

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
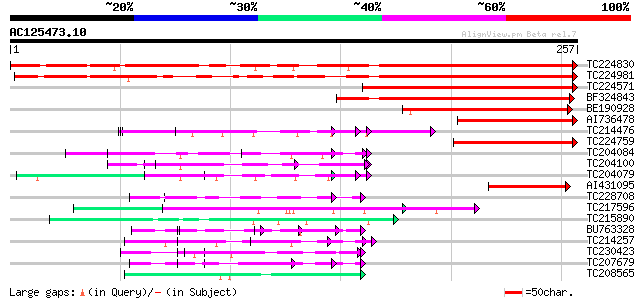
Query= AC125473.10 + phase: 0
(257 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC224830 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-... 258 2e-69
TC224981 similar to UP|O50000 (O50000) Abscisic stress ripening ... 238 3e-63
TC224571 similar to UP|Q9ZRB6 (Q9ZRB6) Ci21A protein, complete 155 2e-38
BF324843 136 8e-33
BE190928 107 5e-24
AI736478 97 9e-21
TC214476 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 70 9e-13
TC224759 similar to UP|Q9SPF3 (Q9SPF3) Ripening induced protein ... 68 5e-12
TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, co... 65 2e-11
TC204100 homologue to UP|Q93WA5 (Q93WA5) Ribosomal protein L32, ... 65 3e-11
TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding ... 65 4e-11
AI431095 weakly similar to PIR|T09789|T097 abscisic acid- and wa... 63 1e-10
TC228708 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wal... 63 2e-10
TC217596 weakly similar to UP|Q8K357 (Q8K357) Procr protein, par... 62 2e-10
TC215890 similar to PIR|F96770|F96770 protein RNA-binding protei... 62 2e-10
BU763328 similar to SP|P10496|GRP2_ Glycine-rich cell wall struc... 62 3e-10
TC214257 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 60 7e-10
TC230423 similar to UP|Q41187 (Q41187) Glycine-rich protein (Fra... 60 7e-10
TC207679 similar to UP|Q7XJI7 (Q7XJI7) Glycine-rich protein TomR... 60 7e-10
TC208565 similar to GB|AAM97131.1|22531255|AY136466 ribonucleopr... 60 1e-09
>TC224830 similar to UP|Q93WZ6 (Q93WZ6) Abscisic stress ripening-like
protein, partial (61%)
Length = 1118
Score = 258 bits (658), Expect = 2e-69
Identities = 148/268 (55%), Positives = 175/268 (65%), Gaps = 11/268 (4%)
Frame = +3
Query: 1 MSEEENRSRGLFHHKKHDDDDNRPIDTDSGYNKTSSYSTGDDDYNK---KTSYGNDNSGG 57
M+EE++ LFHH H D+DN+P++TD+GY+ TS YS DDY+ K SY ++SGG
Sbjct: 171 MAEEKHHKHRLFHH--HKDEDNKPVETDTGYDNTS-YSKPSDDYDSGFNKPSY--ESSGG 335
Query: 58 DYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG-GGYGDS 116
YE GYNK + + + GGY G KT GY GG + T GGYG GG G S
Sbjct: 336 GYEGGYNKTSYSSDEPASGGGYEGGYKKT----GYSGGIDE------TSGGYGAGGGGYS 485
Query: 117 DTKTTTGGGYG---GGYGDSDTKTTTGGGYGGGYGDSDT----KTTTGGGYGDDRREDVD 169
D T GGYG GGYG G GG Y D+ T T+GGGY D+ VD
Sbjct: 486 DV---TDGGYGKPSGGYG--------AAGGGGVYPDTTTDGGYSKTSGGGYDDE----VD 620
Query: 170 YEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGF 229
Y+KEEK H EHLGELGAA+A +AL+EKHK+EKDPE+AHRHKIEEEVAA AAVGSGGF
Sbjct: 621 YKKEEKHHKHLEHLGELGAASAAAYALHEKHKAEKDPEHAHRHKIEEEVAAAAAVGSGGF 800
Query: 230 AFHEHHEKKETKEEEEEAHGKKKHHFFG 257
AFHEHHEKKE KE++EEAHGKK HH FG
Sbjct: 801 AFHEHHEKKEAKEQDEEAHGKKHHHLFG 884
>TC224981 similar to UP|O50000 (O50000) Abscisic stress ripening protein
homolog, partial (50%)
Length = 849
Score = 238 bits (606), Expect = 3e-63
Identities = 137/256 (53%), Positives = 168/256 (65%), Gaps = 1/256 (0%)
Frame = +1
Query: 3 EEENRSRGLFHHKKHDDDDNRPIDTDSGYNKTSSYSTGDDDYNKKTSYGN-DNSGGDYET 61
EE++ LFH +H D+DN+P +T++GY+ TS YS DDY+ S + + SGG YET
Sbjct: 82 EEKHHKHHLFH--QHKDEDNKPTETETGYDNTS-YSKPSDDYDSGFSNPSYETSGGAYET 252
Query: 62 GYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTT 121
G+NK + YS+++ + +GGGY K T GY GG + T+
Sbjct: 253 GHNKTS--YSSDD----------QPASGGGY---------KKT---GYSGGIDE----TS 348
Query: 122 TGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHE 181
GGGYGGG G + TTT GGY T+GG YGD+ VDY+KEEK H E
Sbjct: 349 GGGGYGGG-GGVYSDTTTDGGY---------TRTSGGRYGDE----VDYKKEEKHHKSLE 486
Query: 182 HLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETK 241
HLGELGAAAAG +AL+EKHK+EKDPE+AHRHKIEEEVAA AAVGSGGFAFHEHHEKKE K
Sbjct: 487 HLGELGAAAAGAYALHEKHKAEKDPEHAHRHKIEEEVAAAAAVGSGGFAFHEHHEKKEAK 666
Query: 242 EEEEEAHGKKKHHFFG 257
E++EEAHGKK HH FG
Sbjct: 667 EQDEEAHGKKHHHLFG 714
>TC224571 similar to UP|Q9ZRB6 (Q9ZRB6) Ci21A protein, complete
Length = 762
Score = 155 bits (392), Expect = 2e-38
Identities = 69/97 (71%), Positives = 82/97 (84%)
Frame = +3
Query: 161 GDDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAA 220
GDD +VDY+KEEK H EHLGELGAAAAG +AL+EKH+++KDPE+AHRHK+EEE+AA
Sbjct: 150 GDDVPGEVDYKKEEKHHKHLEHLGELGAAAAGAYALHEKHEAKKDPEHAHRHKVEEEIAA 329
Query: 221 VAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFFG 257
A VG+GGF HEHHEKKE K+E+EEAHGKK HH FG
Sbjct: 330 AATVGAGGFVLHEHHEKKEVKKEDEEAHGKKHHHLFG 440
>BF324843
Length = 313
Score = 136 bits (343), Expect = 8e-33
Identities = 64/108 (59%), Positives = 79/108 (72%)
Frame = +2
Query: 149 DSDTKTTTGGGYGDDRREDVDYEKEEKRHSKHEHLGELGAAAAGGFALYEKHKSEKDPEN 208
D T+GGGY D+ VDY KE H EHLGEL AA+A ++L++KHK++KDPE+
Sbjct: 2 DGGYSKTSGGGYDDE----VDYNKENNLHMHLEHLGELSAASAAAYSLHDKHKAQKDPEH 169
Query: 209 AHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFF 256
AH HKI EEVAA AAVGS GFAFHEHH+K E ++++EEAH KK HH F
Sbjct: 170 AHTHKI*EEVAAAAAVGSDGFAFHEHHDKNEAQDQDEEAHQKKHHHLF 313
>BE190928
Length = 235
Score = 107 bits (267), Expect = 5e-24
Identities = 54/78 (69%), Positives = 61/78 (77%), Gaps = 1/78 (1%)
Frame = -1
Query: 179 KH-EHLGELGAAAAGGFALYEKHKSEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEK 237
KH EHL LG A+ AL+ KHK+EKDPE+A R KIEE+VAA AAVGSGGFAFH HH K
Sbjct: 235 KHLEHLS*LGPASVAASALHYKHKAEKDPEHALRPKIEEQVAAAAAVGSGGFAFHLHHAK 56
Query: 238 KETKEEEEEAHGKKKHHF 255
K KE++EEAHGKK HHF
Sbjct: 55 K*AKEQDEEAHGKKLHHF 2
>AI736478
Length = 419
Score = 96.7 bits (239), Expect = 9e-21
Identities = 40/54 (74%), Positives = 47/54 (86%)
Frame = +2
Query: 204 KDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFFG 257
+DPE+AHRHK+EEE+AA A VG+GGF HEHHEKKE K+E+EEAHGKK HH FG
Sbjct: 5 EDPEHAHRHKVEEEIAAAATVGAGGFVLHEHHEKKEVKKEDEEAHGKKHHHLFG 166
>TC214476 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (14%)
Length = 642
Score = 70.1 bits (170), Expect = 9e-13
Identities = 49/145 (33%), Positives = 65/145 (44%), Gaps = 3/145 (2%)
Frame = -3
Query: 52 NDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGG 111
++ GG G +K Y + GG GS G + GG GG G + + GGG GG
Sbjct: 640 SEGGGGSAGGGGSKGGGGYGGGGSQGGGGSAGGGGS*GGS-GGSAGGGEGSGSQGGGSGG 464
Query: 112 GYGDSDTKTTTGGGYGGG-YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGY--GDDRREDV 168
G + GGGYGGG G S+ ++GGG GGG G GGGY G D+ +
Sbjct: 463 GGSQGGGSGSGGGGYGGGGSGGSEGGGSSGGGGGGGGGGG------GGGYGGGGDKGGGI 302
Query: 169 DYEKEEKRHSKHEHLGELGAAAAGG 193
+K + H K + G GG
Sbjct: 301 GGDKGDYHHGKGGKGDKDGGDKGGG 227
Score = 64.7 bits (156), Expect = 4e-11
Identities = 45/116 (38%), Positives = 53/116 (44%), Gaps = 1/116 (0%)
Frame = -3
Query: 50 YGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGY 109
YG S G + + S GG GSG + GGG GGG + GGGY
Sbjct: 586 YGGGGSQGGGGSAGGGGS*GGSGGSAGGGEGSG----SQGGGSGGGGSQGGGSGSGGGGY 419
Query: 110 -GGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDR 164
GGG G S+ ++GGG GGG GGG GGGYG K GGG G D+
Sbjct: 418 GGGGSGGSEGGGSSGGGGGGG----------GGGGGGGYGGGGDK---GGGIGGDK 290
Score = 59.3 bits (142), Expect = 2e-09
Identities = 43/96 (44%), Positives = 47/96 (48%), Gaps = 12/96 (12%)
Frame = -3
Query: 76 SGGYGS-GGAKTTTGGGYGGG-YGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDS 133
SGG GS GG + GGGYGGG G S+ ++GGG GGG GGG GGGYG
Sbjct: 472 SGGGGSQGGGSGSGGGGYGGGGSGGSEGGGSSGGGGGGG----------GGGGGGGYGGG 323
Query: 134 DTKTTTGGGYG----------GGYGDSDTKTTTGGG 159
K GGG G GG GD D GGG
Sbjct: 322 GDK---GGGIGGDKGDYHHGKGGKGDKDGGDKGGGG 224
Score = 48.1 bits (113), Expect = 4e-06
Identities = 37/98 (37%), Positives = 44/98 (44%)
Frame = -3
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG 110
G+ + GG Y G + ++ ++ GG G GG GGGYGGG GD GGG G
Sbjct: 445 GSGSGGGGYGGGGSGGSEGGGSSGGGGGGGGGGG----GGGYGGG-GDK------GGGIG 299
Query: 111 GGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 148
G GD GG GD D GG GGG G
Sbjct: 298 GDKGDYHHGK-------GGKGDKD-----GGDKGGGGG 221
>TC224759 similar to UP|Q9SPF3 (Q9SPF3) Ripening induced protein (Fragment),
partial (78%)
Length = 427
Score = 67.8 bits (164), Expect = 5e-12
Identities = 31/56 (55%), Positives = 36/56 (63%)
Frame = +1
Query: 202 SEKDPENAHRHKIEEEVAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHHFFG 257
SE P+ E+ A VG+GGF HEHHEKKE K+E+EEAHGKK HH FG
Sbjct: 175 SEIVPDTVVFDDFEKFPPTAATVGAGGFVLHEHHEKKEVKKEDEEAHGKKHHHLFG 342
>TC204084 UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein, complete
Length = 1937
Score = 65.5 bits (158), Expect = 2e-11
Identities = 51/143 (35%), Positives = 62/143 (42%), Gaps = 6/143 (4%)
Frame = +2
Query: 26 DTDSGYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETS------GGY 79
D ++G ++ + T + + K + G N G N + N + NE GG
Sbjct: 1343 DRETGRSRGFGFVTFASEQSMKDAIGAMN-------GQNLDGRNITVNEAQSRGGGGGGG 1501
Query: 80 GSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTT 139
G GG GGGYGG G GGG GGGY D GGYGGGYG
Sbjct: 1502 GGGGGYNRGGGGYGGRSG--------GGGGGGGYRSRD------GGYGGGYGGGG----- 1624
Query: 140 GGGYGGGYGDSDTKTTTGGGYGD 162
GGGYGGG D + GG GD
Sbjct: 1625 GGGYGGG---RDRGYSRGGDGGD 1684
Score = 57.4 bits (137), Expect = 6e-09
Identities = 40/104 (38%), Positives = 45/104 (42%)
Frame = +2
Query: 45 NKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTT 104
N+ S G GG GYN+ GGYG + GGG GGGY D
Sbjct: 1463 NEAQSRGGGGGGGGGGGGYNRG---------GGGYGG----RSGGGGGGGGYRSRD---- 1591
Query: 105 TGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 148
GGYGGGY GGG GGGYG + + GG GG G
Sbjct: 1592 --GGYGGGY---------GGGGGGGYGGGRDRGYSRGGDGGDGG 1690
Score = 43.5 bits (101), Expect = 9e-05
Identities = 31/69 (44%), Positives = 31/69 (44%), Gaps = 10/69 (14%)
Frame = +2
Query: 106 GGGYGGGYGDSDTKTTTGGGY---GGGYGDSDTKTTTG-------GGYGGGYGDSDTKTT 155
GGG GGG G GGGY GGGYG G GGYGGGYG
Sbjct: 1481 GGGGGGGGG--------GGGYNRGGGGYGGRSGGGGGGGGYRSRDGGYGGGYGGGG---- 1624
Query: 156 TGGGYGDDR 164
GGGYG R
Sbjct: 1625 -GGGYGGGR 1648
Score = 32.3 bits (72), Expect = 0.22
Identities = 20/48 (41%), Positives = 22/48 (45%)
Frame = +2
Query: 123 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDY 170
GGG GGG G GGGY G G ++ GGG G R D Y
Sbjct: 1481 GGGGGGGGG--------GGGYNRGGGGYGGRSGGGGGGGGYRSRDGGY 1600
>TC204100 homologue to UP|Q93WA5 (Q93WA5) Ribosomal protein L32, complete
Length = 1145
Score = 65.1 bits (157), Expect = 3e-11
Identities = 41/97 (42%), Positives = 45/97 (46%)
Frame = -3
Query: 67 TDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGY 126
T N + + GG G GG GGGYGG G GGG GGGY D GGY
Sbjct: 414 TVNEAQSRGGGGGGGGGGYNRGGGGYGGRSG--------GGGGGGGYRSRD------GGY 277
Query: 127 GGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDD 163
GGGYG GG GGGYG + + GG G D
Sbjct: 276 GGGYG---------GGGGGGYGGGRDRGYSRGGDGGD 193
Score = 57.8 bits (138), Expect = 5e-09
Identities = 44/105 (41%), Positives = 46/105 (42%), Gaps = 2/105 (1%)
Frame = -3
Query: 62 GYNKNTDNYSNNETS--GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTK 119
G N + N + NE GG G GG GGGY G GGGYGG G
Sbjct: 441 GQNLDGRNITVNEAQSRGGGGGGG-----GGGYNRG----------GGGYGGRSG----- 322
Query: 120 TTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDR 164
GGG GGGY D GGYGGGYG GGGYG R
Sbjct: 321 ---GGGGGGGYRSRD------GGYGGGYGGGG-----GGGYGGGR 229
Score = 55.8 bits (133), Expect = 2e-08
Identities = 35/87 (40%), Positives = 40/87 (45%)
Frame = -3
Query: 45 NKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTT 104
N+ S G GG GYN+ Y SGG G GG + GGYGGGYG
Sbjct: 408 NEAQSRGGGGGGGG--GGYNRGGGGYGGR--SGGGGGGGGYRSRDGGYGGGYG------- 262
Query: 105 TGGGYGGGYGDSDTKTTTGGGYGGGYG 131
GG GGGYG + + GG GG G
Sbjct: 261 --GGGGGGYGGGRDRGYSRGGDGGDGG 187
Score = 30.8 bits (68), Expect = 0.63
Identities = 27/93 (29%), Positives = 33/93 (35%), Gaps = 1/93 (1%)
Frame = -3
Query: 21 DNRPIDTDSGYNKTSSYSTGDDDYNKKTS-YGNDNSGGDYETGYNKNTDNYSNNETSGGY 79
D R I + ++ G YN+ YG + GG GY Y GG
Sbjct: 429 DGRNITVNEAQSRGGGGGGGGGGYNRGGGGYGGRSGGGGGGGGYRSRDGGYGGGY-GGG- 256
Query: 80 GSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGG 112
GGGYGGG D + GG G G
Sbjct: 255 --------GGGGYGGG---RDRGYSRGGDGGDG 190
Score = 30.4 bits (67), Expect = 0.83
Identities = 19/48 (39%), Positives = 21/48 (43%)
Frame = -3
Query: 123 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDRREDVDY 170
GGG GGG GGGY G G ++ GGG G R D Y
Sbjct: 390 GGGGGGG----------GGGYNRGGGGYGGRSGGGGGGGGYRSRDGGY 277
>TC204079 similar to UP|Q9SWA8 (Q9SWA8) Glycine-rich RNA-binding protein,
complete
Length = 748
Score = 64.7 bits (156), Expect = 4e-11
Identities = 44/101 (43%), Positives = 50/101 (48%), Gaps = 3/101 (2%)
Frame = +1
Query: 62 GYNKNTDNYSNNETS---GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDT 118
G N + N + NE GG G GG GGG GGGYG GGG GGG G +
Sbjct: 277 GQNLDGRNITVNEAQSRGGGGGGGGGGGGYGGGRGGGYG--------GGGRGGGGGYN-- 426
Query: 119 KTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGG 159
++ GGGYGGG G GGG GGGYG + GGG
Sbjct: 427 RSGGGGGYGGGGG-------YGGGGGGGYGGGRDRGYGGGG 528
Score = 52.8 bits (125), Expect = 2e-07
Identities = 37/81 (45%), Positives = 39/81 (47%), Gaps = 5/81 (6%)
Frame = +1
Query: 89 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGG---GYGDSDTKTTTGGGY-- 143
GGG GGG G GGGYGGG GGGYGG G G ++ GGGY
Sbjct: 328 GGGGGGGGG--------GGGYGGG---------RGGGYGGGGRGGGGGYNRSGGGGGYGG 456
Query: 144 GGGYGDSDTKTTTGGGYGDDR 164
GGGYG GGGYG R
Sbjct: 457 GGGYGGGG-----GGGYGGGR 504
Score = 49.3 bits (116), Expect = 2e-06
Identities = 51/156 (32%), Positives = 60/156 (37%), Gaps = 11/156 (7%)
Frame = +1
Query: 4 EENRSRGL-FHHKKHDDDDNRPIDTDSGYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETG 62
E RSRG F + I+ +G N T ++ ++ G GG Y G
Sbjct: 196 ETGRSRGFGFVTFASEQSMKDAIEGMNGQNLDGRNITVNEAQSRGGGGGGGGGGGGYGGG 375
Query: 63 YNKNTDNYSNNETSGGYGSGGAKTTTGGGY-----GGGYGDSDTKTTTGGGYG-GGYGDS 116
GGYG GG GGGY GGGYG GGGYG GG
Sbjct: 376 ------------RGGGYGGGGRGG--GGGYNRSGGGGGYGG-------GGGYGGGG---- 480
Query: 117 DTKTTTGGGYGG----GYGDSDTKTTTGGGYGGGYG 148
GGGYGG GYG + + GG GG G
Sbjct: 481 ------GGGYGGGRDRGYGGGGDRGYSRGGDGGDGG 570
>AI431095 weakly similar to PIR|T09789|T097 abscisic acid- and
water-stress-inducible protein LP3-1 - loblolly pine,
partial (30%)
Length = 115
Score = 63.2 bits (152), Expect = 1e-10
Identities = 27/37 (72%), Positives = 31/37 (82%)
Frame = -1
Query: 218 VAAVAAVGSGGFAFHEHHEKKETKEEEEEAHGKKKHH 254
VAA A VG GGFA HEHH+KK KE++EEAHGKK+HH
Sbjct: 115 VAAAAQVGYGGFALHEHHDKKIAKEQDEEAHGKKQHH 5
>TC228708 similar to UP|GRP2_PHAVU (P10496) Glycine-rich cell wall structural
protein 1.8 precursor (GRP 1.8), partial (15%)
Length = 494
Score = 62.8 bits (151), Expect = 2e-10
Identities = 39/91 (42%), Positives = 45/91 (48%)
Frame = +3
Query: 71 SNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGY 130
+ E GYG G +GGG GGGYGD GG +GGGYG G G GGGY
Sbjct: 9 AGGEHGXGYGGAGG---SGGGSGGGYGD-------GGAHGGGYGGG-----AGAGGGGGY 143
Query: 131 GDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
G GGG+GGG G +GGG+G
Sbjct: 144 GAGGAH---GGGFGGGGG-------SGGGHG 206
Score = 60.8 bits (146), Expect = 6e-10
Identities = 39/94 (41%), Positives = 46/94 (48%)
Frame = +3
Query: 55 SGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 114
+GG++ GY S + GGYG GGA GGGYGGG G G GGGYG
Sbjct: 9 AGGEHGXGYGGAGG--SGGGSGGGYGDGGAH---GGGYGGG---------AGAGGGGGYG 146
Query: 115 DSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 148
GG +GGG+G GGG GGG+G
Sbjct: 147 -------AGGAHGGGFGG-------GGGSGGGHG 206
>TC217596 weakly similar to UP|Q8K357 (Q8K357) Procr protein, partial (9%)
Length = 988
Score = 62.4 bits (150), Expect = 2e-10
Identities = 47/161 (29%), Positives = 65/161 (40%), Gaps = 17/161 (10%)
Frame = +3
Query: 70 YSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGG----- 124
Y + YGSG YG GYG ++ G GY G +S+ + GG
Sbjct: 93 YGGRKXESEYGSGYGGRKEESEYGSGYGGRKEESEYGSGYDGRKEESEYGSGYGGRKQES 272
Query: 125 GYGGGYGDSDTKTTTGGGYGG--------GYGDSDTKTTTGGG---YGDDRREDVDYEKE 173
YG GYG ++ G GYGG YG S+ ++ GGG YG + R E+E
Sbjct: 273 EYGSGYGGRKQESEYGSGYGGRSEYEEKPSYGRSNYESQGGGGGVEYGYEGRPPRQEEEE 452
Query: 174 EKRHSKHEHLGELGAAAAG-GFALYEKHKSEKDPENAHRHK 213
E+ + K + + G G Y + D H HK
Sbjct: 453 EEGYRKPSYQRKDDDDEEGYGRKKYGDDSDDDDGRKKHHHK 575
Score = 54.3 bits (129), Expect = 5e-08
Identities = 49/166 (29%), Positives = 66/166 (39%), Gaps = 15/166 (9%)
Frame = +3
Query: 30 GYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTG 89
G S Y +G +++ YG+ G E+ Y D E+ G G GG K +
Sbjct: 99 GRKXESEYGSGYGGRKEESEYGSGYGGRKEESEYGSGYDG-RKEESEYGSGYGGRKQESE 275
Query: 90 GGYGGGYGDSDTKTTTGGGYGG--------GYGDSDTKTTTGGG---YG-GGYGDSDTKT 137
YG GYG ++ G GYGG YG S+ ++ GGG YG G +
Sbjct: 276 --YGSGYGGRKQESEYGSGYGGRSEYEEKPSYGRSNYESQGGGGGVEYGYEGRPPRQEEE 449
Query: 138 TTGGGYGGGYGDSDTKTTTGGG---YGDDRREDVDYEKEEKRHSKH 180
G Y D G G YGDD +D + +K H KH
Sbjct: 450 EEEGYRKPSYQRKDDDDEEGYGRKKYGDDSDDD---DGRKKHHHKH 578
>TC215890 similar to PIR|F96770|F96770 protein RNA-binding protein F1O17.10
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (17%)
Length = 913
Score = 62.4 bits (150), Expect = 2e-10
Identities = 55/174 (31%), Positives = 69/174 (39%), Gaps = 16/174 (9%)
Frame = +2
Query: 19 DDDNRPIDTDSGYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGG 78
D RPI + + Y G G G Y GY DNYS N+ SG
Sbjct: 62 DLHGRPIRVNYANERPRGYGGGGFGSYGAVGGGGYEGGSSYRGGYGG--DNYSRNDGSG- 232
Query: 79 YGSGGAKTTTGGGYGGGYGDSDTKTTTGGGY---GGGYGDSDTKT----------TTGGG 125
YG GG + G GG YGDS + GGY GG G+ ++ T GG
Sbjct: 233 YGYGGGRY----GSGGNYGDSGSGNNYSGGYAGNAGGVGNHESSTGFASNGYDGSVVDGG 400
Query: 126 YGGGYGDS--DTKTTTGGGYGGGYGDSDTKTTTGGGYG-DDRREDVDYEKEEKR 176
G G G S D + G G G D+K ++ G D R+D D + KR
Sbjct: 401 VGAGSGTSFADGYDGSAGSEFGSSGQLDSKASSKGDEDFGDYRDDNDADDFAKR 562
>BU763328 similar to SP|P10496|GRP2_ Glycine-rich cell wall structural
protein 1.8 precursor (GRP 1.8). [Kidney bean French
bean], partial (11%)
Length = 169
Score = 61.6 bits (148), Expect = 3e-10
Identities = 37/73 (50%), Positives = 39/73 (52%)
Frame = -3
Query: 78 GYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKT 137
GYG GGA GGGYGGG G GG GGGYG GG +GGGYG +
Sbjct: 167 GYGDGGAH---GGGYGGGQG---------GGAGGGYG-------AGGEHGGGYGGGE--- 54
Query: 138 TTGGGYGGGYGDS 150
GGG GGGYG S
Sbjct: 53 --GGGAGGGYGAS 21
Score = 44.7 bits (104), Expect = 4e-05
Identities = 29/57 (50%), Positives = 30/57 (51%)
Frame = -3
Query: 77 GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDS 133
GGYG GG GGGYG G G+ GGGYGGG GGG GGGYG S
Sbjct: 140 GGYG-GGQGGGAGGGYGAG-GEH------GGGYGGG---------EGGGAGGGYGAS 21
Score = 41.2 bits (95), Expect = 5e-04
Identities = 26/61 (42%), Positives = 28/61 (45%)
Frame = -3
Query: 56 GGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGD 115
GG + GY GGYG+GG GGGYGGG GGG GGGYG
Sbjct: 155 GGAHGGGYGGG----QGGGAGGGYGAGGEH---GGGYGGG---------EGGGAGGGYGA 24
Query: 116 S 116
S
Sbjct: 23 S 21
Score = 40.4 bits (93), Expect = 8e-04
Identities = 24/51 (47%), Positives = 27/51 (52%), Gaps = 1/51 (1%)
Frame = -3
Query: 112 GYGDSDTKTTTGGGYGGGY-GDSDTKTTTGGGYGGGYGDSDTKTTTGGGYG 161
GYGD GGGYGGG G + GG +GGGYG + GGGYG
Sbjct: 167 GYGDGGAH---GGGYGGGQGGGAGGGYGAGGEHGGGYGGGE-GGGAGGGYG 27
>TC214257 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (7%)
Length = 607
Score = 60.5 bits (145), Expect = 7e-10
Identities = 40/91 (43%), Positives = 44/91 (47%), Gaps = 5/91 (5%)
Frame = +2
Query: 77 GGYGSGGAKTTTGGGY-----GGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 131
GGYG GG + GGGY GGGYG GGGYGGG G GGGYGGG
Sbjct: 14 GGYGGGGRREGGGGGYNRNGGGGGYGG-------GGGYGGGGGHGG-----GGGYGGGGR 157
Query: 132 DSDTKTTTGGGYGGGYGDSDTKTTTGGGYGD 162
D GYG GD ++ + GGG D
Sbjct: 158 DR--------GYG---GDGGSRYSRGGGASD 217
Score = 49.3 bits (116), Expect = 2e-06
Identities = 36/96 (37%), Positives = 41/96 (42%), Gaps = 2/96 (2%)
Frame = +2
Query: 53 DNSGGDYETGYNKNTDN--YSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYG 110
D GG Y G + Y+ N GGYG GGGYGGG G GGGYG
Sbjct: 2 DRGGGGYGGGGRREGGGGGYNRNGGGGGYGG-------GGGYGGGGGHGG-----GGGYG 145
Query: 111 GGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGG 146
GG D GYG GD ++ + GGG G
Sbjct: 146 GGGRDR--------GYG---GDGGSRYSRGGGASDG 220
Score = 46.2 bits (108), Expect = 1e-05
Identities = 28/62 (45%), Positives = 30/62 (48%), Gaps = 5/62 (8%)
Frame = +2
Query: 110 GGGYGDSDTKTTTGGGY-----GGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGDDR 164
GGGYG + GGGY GGGYG GGGYGGG G GGGYG
Sbjct: 11 GGGYGGGGRREGGGGGYNRNGGGGGYGG-------GGGYGGGGGHGG-----GGGYGGGG 154
Query: 165 RE 166
R+
Sbjct: 155 RD 160
Score = 33.5 bits (75), Expect = 0.098
Identities = 25/83 (30%), Positives = 30/83 (36%)
Frame = +2
Query: 30 GYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTG 89
GY G YN+ G GG Y G + GGYG GG
Sbjct: 17 GYGGGGRREGGGGGYNRNGGGGGYGGGGGYGGG--------GGHGGGGGYGGGGRDR--- 163
Query: 90 GGYGGGYGDSDTKTTTGGGYGGG 112
GYG GD ++ + GGG G
Sbjct: 164 -GYG---GDGGSRYSRGGGASDG 220
>TC230423 similar to UP|Q41187 (Q41187) Glycine-rich protein (Fragment),
partial (40%)
Length = 522
Score = 60.5 bits (145), Expect = 7e-10
Identities = 37/85 (43%), Positives = 42/85 (48%)
Frame = +2
Query: 77 GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTK 136
GG G GG GGG+GGG+G GGG GGG G GGG+GGG G
Sbjct: 20 GGIGGGGG---AGGGFGGGHGGG-----VGGGIGGGAGGGG---GAGGGFGGGAGGG--- 157
Query: 137 TTTGGGYGGGYGDSDTKTTTGGGYG 161
GGG+GGG G GGG+G
Sbjct: 158 --AGGGFGGGAGGG-----AGGGFG 211
Score = 57.8 bits (138), Expect = 5e-09
Identities = 36/83 (43%), Positives = 41/83 (49%), Gaps = 1/83 (1%)
Frame = +2
Query: 80 GSGGAKTTTGGGYGGGYGDS-DTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTT 138
G GGA GGG+GGG G GGG GGG+G GGG GGG+G
Sbjct: 32 GGGGAGGGFGGGHGGGVGGGIGGGAGGGGGAGGGFGGG-----AGGGAGGGFGGG----- 181
Query: 139 TGGGYGGGYGDSDTKTTTGGGYG 161
GGG GGG+G GGG+G
Sbjct: 182 AGGGAGGGFG-GGAGGGAGGGFG 247
Score = 56.6 bits (135), Expect = 1e-08
Identities = 40/111 (36%), Positives = 48/111 (43%), Gaps = 1/111 (0%)
Frame = +2
Query: 51 GNDNSGGDYETGYNKNTDNYSNNETSGGYGSG-GAKTTTGGGYGGGYGDSDTKTTTGGGY 109
G +GG + G+ GG G G G GGG GGG+G GGG
Sbjct: 32 GGGGAGGGFGGGHG------------GGVGGGIGGGAGGGGGAGGGFGGG-----AGGGA 160
Query: 110 GGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGY 160
GGG+G GGG GGG+G GGG GGG+G GGG+
Sbjct: 161 GGGFGGG-----AGGGAGGGFGGG-----AGGGAGGGFGG-------GGGF 262
Score = 53.1 bits (126), Expect = 1e-07
Identities = 32/73 (43%), Positives = 36/73 (48%)
Frame = +2
Query: 89 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 148
GGG GGG G GGG+GGG+G GGG GGG G GGG+GGG G
Sbjct: 5 GGGVGGGIGGGG---GAGGGFGGGHGGG-----VGGGIGGGAGGGG---GAGGGFGGGAG 151
Query: 149 DSDTKTTTGGGYG 161
GGG+G
Sbjct: 152 GG-----AGGGFG 175
>TC207679 similar to UP|Q7XJI7 (Q7XJI7) Glycine-rich protein TomR2, partial
(22%)
Length = 425
Score = 60.5 bits (145), Expect = 7e-10
Identities = 34/72 (47%), Positives = 38/72 (52%)
Frame = +3
Query: 77 GGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTK 136
GGYGSGG +GGGYGGG GGG GG G + GG +GGGYG
Sbjct: 24 GGYGSGGG---SGGGYGGG--------AAGGGGGGSGGGGGAGSGGGGAHGGGYGGG--- 161
Query: 137 TTTGGGYGGGYG 148
GGG GGG+G
Sbjct: 162 --AGGGEGGGHG 191
Score = 55.5 bits (132), Expect = 2e-08
Identities = 32/73 (43%), Positives = 36/73 (48%)
Frame = +3
Query: 89 GGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 148
GG + GGYG +GGG GGGYG GG GGG S GGGYGGG G
Sbjct: 9 GGAHEGGYG-------SGGGSGGGYGGGAAGGGGGGSGGGGGAGSGGGGAHGGGYGGGAG 167
Query: 149 DSDTKTTTGGGYG 161
+ GGG+G
Sbjct: 168 GGE-----GGGHG 191
Score = 53.9 bits (128), Expect = 7e-08
Identities = 32/76 (42%), Positives = 37/76 (48%)
Frame = +3
Query: 55 SGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGGGYGDSDTKTTTGGGYGGGYG 114
+GG +E GY S + GGYG G A GG GGG S GGGYGGG G
Sbjct: 6 AGGAHEGGYG------SGGGSGGGYGGGAAGGGGGGSGGGGGAGSGGGGAHGGGYGGGAG 167
Query: 115 DSDTKTTTGGGYGGGY 130
+ GGG+GG Y
Sbjct: 168 GGE-----GGGHGGYY 200
>TC208565 similar to GB|AAM97131.1|22531255|AY136466 ribonucleoprotein-like
{Arabidopsis thaliana;} , partial (34%)
Length = 936
Score = 60.1 bits (144), Expect = 1e-09
Identities = 43/124 (34%), Positives = 48/124 (38%), Gaps = 15/124 (12%)
Frame = +2
Query: 53 DNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTGGGYGG------GYGD-------- 98
++S Y GY D + N GGY SGGA G YGG GYG
Sbjct: 194 NDSRSSYGGGYGDAYDGFGGNFGMGGYRSGGAYGGRGSAYGGFGSEFGGYGGYAGAMGPY 373
Query: 99 -SDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTG 157
D G YGGGYG + GGYGG GGG G S + G
Sbjct: 374 RGDPSLGYAGRYGGGYG----RGYDLGGYGGPSDGYGAYGGVGGGSSGSAYGSSYDASLG 541
Query: 158 GGYG 161
GGYG
Sbjct: 542 GGYG 553
Score = 39.3 bits (90), Expect = 0.002
Identities = 34/99 (34%), Positives = 36/99 (36%)
Frame = +2
Query: 30 GYNKTSSYSTGDDDYNKKTSYGNDNSGGDYETGYNKNTDNYSNNETSGGYGSGGAKTTTG 89
GY GD YG G GY +D Y G YG G G
Sbjct: 347 GYAGAMGPYRGDPSLGYAGRYGGGYGRGYDLGGYGGPSDGY------GAYGGVG-----G 493
Query: 90 GGYGGGYGDSDTKTTTGGGYGGGYGDSDTKTTTGGGYGG 128
G G YG S + GGGYGG G S T GGY G
Sbjct: 494 GSSGSAYG-SSYDASLGGGYGGPAGGSFYGTR--GGYAG 601
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.303 0.131 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,660,273
Number of Sequences: 63676
Number of extensions: 180941
Number of successful extensions: 6884
Number of sequences better than 10.0: 493
Number of HSP's better than 10.0 without gapping: 1615
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3448
length of query: 257
length of database: 12,639,632
effective HSP length: 95
effective length of query: 162
effective length of database: 6,590,412
effective search space: 1067646744
effective search space used: 1067646744
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC125473.10


