

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
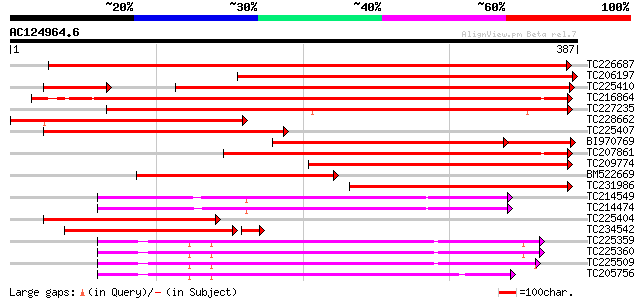
Query= AC124964.6 - phase: 0
(387 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC226687 homologue to UP|MMK2_MEDSA (Q40353) Mitogen-activated p... 555 e-158
TC206197 homologue to UP|MAPK_PEA (Q06060) Mitogen-activated pro... 470 e-133
TC225410 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, partial (... 455 e-128
TC216864 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, com... 422 e-118
TC227235 homologue to UP|Q6TAR9 (Q6TAR9) Mitogen activated prote... 340 6e-94
TC228662 homologue to UP|MAPK_PEA (Q06060) Mitogen-activated pro... 311 2e-85
TC225407 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, partial (... 287 5e-78
BI970769 similar to GP|4456682|emb MAP kinase {Medicago sativa},... 246 2e-76
TC207861 similar to UP|MPK7_ARATH (Q39027) Mitogen-activated pro... 280 8e-76
TC209774 similar to UP|Q84XZ6 (Q84XZ6) Mitogen-activated protein... 261 5e-70
BM522669 251 5e-67
TC231986 similar to UP|MMK2_MEDSA (Q40353) Mitogen-activated pro... 213 2e-55
TC214549 homologue to UP|O82135 (O82135) Cdc2, complete 206 2e-53
TC214474 homologue to UP|CDC2_VIGAC (Q41639) Cell division contr... 202 2e-52
TC225404 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, partial (... 198 3e-51
TC234542 similar to UP|Q8GZR5 (Q8GZR5) Mitogen-activated protein... 181 2e-50
TC225359 homologue to UP|MSK1_MEDSA (P51137) Glycogen synthase k... 190 1e-48
TC225360 homologue to UP|MSK1_MEDSA (P51137) Glycogen synthase k... 188 3e-48
TC225509 similar to UP|KSG7_ARATH (Q39011) Shaggy-related protei... 187 5e-48
TC205756 homologue to UP|Q8H6X8 (Q8H6X8) GSK-3-like protein MsK4... 187 7e-48
>TC226687 homologue to UP|MMK2_MEDSA (Q40353) Mitogen-activated protein
kinase homolog MMK2 , partial (97%)
Length = 1783
Score = 555 bits (1429), Expect = e-158
Identities = 255/358 (71%), Positives = 314/358 (87%), Gaps = 1/358 (0%)
Frame = +3
Query: 27 NIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKI 86
NI +HGGR++QYNI+GN+FEV+ KY PPI P+G+GAYGIVC+A N+ET E VA+KKI
Sbjct: 258 NIRGVPTHGGRYVQYNIYGNLFEVSRKYVPPIRPVGRGAYGIVCAAVNAETGEEVAIKKI 437
Query: 87 ANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQI 146
NAFDN+IDAKRTLREIKLLRHMDH N+++I+DI+ PPQ+E FNDVY+ ELMDTDLHQI
Sbjct: 438 GNAFDNRIDAKRTLREIKLLRHMDHANIMSIKDIIRPPQKENFNDVYLVSELMDTDLHQI 617
Query: 147 IRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVT 206
IRSNQ L+++HC+YFLYQ+LRGLKY+HSANVLHRDLKPSNLLLNANCDLKI DFGLAR T
Sbjct: 618 IRSNQQLTDDHCRYFLYQLLRGLKYVHSANVLHRDLKPSNLLLNANCDLKIADFGLARTT 797
Query: 207 SETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLR 266
SETDFMTEYVVTRWYRAPELLLN S+YTAAID+WSVGCI E++ R+PLFPG+D+VHQLR
Sbjct: 798 SETDFMTEYVVTRWYRAPELLLNCSEYTAAIDIWSVGCILGEIITRQPLFPGKDYVHQLR 977
Query: 267 LLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLVEKMLTFDP 325
L+ ELIG+P + LGFL ++NA+RY++QLP Y +Q+F +FP + P A+DL+EKML FDP
Sbjct: 978 LITELIGSPDDASLGFLRSDNARRYVKQLPQYPKQNFSARFPTMSPGAVDLLEKMLIFDP 1157
Query: 326 RKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYREALAFNP 383
+RITV++AL+HPY++ LHDI++EPVC PFSFDFEQ + TEE +KELI+RE++ FNP
Sbjct: 1158NRRITVDEALSHPYMSPLHDINEEPVCTRPFSFDFEQPSFTEEDIKELIWRESVKFNP 1331
>TC206197 homologue to UP|MAPK_PEA (Q06060) Mitogen-activated protein kinase
homolog D5 , partial (59%)
Length = 940
Score = 470 bits (1210), Expect = e-133
Identities = 228/232 (98%), Positives = 229/232 (98%)
Frame = +1
Query: 156 EHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVTSETDFMTEY 215
EHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVTSETDFMTEY
Sbjct: 1 EHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVTSETDFMTEY 180
Query: 216 VVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTP 275
VVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTP
Sbjct: 181 VVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTP 360
Query: 276 SEDDLGFLNENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDAL 335
SE DLGFLNENAKRYIRQLP YRRQSFQEKFP VHPEAIDLVEKMLTFDPRKRITVEDAL
Sbjct: 361 SEADLGFLNENAKRYIRQLPLYRRQSFQEKFPHVHPEAIDLVEKMLTFDPRKRITVEDAL 540
Query: 336 AHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYREALAFNPEYQQ 387
AHPYLTSLHDISDEPVC+TPFSFDFEQHALTEEQMKELIYREALAFNPEYQQ
Sbjct: 541 AHPYLTSLHDISDEPVCLTPFSFDFEQHALTEEQMKELIYREALAFNPEYQQ 696
>TC225410 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, partial (88%)
Length = 1375
Score = 455 bits (1171), Expect = e-128
Identities = 217/273 (79%), Positives = 242/273 (88%), Gaps = 1/273 (0%)
Frame = +3
Query: 114 VVAIRDIVPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIH 173
V+ +RD++PPP R FNDVYIA ELMDTDLH IIRSNQ LSEEHCQYFLYQILRGLKYIH
Sbjct: 171 VIGLRDVIPPPLRREFNDVYIATELMDTDLHHIIRSNQNLSEEHCQYFLYQILRGLKYIH 350
Query: 174 SANVLHRDLKPSNLLLNANCDLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDY 233
SANV+HRDLKPSNLLLN+NCDLKI DFGLAR T E+DFMTEYVVTRWYRAPELLLNSSDY
Sbjct: 351 SANVIHRDLKPSNLLLNSNCDLKIIDFGLARPTLESDFMTEYVVTRWYRAPELLLNSSDY 530
Query: 234 TAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIR 292
T+AIDVWSVGCIFMELM++KPLFPG+DHVHQ+RLL EL+GTP+E DLG + NE+A+RYIR
Sbjct: 531 TSAIDVWSVGCIFMELMNKKPLFPGKDHVHQMRLLTELLGTPTEADLGLVKNEDARRYIR 710
Query: 293 QLPPYRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVC 352
QLP Y RQ + FP VHP AIDLV+KMLT DP KRITVE+ALAHPYL LHD++DEP+C
Sbjct: 711 QLPQYPRQPLAQVFPHVHPAAIDLVDKMLTVDPTKRITVEEALAHPYLEKLHDVADEPIC 890
Query: 353 MTPFSFDFEQHALTEEQMKELIYREALAFNPEY 385
M PFSFDFEQ L EEQ+KE+IYREALA NPEY
Sbjct: 891 MEPFSFDFEQQQLDEEQIKEMIYREALALNPEY 989
Score = 84.0 bits (206), Expect = 1e-16
Identities = 35/46 (76%), Positives = 43/46 (93%)
Frame = +1
Query: 24 GIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIV 69
G+ + PAT +HGG+FIQYNIFGN+FEVTAKY+PPIMP+G+GAYGIV
Sbjct: 31 GVADFPATPTHGGQFIQYNIFGNLFEVTAKYRPPIMPVGRGAYGIV 168
>TC216864 homologue to UP|Q9M6R8 (Q9M6R8) MAP kinase PsMAPK2, complete
Length = 1735
Score = 422 bits (1086), Expect = e-118
Identities = 207/371 (55%), Positives = 275/371 (73%), Gaps = 2/371 (0%)
Frame = +2
Query: 16 AAPAPPQMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNS 75
A P P GI + G + Y+++ +FE+ +KY P I PIG+GAYGIVCS+ N
Sbjct: 302 ATPVEPPNGIR------TEGKHY--YSMWQTLFEIDSKYVP-IKPIGRGAYGIVCSSVNR 454
Query: 76 ETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIA 135
E NE VA+KKI NAF+N++DA RTLRE+KLLRH+ HENV+A++DI+ P R F DVY+
Sbjct: 455 EINEKVAIKKIQNAFENRVDALRTLRELKLLRHLHHENVIALKDIMMPVHRNSFKDVYLV 634
Query: 136 YELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDL 195
YELMDTDLHQII+S+QALS +HCQYFL+Q+LRGLKY+HSAN+LHRDLKP NLL+NANCDL
Sbjct: 635 YELMDTDLHQIIKSSQALSNDHCQYFLFQLLRGLKYLHSANILHRDLKPGNLLINANCDL 814
Query: 196 KICDFGLARVT-SETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKP 254
KICDFGLAR S+ FMTEYVVTRWYRAPELLL +Y +IDVWSVGCIF EL+ RKP
Sbjct: 815 KICDFGLARTNCSKNQFMTEYVVTRWYRAPELLLCCDNYGTSIDVWSVGCIFAELLGRKP 994
Query: 255 LFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEA 313
+FPG + ++QL+L++ ++G+ E+D+ F+ N AK+YI+ LP + +P HP A
Sbjct: 995 IFPGSECLNQLKLIINILGSQREEDIEFIDNPKAKKYIKSLPYSPGTPLSQLYPNAHPLA 1174
Query: 314 IDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKEL 373
IDL+ KML FDP KRI+V AL HPY+ L+D + +P + P D ++ L EE ++++
Sbjct: 1175IDLLAKMLVFDPTKRISVTQALQHPYMAPLYDPNCDPPAVIPIDLDIDED-LGEEMIRDM 1351
Query: 374 IYREALAFNPE 384
+++E L ++PE
Sbjct: 1352MWKEMLHYHPE 1384
>TC227235 homologue to UP|Q6TAR9 (Q6TAR9) Mitogen activated protein kinase 6,
partial (72%)
Length = 2041
Score = 340 bits (872), Expect = 6e-94
Identities = 169/326 (51%), Positives = 231/326 (70%), Gaps = 8/326 (2%)
Frame = +3
Query: 67 GIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQR 126
G+VCSA+++ T E VA+KKI + F++ DA R LREIKLLR + H ++V I+ I+ PP R
Sbjct: 3 GVVCSAYDTHTGEKVAIKKINDIFEHVSDATRILREIKLLRLLRHPDIVEIKHILLPPSR 182
Query: 127 EVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSN 186
F D+Y+ +ELM++DLHQ+I++N L+ EH Q+FLYQ+LRG+KYIH+ANV HRDLKP N
Sbjct: 183 REFKDIYVVFELMESDLHQVIKANDDLTPEHYQFFLYQLLRGMKYIHTANVFHRDLKPKN 362
Query: 187 LLLNANCDLKICDFGLARV----TSETDFMTEYVVTRWYRAPELLLN-SSDYTAAIDVWS 241
+L NA+C LKICDFGLARV T F T+YV TRWYRAPEL + S YT AID+WS
Sbjct: 363 ILANADCKLKICDFGLARVAFNDTPTAIFWTDYVATRWYRAPELCGSFFSKYTPAIDIWS 542
Query: 242 VGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQ 300
+GCIF EL+ KPLFPG++ VHQL L+ +L+GTPS + + + NE A+RY+ + +
Sbjct: 543 IGCIFAELLTGKPLFPGKNVVHQLDLMTDLLGTPSLEAIARVRNEKARRYLSSMRKKKPV 722
Query: 301 SFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVC--MTPFSF 358
+KFP P A+ L+EKML F+P+ R T E+ALA PY L + EP +T F
Sbjct: 723 PLSQKFPNADPLALRLLEKMLAFEPKDRPTAEEALADPYFKGLAKVEREPSAQPVTKMEF 902
Query: 359 DFEQHALTEEQMKELIYREALAFNPE 384
+FE+ +T+E ++ELIYRE L ++P+
Sbjct: 903 EFERRRITKEDVRELIYREILEYHPK 980
>TC228662 homologue to UP|MAPK_PEA (Q06060) Mitogen-activated protein kinase
homolog D5 , partial (42%)
Length = 679
Score = 311 bits (798), Expect = 2e-85
Identities = 156/166 (93%), Positives = 158/166 (94%), Gaps = 4/166 (2%)
Frame = +1
Query: 1 MEGGGA-PPADTVMSDAAPAPPQ---MGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKP 56
MEGGGA PPADTVMSDAAP P Q MGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKP
Sbjct: 181 MEGGGAAPPADTVMSDAAPPPQQAMAMGIENIPATLSHGGRFIQYNIFGNIFEVTAKYKP 360
Query: 57 PIMPIGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVA 116
PIMPIGKGAYGIVCSA NSETNEHVA+KKIANAFDNKIDAKRTLREIKLLRHMDHENVVA
Sbjct: 361 PIMPIGKGAYGIVCSALNSETNEHVAIKKIANAFDNKIDAKRTLREIKLLRHMDHENVVA 540
Query: 117 IRDIVPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQYFL 162
IRDIVPPPQRE+FNDVYIAYELMDTDLHQIIRSNQ LSEEHCQYFL
Sbjct: 541 IRDIVPPPQREIFNDVYIAYELMDTDLHQIIRSNQGLSEEHCQYFL 678
>TC225407 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, partial (47%)
Length = 686
Score = 287 bits (735), Expect = 5e-78
Identities = 136/167 (81%), Positives = 152/167 (90%)
Frame = +2
Query: 24 GIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAV 83
G+ + PAT +HGG+FIQYNIFGN+FEVTAKY+PPIMP+G+GAYGIVCS N+ETNE VAV
Sbjct: 185 GVADFPATPTHGGQFIQYNIFGNLFEVTAKYRPPIMPVGRGAYGIVCSLLNTETNELVAV 364
Query: 84 KKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDL 143
KKIANAFDN +DAKRTLREIKLLRH+DHENV+ +RD++PPP R FNDVYIA ELMDTDL
Sbjct: 365 KKIANAFDNHMDAKRTLREIKLLRHLDHENVIGLRDVIPPPLRREFNDVYIATELMDTDL 544
Query: 144 HQIIRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLN 190
H IIRSNQ LSEEH QYFLYQILRGLKYIHSANV+HRDLKPSNLLLN
Sbjct: 545 HHIIRSNQNLSEEHSQYFLYQILRGLKYIHSANVIHRDLKPSNLLLN 685
>BI970769 similar to GP|4456682|emb MAP kinase {Medicago sativa}, partial
(54%)
Length = 725
Score = 246 bits (628), Expect(2) = 2e-76
Identities = 116/162 (71%), Positives = 139/162 (85%), Gaps = 1/162 (0%)
Frame = -2
Query: 180 RDLKPSNLLLNANCDLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDV 239
RDL S LLNANCDLKICDFGLAR TSETDFMTEYVVTRWYRAPELLLN S+YTAAID+
Sbjct: 721 RDLXXSXXLLNANCDLKICDFGLARTTSETDFMTEYVVTRWYRAPELLLNCSEYTAAIDI 542
Query: 240 WSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYR 298
WSVGCI ME++ R+PLFPG+D+V QL L+ EL+G+P++ DLGFL ++NAK+Y++QLP
Sbjct: 541 WSVGCILMEIVRREPLFPGKDYVQQLALITELLGSPNDSDLGFLRSDNAKKYVKQLPHVE 362
Query: 299 RQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYL 340
+QSF E+FP++ P AIDL EKML FDP KRITVE+AL HPY+
Sbjct: 361 KQSFAERFPEMSPLAIDLAEKMLVFDPSKRITVEEALNHPYM 236
Score = 57.8 bits (138), Expect(2) = 2e-76
Identities = 25/49 (51%), Positives = 35/49 (71%)
Frame = -3
Query: 338 PYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYREALAFNPEYQ 386
P SLH+I++EP C TPF F FE L EE +KELI++E+L F+ ++Q
Sbjct: 243 PICASLHEINEEPTCPTPFIFSFEXXILKEEDIKELIWKESLNFSQDHQ 97
>TC207861 similar to UP|MPK7_ARATH (Q39027) Mitogen-activated protein kinase
homolog 7 (MAP kinase 7) (AtMPK7) , partial (67%)
Length = 911
Score = 280 bits (716), Expect = 8e-76
Identities = 134/240 (55%), Positives = 179/240 (73%), Gaps = 2/240 (0%)
Frame = +2
Query: 147 IRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVT 206
I+S+Q LS +HC+YFL+Q+LRGLKY+HSAN+LHRDLKP NLL+NANCDLKICDFGLAR
Sbjct: 2 IKSSQPLSNDHCKYFLFQLLRGLKYLHSANILHRDLKPGNLLVNANCDLKICDFGLARTN 181
Query: 207 S-ETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQL 265
+ FMTEYVVTRWYRAPELLL +Y +IDVWSVGCIF E++ RKP+FPG + ++QL
Sbjct: 182 GVDGQFMTEYVVTRWYRAPELLLCCDNYGTSIDVWSVGCIFAEILGRKPIFPGTECLNQL 361
Query: 266 RLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLVEKMLTFD 324
+L++ ++G+ E L F+ N A+R+I+ LP R + F + +PQ P AIDL++KML FD
Sbjct: 362 KLIISVLGSQHESHLEFIDNAKARRFIKSLPYTRGRHFSQLYPQADPLAIDLLQKMLVFD 541
Query: 325 PRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYREALAFNPE 384
P KRITV +AL HPY+ SL+D +P P S D ++H E ++E+ + E L ++PE
Sbjct: 542 PTKRITVLEALQHPYMASLYDPRCDPPAQVPISLDIDEH-WGEPMIREMFWNEMLHYHPE 718
>TC209774 similar to UP|Q84XZ6 (Q84XZ6) Mitogen-activated protein kinase 4,
partial (48%)
Length = 874
Score = 261 bits (666), Expect = 5e-70
Identities = 120/181 (66%), Positives = 152/181 (83%), Gaps = 1/181 (0%)
Frame = +2
Query: 205 VTSETDFMTEYVVTRWYRAPELLLNSSDYTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQ 264
+ +TDFMTEYVVTRWYRAPELLLN S+YT+AIDVWSVGCI E+M R+PLFPG+D+VHQ
Sbjct: 2 IRHQTDFMTEYVVTRWYRAPELLLNCSEYTSAIDVWSVGCILGEIMTREPLFPGKDYVHQ 181
Query: 265 LRLLMELIGTPSEDDLGFL-NENAKRYIRQLPPYRRQSFQEKFPQVHPEAIDLVEKMLTF 323
LRL+ EL+G+P + L FL ++NA+RYIRQLP YR+Q F +FP + PEA+DL+EKML F
Sbjct: 182 LRLITELLGSPDDASLEFLRSDNARRYIRQLPQYRKQKFSARFPNMLPEALDLLEKMLIF 361
Query: 324 DPRKRITVEDALAHPYLTSLHDISDEPVCMTPFSFDFEQHALTEEQMKELIYREALAFNP 383
DP KRITV++AL HPYL+SLH+I+DEPVC PFSFDF+Q EE +KELI++E++ FNP
Sbjct: 362 DPNKRITVDEALCHPYLSSLHNINDEPVCPRPFSFDFDQPTCAEEHVKELIWKESVKFNP 541
Query: 384 E 384
+
Sbjct: 542 D 544
>BM522669
Length = 442
Score = 251 bits (640), Expect = 5e-67
Identities = 115/138 (83%), Positives = 129/138 (93%)
Frame = +3
Query: 87 ANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQI 146
A FDN IDAKRTLREIKLLRHMDHEN++AIRDI+ PP+++ FNDVYI YELMDTDLHQI
Sbjct: 27 ARGFDNIIDAKRTLREIKLLRHMDHENIIAIRDIIRPPRKDAFNDVYIVYELMDTDLHQI 206
Query: 147 IRSNQALSEEHCQYFLYQILRGLKYIHSANVLHRDLKPSNLLLNANCDLKICDFGLARVT 206
IRS+Q L+++HCQYFLYQ+LRGLKY+HSAN+LHRDLKPSNLLLN+NCDLKI DFGLAR T
Sbjct: 207 IRSDQPLNDDHCQYFLYQLLRGLKYVHSANILHRDLKPSNLLLNSNCDLKIADFGLARTT 386
Query: 207 SETDFMTEYVVTRWYRAP 224
SETDFMTEYVVTRWYRAP
Sbjct: 387 SETDFMTEYVVTRWYRAP 440
>TC231986 similar to UP|MMK2_MEDSA (Q40353) Mitogen-activated protein kinase
homolog MMK2 , partial (42%)
Length = 710
Score = 213 bits (541), Expect = 2e-55
Identities = 97/153 (63%), Positives = 125/153 (81%), Gaps = 1/153 (0%)
Frame = +1
Query: 233 YTAAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFL-NENAKRYI 291
YTAAID+WSVGCI E++ R+PLF G+D+VHQLRL+ ELIG+P + LGFL ++NA+RY+
Sbjct: 1 YTAAIDIWSVGCILGEIITRQPLFLGKDYVHQLRLITELIGSPDDPSLGFLRSDNARRYV 180
Query: 292 RQLPPYRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPV 351
RQLP Y RQ F +FP + P A+DL+EKML FDP +RITVE+AL HPYL LHDI++EP
Sbjct: 181 RQLPQYPRQQFAARFPSMSPGAVDLLEKMLVFDPNRRITVEEALCHPYLAPLHDINEEPA 360
Query: 352 CMTPFSFDFEQHALTEEQMKELIYREALAFNPE 384
C+ PFSFDFEQ + TEE +KELI+RE++ FNP+
Sbjct: 361 CVRPFSFDFEQPSFTEEDIKELIWRESVLFNPD 459
>TC214549 homologue to UP|O82135 (O82135) Cdc2, complete
Length = 1216
Score = 206 bits (523), Expect = 2e-53
Identities = 111/287 (38%), Positives = 168/287 (57%), Gaps = 4/287 (1%)
Frame = +3
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IG+G YG+V + TNE +A+KKI +++ +REI LL+ M H N+V ++D+
Sbjct: 183 IGEGTYGVVYKGRDRVTNETIALKKIRLEQEDEGVPSTAIREISLLKEMQHRNIVRLQDV 362
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQY--FLYQILRGLKYIHSANVL 178
V + +Y+ +E +D DL + + S+ +++ Q FLYQIL G+ Y HS VL
Sbjct: 363 VHDEK-----SLYLVFEYLDLDLKKHMDSSPEFAKDPRQVKMFLYQILCGIAYCHSHRVL 527
Query: 179 HRDLKPSNLLLNANCD-LKICDFGLARVTS-ETDFMTEYVVTRWYRAPELLLNSSDYTAA 236
HRDLKP NLL++ + + LK+ DFGLAR T VVT WYRAPE+LL S Y+
Sbjct: 528 HRDLKPQNLLIDRSTNALKLADFGLARAFGIPVRTFTHEVVTLWYRAPEILLGSRQYSTP 707
Query: 237 IDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPP 296
+D+WSVGCIF E+++++PLFPG + +L + ++GTP+ED + + + P
Sbjct: 708 VDIWSVGCIFAEMVNQRPLFPGDSEIDELFKIFRIMGTPNEDTWPGVT-SLPDFKSAFPK 884
Query: 297 YRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSL 343
++ + + P + P +DL+ ML DP KRIT AL H Y +
Sbjct: 885 WQPKDLKNVVPNLEPAGLDLLSSMLYLDPSKRITARSALEHEYFKDI 1025
>TC214474 homologue to UP|CDC2_VIGAC (Q41639) Cell division control protein 2
homolog (p34cdc2) , complete
Length = 1204
Score = 202 bits (514), Expect = 2e-52
Identities = 112/287 (39%), Positives = 165/287 (57%), Gaps = 4/287 (1%)
Frame = +2
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
IG+G YG+V A + TNE +A+KKI +++ +REI LL+ M H N+V ++D+
Sbjct: 107 IGEGTYGVVYKARDRATNETIALKKIRLEQEDEGVPSTAIREISLLKEMQHRNIVRLQDV 286
Query: 121 VPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSEEHCQY--FLYQILRGLKYIHSANVL 178
V +R +Y+ +E +D DL + + S+ ++ Q FLYQIL G+ Y HS VL
Sbjct: 287 VHSEKR-----LYLVFEYLDLDLKKHMDSSPEFVKDPRQVKMFLYQILCGIAYCHSHRVL 451
Query: 179 HRDLKPSNLLLNANCD-LKICDFGLARVTS-ETDFMTEYVVTRWYRAPELLLNSSDYTAA 236
HRDLKP NLL++ + LK+ DFGLAR T VVT WYRAPE+LL S Y+
Sbjct: 452 HRDLKPQNLLIDRRTNSLKLADFGLARAFGIPVRTFTHEVVTLWYRAPEILLGSRHYSTP 631
Query: 237 IDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQLPP 296
+DVWSVGCIF E+++R+PLFPG + +L + ++GTP+ED + + + P
Sbjct: 632 VDVWSVGCIFAEMVNRRPLFPGDSEIDELFKIFRILGTPNEDTWPGVT-SLPDFKSTFPK 808
Query: 297 YRRQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSL 343
+ + P + ++L+ ML DP KRIT A+ H Y +
Sbjct: 809 WPSKDLANVVPNLDAAGLNLLSSMLCLDPSKRITARSAVEHEYFKDI 949
>TC225404 homologue to UP|Q9M6S1 (Q9M6S1) MAP kinase 3, partial (35%)
Length = 473
Score = 198 bits (504), Expect = 3e-51
Identities = 92/121 (76%), Positives = 106/121 (87%)
Frame = +2
Query: 24 GIENIPATLSHGGRFIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAV 83
G+ + A +HGG+FIQYNIFGN+FEVT KY+PPIMPIG+GAYGIVCS N+ETNE VAV
Sbjct: 107 GVADFAAVPTHGGQFIQYNIFGNLFEVTTKYRPPIMPIGRGAYGIVCSLLNTETNELVAV 286
Query: 84 KKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDL 143
KKIANAFDN +DAKRTLREIKLLRH+DHENV+ +RD++PPP R FNDVYIA ELMDTDL
Sbjct: 287 KKIANAFDNHMDAKRTLREIKLLRHLDHENVIGLRDVIPPPLRREFNDVYIATELMDTDL 466
Query: 144 H 144
H
Sbjct: 467 H 469
>TC234542 similar to UP|Q8GZR5 (Q8GZR5) Mitogen-activated protein kinase,
partial (32%)
Length = 552
Score = 181 bits (460), Expect(2) = 2e-50
Identities = 82/118 (69%), Positives = 105/118 (88%)
Frame = +3
Query: 38 FIQYNIFGNIFEVTAKYKPPIMPIGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAK 97
++QYN++GN+FEV++ Y PPI PIG+GAYGIVC+A N +T+E VA+KKI NAF+N IDAK
Sbjct: 114 YVQYNVYGNLFEVSSNYVPPIRPIGRGAYGIVCAAVNCDTHEEVAIKKIGNAFNNIIDAK 293
Query: 98 RTLREIKLLRHMDHENVVAIRDIVPPPQREVFNDVYIAYELMDTDLHQIIRSNQALSE 155
RTLREIKLLRHMD EN++AIRDI+ PP+++ F+DVYI YELMDTDLHQIIRS+Q L++
Sbjct: 294 RTLREIKLLRHMDLENIIAIRDIIRPPRKDAFDDVYIVYELMDTDLHQIIRSDQPLND 467
Score = 36.2 bits (82), Expect(2) = 2e-50
Identities = 14/16 (87%), Positives = 16/16 (99%)
Frame = +2
Query: 159 QYFLYQILRGLKYIHS 174
QYFLYQ+LRGLKY+HS
Sbjct: 503 QYFLYQLLRGLKYVHS 550
>TC225359 homologue to UP|MSK1_MEDSA (P51137) Glycogen synthase kinase-3
homolog MsK-1 , complete
Length = 1668
Score = 190 bits (482), Expect = 1e-48
Identities = 110/314 (35%), Positives = 180/314 (57%), Gaps = 9/314 (2%)
Frame = +2
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
+G G++G+V A ET E VA+KK+ D + RE++ +R +DH NVVA++
Sbjct: 359 VGHGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVALKHC 520
Query: 121 V--PPPQREVFNDVYIAY--ELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSA- 175
+ E++ ++ + Y E ++ + + NQ + + + + YQI R L YIH
Sbjct: 521 FFSTTEKDELYLNLVLEYVPETVNRVIKHYNKLNQRMPLIYVKLYTYQIFRALSYIHRCI 700
Query: 176 NVLHRDLKPSNLLLNANC-DLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYT 234
V HRD+KP NLL+N + +K+CDFG A+V + + Y+ +R+YRAPEL+ +++YT
Sbjct: 701 GVCHRDIKPQNLLVNPHTHQVKLCDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT 880
Query: 235 AAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQL 294
AIDVWSVGC+ EL+ +PLFPG V QL +++++GTP+ +++ +N N + +
Sbjct: 881 TAIDVWSVGCVLAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEF--KF 1054
Query: 295 PPYRRQSFQEKF-PQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDE--PV 351
P + + + F ++ PEA+DLV ++L + P R T DAL HP+ L D +
Sbjct: 1055PQIKAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRCTALDALTHPFFDELRDPNSRLPNG 1234
Query: 352 CMTPFSFDFEQHAL 365
P F+F+ H L
Sbjct: 1235RFLPPLFNFKSHEL 1276
>TC225360 homologue to UP|MSK1_MEDSA (P51137) Glycogen synthase kinase-3
homolog MsK-1 , partial (97%)
Length = 1594
Score = 188 bits (478), Expect = 3e-48
Identities = 110/314 (35%), Positives = 178/314 (56%), Gaps = 9/314 (2%)
Frame = +1
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
+G G++G+V A ET E VA+KK+ D + RE++ +R +DH NVV ++
Sbjct: 394 VGHGSFGVVFQAKCLETGETVAIKKVLQ------DKRYKNRELQTMRLLDHPNVVCLKHC 555
Query: 121 V--PPPQREVFNDVYIAY--ELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSA- 175
+ E++ ++ + Y E ++ + + NQ + + + + YQI R L YIH
Sbjct: 556 FFSTTEKDELYLNLVLEYVPETVNRVIKHYNKLNQRMPLIYVKLYTYQIFRALSYIHRCI 735
Query: 176 NVLHRDLKPSNLLLNANC-DLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYT 234
V HRD+KP NLL+N + +KICDFG A+V + + Y+ +R+YRAPEL+ +++YT
Sbjct: 736 GVCHRDIKPQNLLVNPHTHQVKICDFGSAKVLVKGEPNISYICSRYYRAPELIFGATEYT 915
Query: 235 AAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQL 294
AIDVWSVGC+ ELM +PLFPG V QL +++++GTP+ +++ +N N + +
Sbjct: 916 TAIDVWSVGCVLAELMLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEF--KF 1089
Query: 295 PPYRRQSFQEKF-PQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDE--PV 351
P + + + F ++ PEA+DLV ++L + P R T D L HP+ L D +
Sbjct: 1090PQIKAHPWHKIFHKRMPPEAVDLVSRLLQYSPNLRCTALDTLTHPFFDELRDPNTRLPNG 1269
Query: 352 CMTPFSFDFEQHAL 365
P F+F+ H L
Sbjct: 1270RFLPPLFNFKSHEL 1311
>TC225509 similar to UP|KSG7_ARATH (Q39011) Shaggy-related protein kinase eta
(ASK-eta) , partial (98%)
Length = 1755
Score = 187 bits (476), Expect = 5e-48
Identities = 110/311 (35%), Positives = 181/311 (57%), Gaps = 9/311 (2%)
Frame = +1
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
+G G++GIV A ET E VA+KK+ D + RE++L+R MDH NV++++
Sbjct: 292 VGTGSFGIVFQAKCLETGEAVAIKKVLQ------DRRYKNRELQLMRVMDHPNVISLKHC 453
Query: 121 V--PPPQREVFNDVYIAY--ELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSA- 175
E+F ++ + Y E M L +NQ + + + ++YQI RGL YIH+
Sbjct: 454 FFSTTSTDELFLNLVMEYVPESMYRVLKHYSNANQRMPIIYVKLYMYQIFRGLAYIHTGP 633
Query: 176 NVLHRDLKPSNLLLNA-NCDLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYT 234
V HRDLKP N+L++ +K+CDFG A+V E + Y+ +R+YRAPEL+ +++YT
Sbjct: 634 KVCHRDLKPQNILVDPLTHQVKLCDFGSAKVLVEGEANISYICSRFYRAPELIFGATEYT 813
Query: 235 AAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRYIRQL 294
++ID+WS GC+ EL+ +PLFPG + V QL +++++GTP+ +++ +N N + +
Sbjct: 814 SSIDIWSAGCVLAELLLGQPLFPGENAVDQLVHIIKVLGTPTREEVRCMNPNYNDF--RF 987
Query: 295 PPYRRQSFQEKF-PQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHDISDEPVCM 353
P + + + F ++ PEAIDL ++L + P R T +A AHP+ L + +
Sbjct: 988 PQIKAHPWHKIFHKKMPPEAIDLASRLLQYSPSLRCTALEACAHPFFDELREPNARLPNG 1167
Query: 354 TPFS--FDFEQ 362
PF F+F+Q
Sbjct: 1168RPFPPLFNFKQ 1200
>TC205756 homologue to UP|Q8H6X8 (Q8H6X8) GSK-3-like protein MsK4, partial
(94%)
Length = 1778
Score = 187 bits (475), Expect = 7e-48
Identities = 104/293 (35%), Positives = 171/293 (57%), Gaps = 8/293 (2%)
Frame = +3
Query: 61 IGKGAYGIVCSAHNSETNEHVAVKKIANAFDNKIDAKRTLREIKLLRHMDHENVVAIRDI 120
+G G++G+V A ET E VA+KK+ D + RE+++++ +DH N+VA+R
Sbjct: 414 VGTGSFGVVFQAKCRETGEIVAIKKVLQ------DKRYKNRELQIMQMLDHPNIVALRHC 575
Query: 121 V--PPPQREVFNDVYIAY--ELMDTDLHQIIRSNQALSEEHCQYFLYQILRGLKYIHSA- 175
+ EV+ ++ + Y E ++ R NQ + + + + YQI R L YIH+
Sbjct: 576 FFSTTDKEEVYLNLVLEYVPETVNRIARSYSRINQRMPLIYVKLYTYQICRALAYIHNCI 755
Query: 176 NVLHRDLKPSNLLLNANC-DLKICDFGLARVTSETDFMTEYVVTRWYRAPELLLNSSDYT 234
+ HRD+KP NLL+N + LK+CDFG A+V + + Y+ +R+YRAPEL+ +++YT
Sbjct: 756 GICHRDIKPQNLLVNPHTHQLKLCDFGSAKVLVKGEPNVSYICSRYYRAPELIFGATEYT 935
Query: 235 AAIDVWSVGCIFMELMDRKPLFPGRDHVHQLRLLMELIGTPSEDDLGFLNENAKRY-IRQ 293
AID+WS GC+ EL+ +PLFPG V QL +++++GTP+ +++ +N N + Q
Sbjct: 936 TAIDIWSTGCVMAELLLGQPLFPGESGVDQLVEIIKVLGTPTREEIKCMNPNYTEFKFPQ 1115
Query: 294 LPPYR-RQSFQEKFPQVHPEAIDLVEKMLTFDPRKRITVEDALAHPYLTSLHD 345
+ P+ + FQ++ P PEA+DLV + + P R T +A HP+ L D
Sbjct: 1116IKPHPWHKVFQKRLP---PEAVDLVCRFFQYSPNLRCTALEACIHPFFDELRD 1265
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.415
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,303,671
Number of Sequences: 63676
Number of extensions: 246722
Number of successful extensions: 2117
Number of sequences better than 10.0: 688
Number of HSP's better than 10.0 without gapping: 1794
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1827
length of query: 387
length of database: 12,639,632
effective HSP length: 99
effective length of query: 288
effective length of database: 6,335,708
effective search space: 1824683904
effective search space used: 1824683904
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC124964.6


