFPA----PRGLSSFQSSQALAFSCAASSSSSSSSSPSFDFLSLCQDELTS 1180
Query: 110 EVSVPENELHL 120
+S+ HL
Sbjct: 1179 SMSLTFLSCHL 1147
>TC207408 homologue to UP|Q6T2Z2 (Q6T2Z2) Cyclin-dependent kinase inhibitor
1;2, partial (97%)
Length = 1041
Score = 28.1 bits (61), Expect = 5.9
Identities = 15/57 (26%), Positives = 29/57 (50%)
Frame = +1
Query: 269 ISESEVWEVQYDRCIKSNTSSTRILPAMICSEDGILGVIEQGEEPIELLAEPCAINS 325
I + V E+ +RC+ +S +PA CS +G +G+ E + ++L E + +
Sbjct: 283 IIPATVTEIVQERCLSPTSSE---IPASCCSSNGSIGLDEDRIKLLDLEVESAQVET 444
>TC221178 similar to UP|PERX_NICSY (Q02200) Lignin forming anionic peroxidase
precursor , partial (61%)
Length = 681
Score = 27.7 bits (60), Expect = 7.7
Identities = 12/27 (44%), Positives = 17/27 (62%)
Frame = -1
Query: 49 EIHSFKPNPLSFEFQSSWTSPSPISSI 75
E+ SF+ N S +Q +W SPSP+ I
Sbjct: 450 EVPSFESNS*SSCYQMNWASPSPVLPI 370
>TC205618 similar to GB|AAL60196.1|18139887|AF441079 O-linked N-acetyl
glucosamine transferase {Arabidopsis thaliana;} ,
partial (15%)
Length = 714
Score = 27.7 bits (60), Expect = 7.7
Identities = 12/29 (41%), Positives = 17/29 (58%)
Frame = -3
Query: 100 ADSTDARLVSEVSVPENELHLEGKCCIDL 128
ADST ++S+ S P +EGK C+ L
Sbjct: 391 ADSTF*NMISQDSSPNGPCQIEGKSCVQL 305
>TC219821 weakly similar to UP|PJA1_MOUSE (O55176) Ubiquitin protein ligase
Praja1 , partial (4%)
Length = 1135
Score = 27.7 bits (60), Expect = 7.7
Identities = 17/50 (34%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Frame = -3
Query: 270 SESEVWEVQYDRCIKSNTSSTRILPAMICS---EDGILGVIEQGEEPIEL 316
+ES + E+Q CIK N R++ M CS D I +I P+ L
Sbjct: 545 TESLLVELQSRDCIKKNRIGFRVINMMFCSISILDSISYIIRDSTPPLSL 396
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,938,671
Number of Sequences: 63676
Number of extensions: 225590
Number of successful extensions: 1363
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1357
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1362
length of query: 339
length of database: 12,639,632
effective HSP length: 98
effective length of query: 241
effective length of database: 6,399,384
effective search space: 1542251544
effective search space used: 1542251544
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC126784.7


