

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
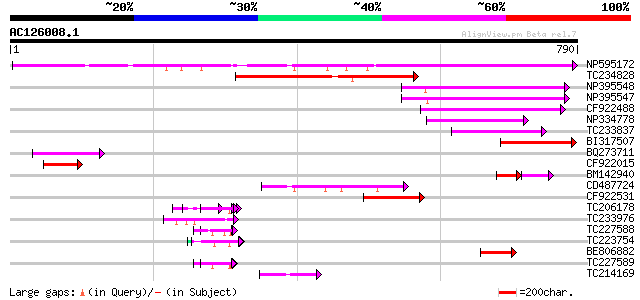
Query= AC126008.1 - phase: 0 /pseudo
(790 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 317 2e-86
TC234828 231 1e-60
NP395548 reverse transcriptase [Glycine max] 151 9e-37
NP395547 reverse transcriptase [Glycine max] 139 6e-33
CF922488 129 5e-30
NP334778 reverse transcriptase [Glycine max] 100 4e-21
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 97 4e-20
BI317507 92 1e-18
BQ273711 81 2e-15
CF922015 78 1e-14
BM142940 42 7e-09
CD487724 58 1e-08
CF922531 57 3e-08
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 56 7e-08
TC233976 55 2e-07
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 54 2e-07
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 51 2e-06
BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, pa... 51 2e-06
TC227589 50 5e-06
TC214169 similar to UP|Q8LF25 (Q8LF25) DNA-damage inducible prot... 49 9e-06
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 317 bits (811), Expect = 2e-86
Identities = 249/845 (29%), Positives = 401/845 (46%), Gaps = 58/845 (6%)
Frame = +1
Query: 4 VEMAGGRDDAAIAQALATMAQVLAQ-GNENTAANRRNEGEAEERRLDRFLRNNPPTFKGR 62
+E+ +D+ +Q A M+QVL + N +++ + + E++R +R+ F R
Sbjct: 136 IEVMQSTNDSQFSQLNAVMSQVLQRLQNIPMSSHGASNSQKEQQRSSFQVRSVKLDFP-R 312
Query: 63 FDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWWINAKGRLELGGDVVTWA 122
FD + W+ E+ F T D ++ +A+ ++ W+ L+ +W
Sbjct: 313 FDGKNVMDWIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWY----QMLQKTEPFSSWQ 480
Query: 123 KFKVEFLRKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEELSRFCPYINAEDAMVSKC 182
F + P + F L Q +V EY +F L +N D + ++
Sbjct: 481 AFTRALELDFGPSAYDCPRATLF-KLNQSA-TVNEYYMQFTAL------VNRVDGLSAEA 636
Query: 183 VK--FESGLRPVIYQYMCVQEIRDFDTLVHKCRMFDD-------------------AGKA 221
+ F SGL+ I + + E R V ++F++ + +
Sbjct: 637 ILDCFVSGLQEEISRDVKAMEPRTLTKAVALAKLFEEKYTSPPKTKTFSNLARNFTSNTS 816
Query: 222 KSNYYKVQNEKKGKGHG-----VGKPYSKDKGKKRESGGGSRPSLADVK-----CYRCGT 271
+ Y N+K + P +K + ++ P+ ++ CY C
Sbjct: 817 ATQKYPPTNQKNDNPKPNLPPLLPTPSTKPFNLRNQNIKKISPAEIQLRREKNLCYFCDE 996
Query: 272 LGHYANECMN-DLNCHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKAAGKVFAL 330
A++C N + + + D +V+ E + + H+S L
Sbjct: 997 KFSPAHKCPNRQVMLLQLEETDEDQTDEQVMVTEEANMD-DDTHHLS------------L 1137
Query: 331 NAKEVEQPDNLIRGMCFINSTPLIAIIDTGATHSFISASCVERLGLVVTPL--LRGMVID 388
NA IR + + ++D G++ +FI + L L V P LR +V
Sbjct: 1138NAMRGSNGVGTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVLV-- 1311
Query: 389 TPASGSG---TTSFMCAKCPVNFGDVDFKLDLVCLPLKNMDVIFGMDWLLAFGVSI---N 442
G+G + + + P++ + K+ + L + DVI G WL G +
Sbjct: 1312----GNGQILSAEGIVQQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYA 1479
Query: 443 CLT-KSVTFSKPVEELGREFLTAEQVK----KSLDGEACVFMMFASLKVSGEKGASVLP- 496
LT K K + G A Q + + L + FA + E L
Sbjct: 1480ALTLKFFQNDKFITLQGEGNSEATQAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKD 1659
Query: 497 -----------VVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGE 545
++ + +VF LP +RE + AI L G+ P+ + PYR ++ +
Sbjct: 1660LPTNIDPELAILLHTYAQVFAVP-ASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQ 1836
Query: 546 LKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD 605
++K ++E+L + I+PS SP P+LLVKKKDGS R C DYR LN +T+K+ +P+P +D+
Sbjct: 1837IEKMIQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDE 2016
Query: 606 *MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFME 665
+D+L GA+ FSK+DLRSGYHQI V+ ED KTAFRT +GHYE+ VMPFG++NAP F
Sbjct: 2017LLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQC 2196
Query: 666 YMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKE 725
MN+IF L +FV+VF DDIL+YS S ++H +HL VLQ LK++QL A+LSKC F E
Sbjct: 2197LMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTE 2376
Query: 726 VSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLALPLT 785
V +LGH +S G+S++ +KV AVL W +P +V +++ GL GYYRRFI+ ++ +A PLT
Sbjct: 2377VDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLT 2556
Query: 786 QLTQK 790
L QK
Sbjct: 2557DLLQK 2571
>TC234828
Length = 857
Score = 231 bits (588), Expect = 1e-60
Identities = 123/259 (47%), Positives = 161/259 (61%), Gaps = 4/259 (1%)
Frame = +3
Query: 315 HISTKPKKAAGKVFALNAKEVEQPDNLIRGMCFINSTPLIAIIDTGATHSFISASCVERL 374
+IS K +VFA++ E D+LIRG C I L + D+GATHSFIS +CVERL
Sbjct: 99 NISGGRPKVPSRVFAMSGSEAAASDDLIRGKCLIADKLLDVLYDSGATHSFISHACVERL 278
Query: 375 GLVVTPLLRGMVIDTPASGSGTTSFMCAKCPVNFGDVDFKLDLVCLPLKNMDVIFGMDWL 434
GL T L MV+ TP S TTS +C KCP+ F DL+CLPL ++DVI GMDWL
Sbjct: 279 GLCATELPYDMVVSTPTSEPVTTSRVCLKCPIIVEGRSFMADLICLPLAHLDVILGMDWL 458
Query: 435 LAFGVSINCLTKSVTFSKPVEELGREFLTAEQVKKSLDGEAC----VFMMFASLKVSGEK 490
+ ++C K + F G + + +E +K+ E +M+ S+ V +
Sbjct: 459 STNHIFLDCKEKMLVF-------GGDVVPSEPLKEDAANEETEDVRTYMVLFSMYVEEDA 617
Query: 491 GASVLPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQL 550
S +PVV EFPEVFP+D+ ELP EREVEF ID+VPG +P+SIAPYRMS EL E+K Q+
Sbjct: 618 EVSCIPVVSEFPEVFPDDVCELPPEREVEFIIDVVPGANPVSIAPYRMSPVELAEVKAQV 797
Query: 551 EELLEKQFIRPSVSPRGAP 569
++LL KQF+RPS SP GAP
Sbjct: 798 QDLLSKQFVRPSASPWGAP 854
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 151 bits (382), Expect = 9e-37
Identities = 92/254 (36%), Positives = 139/254 (54%), Gaps = 19/254 (7%)
Frame = +1
Query: 546 LKKQLEELLEKQFIRP-SVSPRGAPVLLVKKKDG------------------SMRLCVDY 586
++K++ +LLE I P S S +PVL+V KK+G S +LC+DY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 587 RQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGH 646
R+LN+ T K+ +PLP +D +++L G + +D GY+QI V +D K AF +G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 647 YEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQV 706
+ Y +PFG+ NAP F M IF +++ + VF+DD V+ S E + L +VLQ
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 707 LKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGL 766
E L KC F ++E LGH IS GI VD +K+D + + P +V I++ G
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 767 DGYYRRFIEGFSKL 780
+YRRFI+ F+K+
Sbjct: 721 ARFYRRFIKDFTKV 762
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 139 bits (349), Expect = 6e-33
Identities = 89/254 (35%), Positives = 132/254 (51%), Gaps = 19/254 (7%)
Frame = +1
Query: 546 LKKQLEELLEKQFIRP-SVSPRGAPVLLVKKKDGSM------------------RLCVDY 586
++K++ +LLE I P S S +PV +V KK G R+C+DY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 587 RQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGH 646
R+LN+ T K+ YPLP +D + +L + +D SGY+QI V +D KTAF +
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 647 YEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQV 706
+ Y MPFG+ NA F M IF +++ + VF+DD + S +L VLQ
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 707 LKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGL 766
+++ L KC F ++E LGH ISK GI V K+D + + P +V I + G
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 767 DGYYRRFIEGFSKL 780
G+YRRFI+ F+K+
Sbjct: 721 VGFYRRFIKDFTKV 762
>CF922488
Length = 741
Score = 129 bits (324), Expect = 5e-30
Identities = 71/202 (35%), Positives = 115/202 (56%)
Frame = +3
Query: 573 VKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKA 632
V K+DG + +CVDYR LN + K+++PLP I+ +D FS +D SGY+QI++
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 633 EDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKS 692
ED+ KT F T +G + Y M FG+ N + M +F + + + V++DD++V S++
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 693 EEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWE 752
EEEH +LR + + L++ +L +KC F +K L + S GI VD +KV +L+
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 753 SPKSVFEIKAIWGLDGYYRRFI 774
P + +++ G Y RFI
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFI 608
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 99.8 bits (247), Expect = 4e-21
Identities = 51/143 (35%), Positives = 83/143 (57%)
Frame = +3
Query: 581 RLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAF 640
R+CVDYR LN+ + K+ +PLP ID M + +FS +D SGY+QI++ ED+ KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 641 RTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHL 700
T +G + Y VM FG+ N + M +F + + + ++D+++ S+ EEEH +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 701 RIVLQVLKENQLCAKLSKCEFWL 723
+ + L++ +L KC F L
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFGL 431
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 96.7 bits (239), Expect = 4e-20
Identities = 49/133 (36%), Positives = 78/133 (57%)
Frame = +2
Query: 616 FSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYL 675
FS +D SGY+QI + ED+ KT F T +G + Y VM FG+ N + M +FH +
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 676 DRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISK 735
+ + V++DD++ S++E EH +L + L++ QL +KC F +K LG ++S+
Sbjct: 182 HKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQ 361
Query: 736 GGISVDPSKVDAV 748
GI +DP KV A+
Sbjct: 362 KGIEIDPEKVKAL 400
>BI317507
Length = 359
Score = 91.7 bits (226), Expect = 1e-18
Identities = 47/106 (44%), Positives = 68/106 (63%), Gaps = 1/106 (0%)
Frame = -1
Query: 685 DILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSK 744
+IL+YS + H HL VL VLK+ +L A KC F + +LGHVISK +++D +K
Sbjct: 353 NILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNK 174
Query: 745 VDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLA-LPLTQLTQ 789
V +V++W PK+V + + L GYYR+FI+ + KLA PLT LT+
Sbjct: 173 VKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAPRPLTDLTK 36
>BQ273711
Length = 409
Score = 80.9 bits (198), Expect = 2e-15
Identities = 42/100 (42%), Positives = 53/100 (53%)
Frame = -2
Query: 33 TAANRRNEGEAEERRLDRFLRNNPPTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRL 92
T R+ AE R L F +N+PP F G +DPEGA+ W+ E+IF AM ++ KV
Sbjct: 300 TLVQERDVEPAEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLY 121
Query: 93 ATHMFAEEAEYWWINAKGRLELGGDVVTWAKFKVEFLRKY 132
AT M EAE WW K G V+ W FK +FL Y
Sbjct: 120 ATFMLQGEAENWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 78.2 bits (191), Expect = 1e-14
Identities = 35/54 (64%), Positives = 42/54 (76%)
Frame = -3
Query: 48 LDRFLRNNPPTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEA 101
LDRF RNNPPTFKG +DPEGA+ W++ +E+IFR M D QKV ATHM A+EA
Sbjct: 164 LDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>BM142940
Length = 422
Score = 42.0 bits (97), Expect(2) = 7e-09
Identities = 17/44 (38%), Positives = 26/44 (58%)
Frame = -3
Query: 714 AKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSV 757
A KC F K+ +L H+IS G+S DP K++ ++ W PK +
Sbjct: 156 ANRKKCSFGCKKFEYLVHMISVKGVSADPKKLEVMVAWPIPKEM 25
Score = 37.0 bits (84), Expect(2) = 7e-09
Identities = 17/34 (50%), Positives = 24/34 (70%)
Frame = -1
Query: 679 VVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQL 712
+ F DIL+YS +E +H +HL+IVLQ KE +L
Sbjct: 260 ICFFSYDILIYSINEGKHEKHLKIVLQTFKEAKL 159
>CD487724
Length = 676
Score = 58.2 bits (139), Expect = 1e-08
Identities = 52/224 (23%), Positives = 96/224 (42%), Gaps = 20/224 (8%)
Frame = +3
Query: 352 PLIAIIDTGATHSFISASCVERLGLVV--TPLLRGMVIDTPASGSG---TTSFMCAKCPV 406
PL+ ++D G+TH+F+ V +LGL TP LR MV G+G + +C P+
Sbjct: 21 PLVYLVDGGSTHNFVQQPLVSQLGLPCRSTPPLRVMV------GNGHHLKCTTICEAIPI 182
Query: 407 NFGDVDFKLDLVCLPLKNMDVIFGMDWLLAFG---VSINCLTKSVTFSKPVEELGRE--- 460
+ +++F + L LP+ +++ G+ WL G V N L+ + + +L E
Sbjct: 183 SIQNIEFLVHLYVLPIVGANIVLGVQWLKTLGPILVDYNSLSMQFFYQHRLVQLKGESEA 362
Query: 461 ---FLTAEQVKKSLDGEACVFMMFASLKVSGEKGASVLPVVQEFPEVFPEDIT------E 511
L Q+++ V ++ S P+ Q + +
Sbjct: 363 QLGLLNHHQLRRLHQTHEPVTYFHIAILTENTSPTSSPPLPQPIQHLLDQFSALFQ*PQG 542
Query: 512 LPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLE 555
LP RE + I L+P + P+++ Y E++ Q+ +L+
Sbjct: 543 LPPARETDHHIHLLP*SEPVNMRLY*YPHY*NNEIEHQVNLMLQ 674
>CF922531
Length = 602
Score = 57.0 bits (136), Expect = 3e-08
Identities = 35/85 (41%), Positives = 52/85 (61%), Gaps = 1/85 (1%)
Frame = -2
Query: 494 VLPVVQEFPEVFPEDIT-ELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEE 552
V ++ EF ++FP++I LP R +E IDLVP S + YR + E E++ Q++E
Sbjct: 256 VQELLHEFGDIFPKEIPLGLPPLRGIEHQIDLVPRASLPNRPTYRTNPQETKEIESQVKE 77
Query: 553 LLEKQFIRPSVSPRGAPVLLVKKKD 577
LLEK +++ S+S VLLV KKD
Sbjct: 76 LLEKGWVQESLSLCVVLVLLVPKKD 2
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 55.8 bits (133), Expect = 7e-08
Identities = 30/77 (38%), Positives = 39/77 (49%)
Frame = +2
Query: 242 PYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVV 301
PY +D + G SR +L C C GHYA EC N CH CG GH A +C
Sbjct: 299 PYRRDSRR-----GFSRDNL----CKNCKRPGHYARECPNVAICHNCGLPGHIASEC--- 442
Query: 302 AREITCYNCGEKGHIST 318
+ C+NC E GH+++
Sbjct: 443 TTKSLCWNCKEPGHMAS 493
Score = 52.4 bits (124), Expect = 8e-07
Identities = 23/58 (39%), Positives = 32/58 (54%), Gaps = 4/58 (6%)
Frame = +2
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVVARE----ITCYNCGEKGHISTK 319
C+ C GH A+ C N+ CH CGKAGH+A +C C NC ++GHI+ +
Sbjct: 458 CWNCKEPGHMASSCPNEGICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAE 631
Score = 48.1 bits (113), Expect = 1e-05
Identities = 28/72 (38%), Positives = 34/72 (46%), Gaps = 1/72 (1%)
Frame = +2
Query: 228 VQNEKKGKGHGVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLN-CH 286
V ++ G G G G G + GGG R DV C C LGH + +CM L CH
Sbjct: 761 VLGDRSGGGGGGG-------GARGGGGGGYR----DVVCRNCQQLGHMSRDCMGPLMICH 907
Query: 287 KCGKAGHKAVDC 298
CG GH A +C
Sbjct: 908 NCGGRGHLAYEC 943
Score = 47.0 bits (110), Expect = 3e-05
Identities = 20/50 (40%), Positives = 25/50 (50%)
Frame = +2
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVVAREITCYNCGEKGH 315
C+ CG GH A+EC C C + GH A C E C+ CG+ GH
Sbjct: 401 CHNCGLPGHIASECTTKSLCWNCKEPGHMASSC---PNEGICHTCGKAGH 541
Score = 46.6 bits (109), Expect = 4e-05
Identities = 22/58 (37%), Positives = 28/58 (47%)
Frame = +2
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKA 323
C C GH A EC N+ C+ C K GH A DC + C C GH++ + KA
Sbjct: 593 CNNCYKQGHIAAECTNEKACNNCRKTGHLARDC---PNDPICNLCNVSGHVARQCPKA 757
Score = 34.7 bits (78), Expect = 0.17
Identities = 19/50 (38%), Positives = 25/50 (50%)
Frame = +2
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKAAGKVFALNAKE 334
C C + GH A +C VA C+NCG GHI++ + K N KE
Sbjct: 344 CKNCKRPGHYARECPNVA---ICHNCGLPGHIAS---ECTTKSLCWNCKE 475
>TC233976
Length = 763
Score = 54.7 bits (130), Expect = 2e-07
Identities = 43/120 (35%), Positives = 53/120 (43%), Gaps = 16/120 (13%)
Frame = +1
Query: 215 FDDAGKAKSNYYKVQN-----EKKGKGHGVGKPYS------KDKGKKRESGG----GSRP 259
FD + K +YK N EKK H KPYS G +R SGG G
Sbjct: 13 FD*DQQEKVAFYKNANASHGKEKKPMTHSRAKPYSAPLEYENHYGGQRTSGGHHLAGGSS 192
Query: 260 SLADVKCYRCGTLGHYANECMND-LNCHKCGKAGHKAVDCRVVAREITCYNCGEKGHIST 318
L + G G A + L C KCG+ GH A +C RE+TC+N KGH+ST
Sbjct: 193 QLVNRVSQPAGRGGSGAPAIVTTPLRCRKCGRLGHNAHECTY--REVTCFNYQGKGHLST 366
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 54.3 bits (129), Expect = 2e-07
Identities = 26/75 (34%), Positives = 41/75 (54%), Gaps = 14/75 (18%)
Frame = +1
Query: 257 SRPSLADVKCYRCGTLGHYANECMN-------DLNCHKCGKAGHKAVDCRVVAREIT--- 306
S+ L +++CY C LGH C+N +++C+KCG+ GH + C + EIT
Sbjct: 319 SQDDLKEIQCYVCKRLGHLC--CVNTDDATPGEISCYKCGQLGHTGLACSRLRDEITSGA 492
Query: 307 ----CYNCGEKGHIS 317
C+ CGE+GH +
Sbjct: 493 TPSSCFKCGEEGHFA 537
Score = 49.7 bits (117), Expect = 5e-06
Identities = 23/55 (41%), Positives = 28/55 (50%), Gaps = 5/55 (9%)
Frame = +1
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDC----RVVAREIT-CYNCGEKGH 315
CY CG LGH A +C +C C K GH+A DC ++ I C CG GH
Sbjct: 127 CYVCGCLGHNARQCSKVQDCFICKKGGHRAKDCPEKHTSTSKSIAICLKCGNSGH 291
Score = 40.8 bits (94), Expect = 0.002
Identities = 16/31 (51%), Positives = 19/31 (60%)
Frame = +1
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGH 315
C CG+ GH AV+C V R+ CY CG GH
Sbjct: 61 CFNCGEEGHAAVNCSAVKRKKPCYVCGCLGH 153
Score = 37.4 bits (85), Expect = 0.026
Identities = 16/46 (34%), Positives = 22/46 (47%), Gaps = 10/46 (21%)
Frame = +1
Query: 263 DVKCYRCGTLGHYANECMN----------DLNCHKCGKAGHKAVDC 298
++ CY+CG LGH C +C KCG+ GH A +C
Sbjct: 409 EISCYKCGQLGHTGLACSRLRDEITSGATPSSCFKCGEEGHFAREC 546
Score = 36.6 bits (83), Expect = 0.044
Identities = 19/53 (35%), Positives = 23/53 (42%), Gaps = 3/53 (5%)
Frame = +1
Query: 266 CYRCGTLGHYANECM---NDLNCHKCGKAGHKAVDCRVVAREITCYNCGEKGH 315
C+ CG GH A C C+ CG GH A C V C+ C + GH
Sbjct: 61 CFNCGEEGHAAVNCSAVKRKKPCYVCGCLGHNARQCSKVQ---DCFICKKGGH 210
Score = 30.4 bits (67), Expect = 3.2
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +1
Query: 252 ESGGGSRPSLADVKCYRCGTLGHYANECMNDLNCHK 287
E G+ PS C++CG GH+A EC + + K
Sbjct: 475 EITSGATPS----SCFKCGEEGHFARECTSSIKSGK 570
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 51.2 bits (121), Expect = 2e-06
Identities = 29/95 (30%), Positives = 40/95 (41%), Gaps = 23/95 (24%)
Frame = +2
Query: 254 GGGSRPSLADVKCYRCGTLGHYANECMNDLN----------CHKCGKAGHKAVDCRVVAR 303
GGGS C+RCG GH A +C C +CG+ GH A DC +
Sbjct: 218 GGGS--------CFRCGGFGHMARDCATGKGNIGGGGSGGGCFRCGEVGHLARDCGMEGG 373
Query: 304 EI-------------TCYNCGEKGHISTKPKKAAG 325
TC+NCG+ GH + + +A+G
Sbjct: 374 RFGGGGGSGGGGGKSTCFNCGKPGHFARECVEASG 478
Score = 45.4 bits (106), Expect = 9e-05
Identities = 29/101 (28%), Positives = 39/101 (37%), Gaps = 21/101 (20%)
Frame = +2
Query: 248 GKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLN-----------CHKCGKAGHKAV 296
G +R GGG+ CY+CG GH A +C N C CG GH A
Sbjct: 11 GARRNGGGGAA-------CYQCGEFGHLARDCNRSSNSGGGGGGSGGGCFNCGGFGHLAR 169
Query: 297 DC-RVVAREI---------TCYNCGEKGHISTKPKKAAGKV 327
DC R + +C+ CG GH++ G +
Sbjct: 170 DCVRGGGGSVGIGGGGGGGSCFRCGGFGHMARDCATGKGNI 292
Score = 32.0 bits (71), Expect = 1.1
Identities = 13/34 (38%), Positives = 18/34 (52%)
Frame = +2
Query: 247 KGKKRESGGGSRPSLADVKCYRCGTLGHYANECM 280
+G + GGGS C+ CG GH+A EC+
Sbjct: 365 EGGRFGGGGGSGGGGGKSTCFNCGKPGHFARECV 466
>BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, partial (3%)
Length = 153
Score = 50.8 bits (120), Expect = 2e-06
Identities = 25/51 (49%), Positives = 33/51 (64%)
Frame = -1
Query: 656 VSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQV 706
++NAP F MN IF L ++V+VF DDILVYS + EH HL +V +V
Sbjct: 153 LTNAPTSFQCLMNHIFQHALRKYVLVFFDDILVYSSTWHEHLCHLEVVFKV 1
>TC227589
Length = 547
Score = 49.7 bits (117), Expect = 5e-06
Identities = 25/75 (33%), Positives = 38/75 (50%), Gaps = 14/75 (18%)
Frame = +2
Query: 257 SRPSLADVKCYRCGTLGHYANECMN-------DLNCHKCGKAGHKAVDCRVVAREIT--- 306
S L +++CY C +GH C+N +++C+KCG+ GH + C + EIT
Sbjct: 305 SPDDLKEIQCYVCKRVGHLC--CVNTDDATPGEISCYKCGQLGHTGLACSKLPDEITSAA 478
Query: 307 ----CYNCGEKGHIS 317
C CGE GH +
Sbjct: 479 TPSSCCKCGEAGHFA 523
Score = 49.3 bits (116), Expect = 7e-06
Identities = 23/55 (41%), Positives = 28/55 (50%), Gaps = 5/55 (9%)
Frame = +2
Query: 266 CYRCGTLGHYANECMNDLNCHKCGKAGHKAVDC--RVVARE---ITCYNCGEKGH 315
CY CG LGH A +C +C C K GH+A DC + +R C CG GH
Sbjct: 113 CYVCGGLGHNARQCTKAQDCFICKKGGHRAKDCLEKHTSRSKSVAICLKCGNSGH 277
Score = 40.8 bits (94), Expect = 0.002
Identities = 17/39 (43%), Positives = 22/39 (55%)
Frame = +2
Query: 285 CHKCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKKA 323
C CG+ GH AV+C R+ CY CG GH + + KA
Sbjct: 47 CFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNARQCTKA 163
Score = 39.7 bits (91), Expect = 0.005
Identities = 23/76 (30%), Positives = 30/76 (39%), Gaps = 13/76 (17%)
Frame = +2
Query: 254 GGGSRPSLADVKCYRCGTLGHYANECMNDLN--------CHKCGKAGHKAVDCR-----V 300
G +R C+ C GH A +C+ C KCG +GH CR
Sbjct: 134 GHNARQCTKAQDCFICKKGGHRAKDCLEKHTSRSKSVAICLKCGNSGHDMFSCRNDYSPD 313
Query: 301 VAREITCYNCGEKGHI 316
+EI CY C GH+
Sbjct: 314 DLKEIQCYVCKRVGHL 361
Score = 38.9 bits (89), Expect = 0.009
Identities = 17/46 (36%), Positives = 23/46 (49%), Gaps = 10/46 (21%)
Frame = +2
Query: 263 DVKCYRCGTLGHYANECMN----------DLNCHKCGKAGHKAVDC 298
++ CY+CG LGH C +C KCG+AGH A +C
Sbjct: 395 EISCYKCGQLGHTGLACSKLPDEITSAATPSSCCKCGEAGHFAQEC 532
Score = 35.8 bits (81), Expect = 0.075
Identities = 18/53 (33%), Positives = 23/53 (42%), Gaps = 3/53 (5%)
Frame = +2
Query: 266 CYRCGTLGHYANECM---NDLNCHKCGKAGHKAVDCRVVAREITCYNCGEKGH 315
C+ CG GH A C C+ CG GH A C + C+ C + GH
Sbjct: 47 CFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNARQC---TKAQDCFICKKGGH 196
>TC214169 similar to UP|Q8LF25 (Q8LF25) DNA-damage inducible protein
DDI1-like, partial (90%)
Length = 1604
Score = 48.9 bits (115), Expect = 9e-06
Identities = 30/91 (32%), Positives = 46/91 (49%), Gaps = 4/91 (4%)
Frame = +2
Query: 348 INSTPLIAIIDTGATHSFISASCVERLGL--VVTPLLRGMVIDTPASGSGTTSFM--CAK 403
+N PL A +D+GA + IS SC ERLGL ++ RG+ A G G + +
Sbjct: 764 VNGVPLKAFVDSGAQSTIISKSCAERLGLLRLLDQRYRGI-----AHGVGQSEILGRIHV 928
Query: 404 CPVNFGDVDFKLDLVCLPLKNMDVIFGMDWL 434
P+ G + + + L NM+ +FG+D L
Sbjct: 929 APIKIGSIFYPCSFLVLDSPNMEFLFGLDML 1021
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,297,714
Number of Sequences: 63676
Number of extensions: 458404
Number of successful extensions: 2276
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 2096
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2219
length of query: 790
length of database: 12,639,632
effective HSP length: 105
effective length of query: 685
effective length of database: 5,953,652
effective search space: 4078251620
effective search space used: 4078251620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC126008.1


