

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123572.11 + phase: 0
(475 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
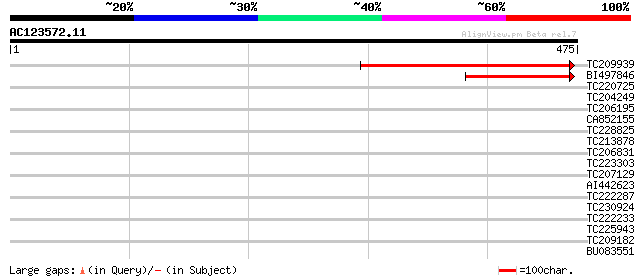
Score E
Sequences producing significant alignments: (bits) Value
TC209939 similar to UP|Q9LU81 (Q9LU81) Arabidopsis thaliana geno... 318 3e-87
BI497846 159 3e-39
TC220725 weakly similar to PIR|D88560|D88560 protein F58A4.1 [im... 34 0.16
TC204249 UP|Q9SB11 (Q9SB11) Glycinin, complete 30 1.8
TC206195 similar to UP|Q8GZT8 (Q8GZT8) PTEN-like protein, partia... 30 2.4
CA852155 similar to GP|15391731|em nitrite transporter {Cucumis ... 30 3.1
TC228825 weakly similar to UP|Q9V5S5 (Q9V5S5) CG33473-PB (Transc... 29 4.0
TC213878 homologue to UP|Q8SCS3 (Q8SCS3) PHIKZ239, partial (9%) 29 4.0
TC206831 similar to UP|Q96400 (Q96400) Nitrite transporter, part... 29 5.2
TC223303 similar to UP|Q8RV26 (Q8RV26) Oligopeptide transporter-... 29 5.2
TC207129 homologue to UP|Q84XV9 (Q84XV9) Phosphoribosylformylgly... 29 5.2
AI442623 29 5.2
TC222287 28 6.8
TC230924 UP|ATPH_LOTJA (P12231) ATP synthase C chain (Lipid-bin... 28 6.8
TC222233 similar to UP|Q6IMT1 (Q6IMT1) SAB, partial (7%) 28 6.8
TC225943 similar to UP|NU4C_LOTJA (Q9BBP3) NAD(P)H-quinone oxido... 28 6.8
TC209182 similar to PIR|T51854|T51854 RING-H2 finger protein RHF... 28 6.8
BU083551 homologue to GP|7301548|gb|A CG6066-PA {Drosophila mela... 28 8.9
>TC209939 similar to UP|Q9LU81 (Q9LU81) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MPE11, partial (49%)
Length = 871
Score = 318 bits (816), Expect = 3e-87
Identities = 150/179 (83%), Positives = 167/179 (92%)
Frame = +1
Query: 295 SEPISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFFDEVHELSKISEHDCVRRI 354
SEPISR+DPNEPLDKLLLNK+FRQSFM FADSCLAGESVHFFDEV+ELSKI E DCV+RI
Sbjct: 1 SEPISRIDPNEPLDKLLLNKRFRQSFMAFADSCLAGESVHFFDEVYELSKIPEDDCVKRI 180
Query: 355 YMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFHNALNEIVHLMKTNLAKDY 414
YMARHIIEKY+VAGAAME+NISHRS+QEILSTS+L R DLFHNALNEI+ LM+TNLA+DY
Sbjct: 181 YMARHIIEKYIVAGAAMEVNISHRSRQEILSTSNLTRPDLFHNALNEIIQLMRTNLARDY 360
Query: 415 WSSMFFLKFQEECDMRCNGYELEQMTGWNYSPRLSSVHGTDDPFHHDHLLKNSECGNDT 473
WSSMFFLKFQE+ ++R N YELEQ+TGWN+SPRLSSVHGTDDPFH DHLLK+S C NDT
Sbjct: 361 WSSMFFLKFQEDSNVRSNEYELEQITGWNFSPRLSSVHGTDDPFHQDHLLKSSGCSNDT 537
>BI497846
Length = 421
Score = 159 bits (401), Expect = 3e-39
Identities = 73/91 (80%), Positives = 80/91 (87%)
Frame = +1
Query: 383 ILSTSDLARADLFHNALNEIVHLMKTNLAKDYWSSMFFLKFQEECDMRCNGYELEQMTGW 442
ILSTS L R DLFHNALNEI+ LMKTNLA+DYWSSMFFLKFQE+ ++R N YELEQMTGW
Sbjct: 1 ILSTSSLTRPDLFHNALNEIIQLMKTNLARDYWSSMFFLKFQEDTNVRSNEYELEQMTGW 180
Query: 443 NYSPRLSSVHGTDDPFHHDHLLKNSECGNDT 473
N+SPRLSSVHG DDPFH DHLLK+S C NDT
Sbjct: 181 NFSPRLSSVHGIDDPFHQDHLLKSSGCSNDT 273
>TC220725 weakly similar to PIR|D88560|D88560 protein F58A4.1 [imported] -
Caenorhabditis elegans {Caenorhabditis elegans;} ,
partial (10%)
Length = 783
Score = 33.9 bits (76), Expect = 0.16
Identities = 18/44 (40%), Positives = 25/44 (55%)
Frame = -2
Query: 122 LYFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHIKKPLSNRCH 165
L +IFVK+ PL+ S LL P +LL + A ++H RCH
Sbjct: 569 LQWIFVKKTFPLLVSLLLSPFLLLTVVAAAFLLH-------RCH 459
>TC204249 UP|Q9SB11 (Q9SB11) Glycinin, complete
Length = 1916
Score = 30.4 bits (67), Expect = 1.8
Identities = 41/157 (26%), Positives = 68/157 (43%), Gaps = 13/157 (8%)
Frame = -3
Query: 16 YVAVTVSILSFILLLIWS-----IFPFIVHKVPRTKGSGFWIPVIQVVASFNLLLSIMTI 70
+V V V +L F L L WS IF F++ V T + ++ ++ + I
Sbjct: 1065 FVLVLVLVL-FTLSLAWSTRTRFIFIFVLVLVLFTLSMAWSARRVRGNLFIFIIFVFIFI 889
Query: 71 SSNLKRAIGGSHVTCGQVSFHNHTLCLFPVWGEGPLGFGLL--------LSSRITQAFQL 122
L +G + + SFH H L F VW L + L+ + + T+
Sbjct: 888 FILLFLPLGADNA---ETSFHCHDLLPFVVWRLKFLSYVLVGVEGLCQEVFAEATEHAAT 718
Query: 123 YFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHIKKP 159
+F+F+ LPL+ S + L+LL + G V++I P
Sbjct: 717 FFLFLLLVLPLLASTMTFLLLLL--LHGLWVLYIWVP 613
>TC206195 similar to UP|Q8GZT8 (Q8GZT8) PTEN-like protein, partial (58%)
Length = 1688
Score = 30.0 bits (66), Expect = 2.4
Identities = 15/34 (44%), Positives = 20/34 (58%), Gaps = 1/34 (2%)
Frame = -3
Query: 398 ALNEIVHLMKTNLAKDYW-SSMFFLKFQEECDMR 430
+L +VH K LAKD W S F+L+ CD+R
Sbjct: 711 SLPSLVHQTKNLLAKDPWFSDAFWLRTTHHCDLR 610
>CA852155 similar to GP|15391731|em nitrite transporter {Cucumis sativus},
partial (22%)
Length = 516
Score = 29.6 bits (65), Expect = 3.1
Identities = 15/59 (25%), Positives = 27/59 (45%)
Frame = +1
Query: 166 MSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTAYI 224
+SV W +P CLH + +V V H+EF +D+ + R + + + Y+
Sbjct: 79 ISVFWLVPQYCLHGVAEVFMV-----VGHLEFLYDQSPESMRSTATALYCITTAIGNYV 240
>TC228825 weakly similar to UP|Q9V5S5 (Q9V5S5) CG33473-PB (Transcription
factor DKLF) (HL07808p), partial (3%)
Length = 1050
Score = 29.3 bits (64), Expect = 4.0
Identities = 25/88 (28%), Positives = 43/88 (48%), Gaps = 11/88 (12%)
Frame = +2
Query: 243 LLVLASILVLAFFSISSSQPLLS------QISLRRRESREFRTM-----GQALGIPDSGV 291
+L + + VL ++ QP+++ +IS +RE R+F+ + GQALGI +
Sbjct: 539 ILQPSGMFVLKRDQLADHQPVVTSGNESQEISAEQREVRDFQHIVTILSGQALGIVEDEW 718
Query: 292 LTQSEPISRVDPNEPLDKLLLNKKFRQS 319
+ + DP L LL+ KF +S
Sbjct: 719 DEEEFDVDDFDPEYAL*SLLV*SKFVKS 802
>TC213878 homologue to UP|Q8SCS3 (Q8SCS3) PHIKZ239, partial (9%)
Length = 619
Score = 29.3 bits (64), Expect = 4.0
Identities = 23/72 (31%), Positives = 29/72 (39%)
Frame = -3
Query: 7 AVKGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGFWIPVIQVVASFNLLLS 66
++K CPT Y S L + W I VH VPR WIP I++ +
Sbjct: 506 SIKARCPTKYKFPVTSNLEQ*VFT-WQITGHAVHGVPRRSLKL*WIPPIKLFEQYP-RYR 333
Query: 67 IMTISSNLKRAI 78
I I N R I
Sbjct: 332 IFLIHGNKSRTI 297
>TC206831 similar to UP|Q96400 (Q96400) Nitrite transporter, partial (68%)
Length = 1500
Score = 28.9 bits (63), Expect = 5.2
Identities = 21/76 (27%), Positives = 33/76 (42%)
Frame = +1
Query: 151 AAVIHIKKPLSNRCHMSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRFDELRDLWRGIL 210
AA H+ +SV W +P CLH + + +V H+EF F++ + R
Sbjct: 883 AAKYHLLDDPKATIPISVFWLVPQYCLHG-----VAEIFMSVGHLEFLFEQSPESMR--- 1038
Query: 211 VSSVSVAVWVTAYILN 226
SS + +T I N
Sbjct: 1039-SSATALYCITTAIGN 1083
>TC223303 similar to UP|Q8RV26 (Q8RV26) Oligopeptide transporter-like
protein, partial (20%)
Length = 432
Score = 28.9 bits (63), Expect = 5.2
Identities = 15/45 (33%), Positives = 24/45 (53%)
Frame = +1
Query: 215 SVAVWVTAYILNEIHDNISWLQVVSRFLLLVLASILVLAFFSISS 259
S ++VT +ILN + DN+ W V F + +A + L F I +
Sbjct: 289 SAGLFVTLFILNYVQDNVGW---VLGFGIPCIAMLTALVIFLIGT 414
>TC207129 homologue to UP|Q84XV9 (Q84XV9) Phosphoribosylformylglycinamidine
synthase, complete
Length = 4134
Score = 28.9 bits (63), Expect = 5.2
Identities = 11/36 (30%), Positives = 16/36 (43%)
Frame = +2
Query: 431 CNGYELEQMTGWNYSPRLSSVHGTDDPFHHDHLLKN 466
CNG +L + GW P++ VHG + N
Sbjct: 3416 CNGCQLMALLGWVPGPQVGGVHGAGGDLSQPRFIHN 3523
>AI442623
Length = 214
Score = 28.9 bits (63), Expect = 5.2
Identities = 19/60 (31%), Positives = 31/60 (51%), Gaps = 9/60 (15%)
Frame = -3
Query: 213 SVSVAVWVTAYILNEIHDNISWLQVVS---------RFLLLVLASILVLAFFSISSSQPL 263
++SV+VW+ Y ++ ++ WL VVS FLL A L+ F++ SS+ L
Sbjct: 182 TLSVSVWLGLYF*SKCKGDVPWLGVVSPLISIPAFLNFLLSFAAFCHFLSAFAVCSSKIL 3
>TC222287
Length = 524
Score = 28.5 bits (62), Expect = 6.8
Identities = 16/52 (30%), Positives = 29/52 (55%)
Frame = +3
Query: 203 RDLWRGILVSSVSVAVWVTAYILNEIHDNISWLQVVSRFLLLVLASILVLAF 254
R++W+ ++ V +TAYI H +S+ S+ +LL A+++VL F
Sbjct: 246 REIWKCATWMTIIVPTSMTAYIYLYSHGEVSY--AASQPVLLYAAAVMVLIF 395
>TC230924 UP|ATPH_LOTJA (P12231) ATP synthase C chain (Lipid-binding
protein) (Subunit III) , complete
Length = 567
Score = 28.5 bits (62), Expect = 6.8
Identities = 15/41 (36%), Positives = 25/41 (60%)
Frame = +2
Query: 116 ITQAFQLYFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHI 156
I+ L++I V R+L I+ FLL+ L+LL W G ++ +
Sbjct: 272 ISLTISLFYI*VTRKLS*IQLFLLLLLLLLGWP*GLLLLDL 394
>TC222233 similar to UP|Q6IMT1 (Q6IMT1) SAB, partial (7%)
Length = 597
Score = 28.5 bits (62), Expect = 6.8
Identities = 24/85 (28%), Positives = 35/85 (40%), Gaps = 2/85 (2%)
Frame = -3
Query: 312 LNKKFRQSFMGFADSCLAG-ESVHFFDEVHELSKISEHDCVRRIYMARHIIE-KYMVAGA 369
L+K R F F C A S+ FF H + + + R + KY+V
Sbjct: 445 LSKSSRGEFPSFGAPCRASTRSITFFP--HSDGGFHADKSISDLTLGRQFLTTKYLVVNR 272
Query: 370 AMEINISHRSKQEILSTSDLARADL 394
AM ++ RSK I+S + DL
Sbjct: 271 AMPTSL*SRSKSYIMSLISASAKDL 197
>TC225943 similar to UP|NU4C_LOTJA (Q9BBP3) NAD(P)H-quinone oxidoreductase
chain 4, chloroplast (NAD(P)H dehydrogenase, chain 4)
(NADH-plastoquinone oxidoreductase chain 4) , partial
(86%)
Length = 1432
Score = 28.5 bits (62), Expect = 6.8
Identities = 16/52 (30%), Positives = 24/52 (45%)
Frame = +3
Query: 409 NLAKDYWSSMFFLKFQEECDMRCNGYELEQMTGWNYSPRLSSVHGTDDPFHH 460
N+++ YW FFL E + L + GWN P ++HG FH+
Sbjct: 1029 NISRIYWRCAFFLGGNE---L**TTSSLSRRNGWNGYPNAKNIHG----FHY 1163
>TC209182 similar to PIR|T51854|T51854 RING-H2 finger protein RHF2a
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (16%)
Length = 649
Score = 28.5 bits (62), Expect = 6.8
Identities = 13/32 (40%), Positives = 21/32 (65%)
Frame = +3
Query: 225 LNEIHDNISWLQVVSRFLLLVLASILVLAFFS 256
L ++ D +SWLQ+ F L++ A +LVL +S
Sbjct: 291 LLQLLDLLSWLQIAKEFTLMIEALLLVLLQWS 386
>BU083551 homologue to GP|7301548|gb|A CG6066-PA {Drosophila melanogaster},
partial (3%)
Length = 409
Score = 28.1 bits (61), Expect = 8.9
Identities = 19/45 (42%), Positives = 24/45 (53%)
Frame = -3
Query: 245 VLASILVLAFFSISSSQPLLSQISLRRRESREFRTMGQALGIPDS 289
+LA I+ F S SSS PLL L R REFR + + + DS
Sbjct: 374 LLAGIIPPIFASSSSSSPLLLL**LFRTFDREFRFLNSSPSLHDS 240
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.140 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,147,904
Number of Sequences: 63676
Number of extensions: 432000
Number of successful extensions: 2921
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 2895
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2919
length of query: 475
length of database: 12,639,632
effective HSP length: 101
effective length of query: 374
effective length of database: 6,208,356
effective search space: 2321925144
effective search space used: 2321925144
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC123572.11


