

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122164.8 + phase: 0 /pseudo
(640 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
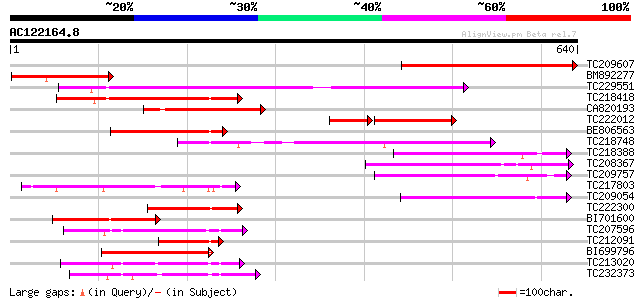
Score E
Sequences producing significant alignments: (bits) Value
TC209607 375 e-104
BM892277 179 3e-45
TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, p... 173 2e-43
TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partia... 155 4e-38
CA820193 similar to PIR|T04229|T04 ABC-type transport protein F1... 138 9e-33
TC222012 similar to UP|Q9FLX5 (Q9FLX5) ABC transporter-like prot... 111 3e-31
BE806563 similar to GP|11994752|db ABC transporter-like protein ... 107 2e-23
TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter ... 107 2e-23
TC218388 similar to UP|Q9LMU4 (Q9LMU4) F2H15.7 protein, partial ... 98 1e-20
TC208367 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {A... 96 5e-20
TC209757 93 4e-19
TC217803 similar to UP|Q9AT00 (Q9AT00) At1g65410/T8F5_19, partia... 91 2e-18
TC209054 similar to UP|Q6K376 (Q6K376) ABC transporter-like prot... 90 4e-18
TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {A... 89 8e-18
BI701600 88 1e-17
TC207596 similar to UP|Q7GBY1 (Q7GBY1) Tonoplast ABC transporter... 87 2e-17
TC212091 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like prot... 85 1e-16
BI699796 similar to PIR|G85167|G85 ABC transporter like protein ... 84 3e-16
TC213020 weakly similar to UP|Q8GU71 (Q8GU71) MDR-like ABC trans... 83 3e-16
TC232373 weakly similar to UP|AB10_MOUSE (Q9JI39) ATP-binding ca... 79 6e-15
>TC209607
Length = 828
Score = 375 bits (962), Expect = e-104
Identities = 179/198 (90%), Positives = 188/198 (94%)
Frame = +1
Query: 443 QEREILMKETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFL 502
QEREILMKETSCGSYRVSSYAIANG VYLPFLLILAILF++PLYWLVGLN NF AFLHFL
Sbjct: 1 QEREILMKETSCGSYRVSSYAIANGXVYLPFLLILAILFSMPLYWLVGLNRNFLAFLHFL 180
Query: 503 LLIWLVLYTANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHY 562
LLIWL+LYTANSVVVCFSALVPNFIVGNSVI GVIGSFFLFSGYFIS EIP+YWIFMHY
Sbjct: 181 LLIWLILYTANSVVVCFSALVPNFIVGNSVIAGVIGSFFLFSGYFISKQEIPNYWIFMHY 360
Query: 563 ISLFKYPFEGFLINEFSNSKKCLEYMFGACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFI 622
ISLFKYPFEGFLINEFSNS KCLEYMFGAC+ GEDVLKEEGYGGE +RWKNVGVTVCFI
Sbjct: 361 ISLFKYPFEGFLINEFSNSGKCLEYMFGACIKSGEDVLKEEGYGGESNRWKNVGVTVCFI 540
Query: 623 MVYRFISYVILRYKCSER 640
+VYRFISYVILRY+CS+R
Sbjct: 541 LVYRFISYVILRYRCSQR 594
>BM892277
Length = 421
Score = 179 bits (454), Expect = 3e-45
Identities = 89/120 (74%), Positives = 99/120 (82%), Gaps = 5/120 (4%)
Frame = +2
Query: 3 MQGCEVEVIGINYKIHTNKAEHPFKIFSKSPQLVNTNV-----QETEEVEKGCSGVRHVL 57
MQGCEV+ IGINY IHT+K+EHPFKIFS ++T +E EVE+ CSGVRHVL
Sbjct: 62 MQGCEVDAIGINYTIHTHKSEHPFKIFSNKSAHLDTEQDGKEPEEEAEVEQSCSGVRHVL 241
Query: 58 KNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPVDKSQFRKLSGYV 117
KNVSFQA+PWEILAIVGPSGAGKSSLLEILAGKH PQ G+V LN KPVDK+QFRKLSGYV
Sbjct: 242 KNVSFQAKPWEILAIVGPSGAGKSSLLEILAGKHSPQSGTVFLNHKPVDKAQFRKLSGYV 421
>TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (34%)
Length = 1436
Score = 173 bits (438), Expect = 2e-43
Identities = 123/472 (26%), Positives = 230/472 (48%), Gaps = 10/472 (2%)
Frame = +3
Query: 56 VLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK---HRPQKGSVLLNQKPVDKSQFRK 112
+L ++ A+P +LAI+GPSG+GKS+LL+ LAG+ + Q G +L+N + +
Sbjct: 75 ILHGLTGYAQPGRLLAIIGPSGSGKSTLLDALAGRLTSNIKQTGKILINGHKQELAY--G 248
Query: 113 LSGYVTQKDTLFPLLTVEETMMFSAKLRLK--LPQQQQCSRVKSLIKELGLDHVAGTRIG 170
SGYVTQ D + LT ET+ +SA L+ + +++ R ++E+GL TR+G
Sbjct: 249 TSGYVTQDDAMLSCLTAGETLYYSAMLQFPNTMSVEEKKERADMTLREMGLQDAINTRVG 428
Query: 171 DDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRG--R 228
+G+SGG+RRR+SI +E++ PK+L LDEPTSGLDS ++ ++ + + + G R
Sbjct: 429 GWNCKGLSGGQRRRLSICIEILTHPKLLFLDEPTSGLDSAASYYVMSGIANLIQRDGIQR 608
Query: 229 TIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAID 288
TI+ S+HQP + +LF+ L LL++G ++ G A + G P N + +
Sbjct: 609 TIVASVHQPSSEVFQLFHDLFLLSSGETVYFGPASDANQFFASNGFPCPPLYNPSDHYLR 788
Query: 289 SIDV-IQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDED 347
I+ Q + TE ++ + + + Q + + ++ K+
Sbjct: 789 IINKDFNQDADEGITTEEATKILVNSYKSSEFSNHVQSEIAKSETDFGACGKKKKI---- 956
Query: 348 IINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFC 407
+ + + +IL R S I+R R + + L +GSIF
Sbjct: 957 ---------------HAAFITQCLILIRRASLQIYRDTNNHRARLVVFIFISLSVGSIFY 1091
Query: 408 NL-KDDLRGTQERVGLFAFILTFL-LSSSIEALPIFLQEREILMKETSCGSYRVSSYAIA 465
+ DLR +R L F L+ + + + + ++E + +E G Y ++++ I+
Sbjct: 1092HSGGPDLRSIMDRGSLLCFFLSVVTFMTLVGGISPLIEEMNVFKRERLNGHYGITAFLIS 1271
Query: 466 NGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVV 517
N +P+ ++ I+ + +L GL+ F+ + +++ + S+++
Sbjct: 1272NIFSAVPYNFLMLIIPGAVVTYLSGLHKGVDNFVFLISVLFATVTWVESLMM 1427
>TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (49%)
Length = 1220
Score = 155 bits (393), Expect = 4e-38
Identities = 92/214 (42%), Positives = 135/214 (62%), Gaps = 5/214 (2%)
Frame = +1
Query: 54 RHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQ---KGSVLLNQKPVDKSQF 110
++VL+ ++ A P A++GPSG+GKS+LL+ L+ + G++LLN + S
Sbjct: 292 QNVLEGLTGYAEPGTFTALMGPSGSGKSTLLDALSSRLAANAFLSGTILLNGRKAKLSF- 468
Query: 111 RKLSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTR 168
+ YVTQ D L LTV ET+ +SA+LRL +P + + V+S I +GL A T
Sbjct: 469 -GTAAYVTQDDNLIGTLTVRETISYSARLRLPDNMPWADKRALVESTIVAMGLQDCADTV 645
Query: 169 IGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGR 228
IG+ +RGISGGE+RRVSI +E++ P++L LDEPTSGLDS SA + L+ +A GR
Sbjct: 646 IGNWHLRGISGGEKRRVSIALEILMRPRLLFLDEPTSGLDSASAFFVTQTLRALARD-GR 822
Query: 229 TIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTA 262
T+I SIHQP + +LF+ L LL++G ++ G A
Sbjct: 823 TVIASIHQPSSEVFELFDQLYLLSSGKTVYFGQA 924
>CA820193 similar to PIR|T04229|T04 ABC-type transport protein F14M19.30 -
Arabidopsis thaliana, partial (23%)
Length = 423
Score = 138 bits (347), Expect = 9e-33
Identities = 70/137 (51%), Positives = 97/137 (70%)
Frame = +1
Query: 152 VKSLIKELGLDHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTS 211
V SL+ EL L H++ TR+ RG+SGGERRRVSIG+ ++HDP VL+LDEPTSGLDSTS
Sbjct: 13 VSSLLSELRLTHLSNTRLA----RGLSGGERRRVSIGLCLLHDPAVLLLDEPTSGLDSTS 180
Query: 212 ALQIIDMLKVMAETRGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRL 271
A +++ +LK +R RTIILSIHQP F+I+ + +LLL+ G V+HHG+ L L
Sbjct: 181 AFKVMRILKQTCVSRNRTIILSIHQPSFKILACIDRILLLSKGQVVHHGSVATLQAFLHS 360
Query: 272 MGLELPLHVNVVEFAID 288
G +P +N +E+A++
Sbjct: 361 NGFTVPHQLNALEYAME 411
>TC222012 similar to UP|Q9FLX5 (Q9FLX5) ABC transporter-like protein, partial
(24%)
Length = 503
Score = 111 bits (278), Expect(2) = 3e-31
Identities = 51/93 (54%), Positives = 75/93 (79%)
Frame = +1
Query: 412 DLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYL 471
D G ++R GLFAF LTFLLSS+ E LPIF+ ER IL++ETS G YR+SSY IAN LV+L
Sbjct: 223 DKEGIEKRFGLFAFTLTFLLSSTTETLPIFINERPILLRETSSGVYRLSSYLIANTLVFL 402
Query: 472 PFLLILAILFTVPLYWLVGLNTNFTAFLHFLLL 504
P+L ++A+++++P+Y+LVGL ++ +F +F+L+
Sbjct: 403 PYLFVVAVIYSIPVYFLVGLCASWLSFAYFVLV 501
Score = 42.4 bits (98), Expect(2) = 3e-31
Identities = 20/48 (41%), Positives = 31/48 (63%)
Frame = +3
Query: 362 FANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNL 409
+ +SR+ E L RF K I+RT++L T + L+ GLVLG+I+ N+
Sbjct: 72 YKSSRVHEIFTLYSRFWKIIYRTRQLLLTNTAEALLVGLVLGTIYINI 215
>BE806563 similar to GP|11994752|db ABC transporter-like protein {Arabidopsis
thaliana}, partial (20%)
Length = 428
Score = 107 bits (267), Expect = 2e-23
Identities = 60/135 (44%), Positives = 89/135 (65%), Gaps = 4/135 (2%)
Frame = +1
Query: 115 GYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRIGDD 172
G+V Q D +P LTV ET+ ++A LRL L ++++ + +I ELGL + +G
Sbjct: 25 GFVPQDDVHYPHLTVLETLTYAALLRLPKSLSREEKKEHAEMVIAELGLTRCRNSPVGGC 204
Query: 173 RV--RGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTI 230
RGISGGER+RVSIG E++ +P +L +DEPTSGLDST+A I+ +L +A GRT+
Sbjct: 205 MALFRGISGGERKRVSIGQEMLVNPSLLFVDEPTSGLDSTTAQLIVSVLHGLARA-GRTV 381
Query: 231 ILSIHQPGFRIVKLF 245
+ +IHQP R+ ++F
Sbjct: 382 VATIHQPSSRLYRMF 426
>TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter 2
(Fragment), partial (37%)
Length = 1384
Score = 107 bits (267), Expect = 2e-23
Identities = 81/371 (21%), Positives = 166/371 (43%), Gaps = 12/371 (3%)
Frame = +3
Query: 190 EVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIHQPGFRIVKLFNSLL 249
E++ +P ++ +DEPTSGLD+ +A ++ ++ +T GRT++ +IHQP I + F+ LL
Sbjct: 9 ELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDT-GRTVVCTIHQPSIDIFESFDELL 185
Query: 250 LLANGSV------LHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSIDVIQQQQQWQVET 303
L+ G L H ++ L++ + G ++ I W +E
Sbjct: 186 LMKQGGQEIYVGPLGHHSSHLINYFEGIQG-------------VNKIKDGYNPATWMLEV 326
Query: 304 ETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDIINKTGTGMD--FSYD 361
T + + + G D + + +L++++K + +++ D F
Sbjct: 327 STSAK------------EMELGIDFAEVYKNSELYRRNKALIKELSTPAPGSKDLYFPSQ 470
Query: 362 FANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNLKDDLRGTQERVG 421
++ S L + M + + +R A R + VLGS+F +L + Q+
Sbjct: 471 YSTSFLTQCMACLWKQHWSYWRNPLYTAIRFLYSTAVAAVLGSMFWDLGSKIDKQQDLFN 650
Query: 422 ----LFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFLLIL 477
++A +L + ++ P+ ER + +E + G Y YA A L+ LP++L+
Sbjct: 651 AMGSMYAAVLLIGIKNANAVQPVVAVERTVFYREKAAGMYSALPYAFAQVLIELPYVLVQ 830
Query: 478 AILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGNSVINGVI 537
A+++ + +Y ++G T + +L ++ T + A+ PN + + V +
Sbjct: 831 AVVYGIIIYAMIGFEWTVTKVIWYLFFMYFTFLTFTYYGMMSVAVTPNQHISSIVSSAFY 1010
Query: 538 GSFFLFSGYFI 548
+ LFSG+ +
Sbjct: 1011AVWNLFSGFIV 1043
>TC218388 similar to UP|Q9LMU4 (Q9LMU4) F2H15.7 protein, partial (38%)
Length = 933
Score = 97.8 bits (242), Expect = 1e-20
Identities = 60/208 (28%), Positives = 109/208 (51%), Gaps = 7/208 (3%)
Frame = +1
Query: 434 SIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNT 493
SI P F+++ ++ +E G Y V+S+ I+N L +PFL+++ L Y++V L+
Sbjct: 13 SIGGFPSFVEDMKVFQRERLNGHYGVTSFVISNTLSAMPFLILITFLSGTICYFMVRLHP 192
Query: 494 NFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEI 553
F +L F+L ++ + S+++ +++VPNF++G + G+ G F L SGYF H+I
Sbjct: 193 GFWHYLFFVLCLYASVTVVESLMMAIASIVPNFLMGIIIGAGIQGIFMLVSGYFRLPHDI 372
Query: 554 PS-YWIF-MHYISLFKYPFEGFLINE-----FSNSKKCLEYMFGACVMKGEDVLKEEGYG 606
P W + M YIS + +G N+ F N L + G +++ K
Sbjct: 373 PKPVWRYPMSYISFHFWALQGQYQNDLRGLVFDNQTPDLPKIPGEYILE-----KVFQID 537
Query: 607 GEGSRWKNVGVTVCFIMVYRFISYVILR 634
S+W N+ V I++YR I +++++
Sbjct: 538 VNRSKWINLSVIFSMIVIYRIIFFIMIK 621
>TC208367 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Arabidopsis
thaliana;} , partial (34%)
Length = 1062
Score = 95.9 bits (237), Expect = 5e-20
Identities = 70/240 (29%), Positives = 118/240 (49%), Gaps = 5/240 (2%)
Frame = +1
Query: 402 LGSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSS 461
+G+IF + R R AFI F+ SI P F++E ++ KE G Y V
Sbjct: 1 VGTIFYGVGSSYRAIFARGACGAFISGFMTFMSIGGFPSFIEEMKVFYKERLNGYYGVGV 180
Query: 462 YAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSA 521
Y ++N L PF+ +++I Y++V T F+ +++ L + + S ++ ++
Sbjct: 181 YILSNFLSSFPFVAVMSIATGTITYYMVRFRTEFSHYVYICLDLIGCIAVVESSMMIIAS 360
Query: 522 LVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHY-ISLFKYPFEGFLINEFSN 580
LVPNF++G + G IG + +GYF ++P IF Y IS Y G L F N
Sbjct: 361 LVPNFLMGLIIGAGYIGVMMMTAGYFRQIPDLPK--IFWRYPISYINYGAWG-LQGAFKN 531
Query: 581 SKKCLEY---MFGACVMKGEDVLKEE-GYGGEGSRWKNVGVTVCFIMVYRFISYVILRYK 636
+E+ G +KGE +LK G E S+W ++ + +++ R + +VIL++K
Sbjct: 532 DMIGMEFDPLEPGGTKLKGEIILKTMLGIRVEISKWWDLAAVMIILVLLRVLFFVILKFK 711
>TC209757
Length = 1116
Score = 92.8 bits (229), Expect = 4e-19
Identities = 69/230 (30%), Positives = 112/230 (48%), Gaps = 7/230 (3%)
Frame = +3
Query: 412 DLRGTQERVGLFAFILTFL-LSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVY 470
D R Q+R+GL FI F + S ++ F QER I MKE + G Y +SSY +A +
Sbjct: 21 DYRNIQDRLGLLFFISIFWGVFPSFNSVFAFPQERAIFMKERASGMYTLSSYFMARIVGD 200
Query: 471 LPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGN 530
LP LIL +F + YW+ GL + AFL LL++ + + + + A + + +
Sbjct: 201 LPMELILPTIFLIVTYWMGGLKPDLWAFLLTLLVVLGYVMVSQGLGLALGAAIMDAKQAS 380
Query: 531 SVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSKK------C 584
+V + +F L GY++ H++PS ++ YIS Y + ++ + KK C
Sbjct: 381 TVAAVTMLAFVLTGGYYV--HKVPSCMAWIKYISTTFYCYRLLTRIQYEDGKKISYLLGC 554
Query: 585 LEYMFGACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILR 634
G C EDV+ + G G +GV + + YR ++Y+ LR
Sbjct: 555 YHGDKGGCRFVEEDVVGQIGTLG------CIGVLLFMFVFYRLLAYLALR 686
>TC217803 similar to UP|Q9AT00 (Q9AT00) At1g65410/T8F5_19, partial (81%)
Length = 1563
Score = 90.5 bits (223), Expect = 2e-18
Identities = 83/276 (30%), Positives = 130/276 (47%), Gaps = 29/276 (10%)
Frame = +3
Query: 14 NYKIHTNKAEHPFKIFSKSPQLVNTNVQETE-EVEKGCS------GVRHVLKNVSFQARP 66
N+K + A H F SKS QL E + +V C G + +L VSF+ +
Sbjct: 240 NFKSQDSSAIH-FNGSSKSEQLSTARDHEDDSDVLIECRDVYKSFGEKKILNGVSFKIKH 416
Query: 67 WEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKP----VDKSQFRKLS-GYVTQKD 121
E + I+GPSG GKS++L+I+AG P KG V + K V L G V Q
Sbjct: 417 GEAVGIIGPSGTGKSTVLKIIAGLLAPDKGEVYIRGKKRVGLVSDDDISGLRIGLVFQSA 596
Query: 122 TLFPLLTVEETMMFSAKLRLKLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVRG-ISGG 180
LF LTV E + F + + Q V + +GL V +DR+ +SGG
Sbjct: 597 ALFDSLTVRENVGFLLYEHSSMSEDQISELVTETLAAVGLKGV------EDRLPSELSGG 758
Query: 181 ERRRVSIGVEVIHD-------PKVLILDEPTSGLDSTSALQIIDMLKVM----AETRGR- 228
++RV++ +I D P+VL+ DEPT+GLD ++ + D+++ + + RG+
Sbjct: 759 MKKRVALARSIICDTTKESIEPEVLLYDEPTAGLDPIASTVVEDLIRSVHIKGQDARGKP 938
Query: 229 ----TIILSIHQPGFRIVKLFNSLLLLANGSVLHHG 260
+ ++ HQ I + + LL L G ++ G
Sbjct: 939 GNISSYVVVTHQHS-TIKRAIDRLLFLHKGKIVWEG 1043
>TC209054 similar to UP|Q6K376 (Q6K376) ABC transporter-like protein, partial
(27%)
Length = 913
Score = 89.7 bits (221), Expect = 4e-18
Identities = 47/196 (23%), Positives = 109/196 (54%), Gaps = 3/196 (1%)
Frame = +1
Query: 442 LQEREILMKETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHF 501
+++ ++ +E G Y V+++ I N +P+LL+++I+ Y+L GL +F F++F
Sbjct: 1 VEDMKVFERERLNGHYSVTAFVIGNTFSAIPYLLLVSIIPGAIAYYLPGLQKDFEHFVYF 180
Query: 502 LLLIWLVLYTANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIP-SYWIF- 559
+ +++ L S+++ +++VPNF++G G+ G + G+F +++P +W +
Sbjct: 181 ICVLFACLMLVESLMMIVASIVPNFLMGIITGAGIQGIMIIGGGFFRLPNDLPRPFWKYP 360
Query: 560 MHYISLFKYPFEGFLINEFSNSKKCLEYMFGACVMKGEDVLKEEGYGGEG-SRWKNVGVT 618
M Y++ +Y ++G NEF + + G + GE++L++ S+W ++G+
Sbjct: 361 MFYVAFHRYAYQGLFKNEFEGLRFATNNVGGGYI-SGEEILRDMWQVNMSYSKWFDLGIL 537
Query: 619 VCFIMVYRFISYVILR 634
+ I++YR + + ++
Sbjct: 538 LGMIILYRVLFLINIK 585
>TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Arabidopsis
thaliana;} , partial (37%)
Length = 801
Score = 88.6 bits (218), Expect = 8e-18
Identities = 47/107 (43%), Positives = 68/107 (62%)
Frame = +1
Query: 156 IKELGLDHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQI 215
I E+GL IG +RGISGGE++R+SI + ++ P++L LDEPTSGLDS SA +
Sbjct: 19 IIEMGLQDCDDRLIGKWHLRGISGGEKKRLSIAL*ILTRPRLLFLDEPTSGLDSASAFFV 198
Query: 216 IDMLKVMAETRGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTA 262
+ L+ +A RT+I SIHQP + L++ L LL+ G ++ G A
Sbjct: 199 VQTLRNVARDE-RTVIYSIHQPDSEVFALYDDLFLLSGGETVYFGEA 336
>BI701600
Length = 426
Score = 88.2 bits (217), Expect = 1e-17
Identities = 56/126 (44%), Positives = 79/126 (62%), Gaps = 4/126 (3%)
Frame = +3
Query: 49 GCSGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK-HRP-QKGSVLLNQKPVD 106
G R +LK V+ A+P EILA++GPSG+GKS+LL LAG+ H P G++L N +
Sbjct: 51 GAPKERTILKGVTGIAQPGEILAVLGPSGSGKSTLLHALAGRLHGPGLTGTILANSSKLT 230
Query: 107 KSQFRKLSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHV 164
K R+ +G+VTQ D L+P LTV ET++F A LRL L + ++ + ++ I ELGL
Sbjct: 231 KPVLRR-TGFVTQDDILYPHLTVRETLVFCAMLRLPRALLRSEKLAAAEAAIAELGLGKC 407
Query: 165 AGTRIG 170
T IG
Sbjct: 408 ENTIIG 425
>TC207596 similar to UP|Q7GBY1 (Q7GBY1) Tonoplast ABC transporter IDI7,
partial (35%)
Length = 1076
Score = 87.4 bits (215), Expect = 2e-17
Identities = 62/214 (28%), Positives = 121/214 (55%), Gaps = 6/214 (2%)
Frame = +2
Query: 61 SFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPV----DKSQFRKLSGY 116
+ + P +A+VGPSG GKS++ ++ + P KG +LLN P+ K RK+S
Sbjct: 2 TLKLHPGSKVALVGPSGGGKSTIANLIERFYDPTKGKILLNGVPLVEISHKHLHRKIS-I 178
Query: 117 VTQKDTLFPLLTVEETMMFSAKLRLKLPQQQQCSRVKSLIKELG-LDHVAGTRIGDDRVR 175
V+Q+ TLF ++EE + + ++ + +++ + + + T +G+ VR
Sbjct: 179 VSQEPTLFNC-SIEENIAYGFDGKVNDVDIENAAKMANAHEFISKFPEKYQTFVGERGVR 355
Query: 176 GISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIH 235
+SGG+++R++I ++ DPK+L+LDE TS LD+ S + D ++ + +GRT+++ H
Sbjct: 356 -LSGGQKQRIAIARALLMDPKILLLDEATSALDAESEYLVQDAMESL--MKGRTVLVIAH 526
Query: 236 QPGFRIVKLFNSLLLLANGSVLHHGT-ADLLSVN 268
+ VK +++ ++++G V+ G +LL+ N
Sbjct: 527 R--LSTVKTADTVAVISDGQVVERGNHEELLNKN 622
>TC212091 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like protein, partial
(12%)
Length = 477
Score = 84.7 bits (208), Expect = 1e-16
Identities = 42/73 (57%), Positives = 56/73 (76%)
Frame = +3
Query: 169 IGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGR 228
IGD+ R +SGGERRRVSIG ++IH+P VL LDEPTS LDSTSA ++ +L+ +A++ G
Sbjct: 12 IGDNGHRSVSGGERRRVSIGTDIIHNPIVLFLDEPTSNLDSTSAFMVVKVLQRIAQS-GN 188
Query: 229 TIILSIHQPGFRI 241
I+SIH P +RI
Sbjct: 189 IFIMSIH*PSYRI 227
>BI699796 similar to PIR|G85167|G85 ABC transporter like protein [imported] -
Arabidopsis thaliana, partial (13%)
Length = 421
Score = 83.6 bits (205), Expect = 3e-16
Identities = 45/129 (34%), Positives = 80/129 (61%), Gaps = 2/129 (1%)
Frame = +1
Query: 104 PVDKSQFRKLSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGL 161
P + F ++SGY Q D P +TVEE++MFSA LRL ++ + + V +I + L
Sbjct: 28 PKVQETFARVSGYCEQNDIHSPNITVEESVMFSAWLRLPSQIDAKTKAEFVNEVIHTIEL 207
Query: 162 DHVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKV 221
D + + +G + G+S +R+R++I VE++ +P ++ +DEPT+GLD+ +A ++ +K
Sbjct: 208 DGIKDSLVGMPNISGLSTEQRKRLTIAVELVANPSIIFMDEPTTGLDARAAAVVMRAVKN 387
Query: 222 MAETRGRTI 230
+ T GRT+
Sbjct: 388 VVGT-GRTV 411
>TC213020 weakly similar to UP|Q8GU71 (Q8GU71) MDR-like ABC transporter,
partial (18%)
Length = 864
Score = 83.2 bits (204), Expect = 3e-16
Identities = 62/213 (29%), Positives = 113/213 (52%), Gaps = 5/213 (2%)
Frame = +1
Query: 58 KNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPVDKSQFRKLS--- 114
+N++ + + LA+VG SG+GKS+++ ++ + P G VL+++ + R L
Sbjct: 1 ENLNLRVPAGKSLAVVGQSGSGKSTVISLVMRFYDPDSGLVLVDECDIKNLNLRSLRLRI 180
Query: 115 GYVTQKDTLFPLLTVEETMMFSAK--LRLKLPQQQQCSRVKSLIKELGLDHVAGTRIGDD 172
G V Q+ LF TV E + + + +++ + + + I + + T +G+
Sbjct: 181 GLVQQEPALFST-TVYENIKYGKEEASEIEVMKAAKAANAHEFISRMPEGYK--TEVGER 351
Query: 173 RVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIIL 232
V+ +SGG+++RV+I ++ DP +L+LDE TS LD+ S + + L + E GRT IL
Sbjct: 352 GVQ-LSGGQKQRVAIARAILKDPSILLLDEATSALDTVSERLVQEALDKLME--GRTTIL 522
Query: 233 SIHQPGFRIVKLFNSLLLLANGSVLHHGTADLL 265
H+ V+ NS+ +L NG V G+ + L
Sbjct: 523 VAHR--LSTVRDANSIAVLQNGRVAEMGSHERL 615
>TC232373 weakly similar to UP|AB10_MOUSE (Q9JI39) ATP-binding cassette,
sub-family B, member 10, mitochondrial precursor
(ATP-binding cassette transporter 10) (ABC transporter
10 protein) (ABC-mitochondrial erythroid protein)
(ABC-me protein), partial (24%)
Length = 742
Score = 79.0 bits (193), Expect = 6e-15
Identities = 64/225 (28%), Positives = 112/225 (49%), Gaps = 9/225 (4%)
Frame = +1
Query: 68 EILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPVDKSQ---FRKLSGYVTQKDTLF 124
E +A+ G SG+GKS+++ +L + P G + L+ + K Q FR+ G V+Q+ LF
Sbjct: 19 ETVALAGESGSGKSTVISLLQRFYEPDSGQITLDGTEIQKLQLKWFRQQMGLVSQEPVLF 198
Query: 125 PLLTVEETMMFSA---KLRLKLPQQQQCSRVKSLIKEL--GLDHVAGTRIGDDRVRGISG 179
T+ + + ++ + + + I L G D + G +R +SG
Sbjct: 199 ND-TIRTNIAYGKGGDATEAEIIAATELANAHTFISSLQQGYDTIVG-----ERGIQLSG 360
Query: 180 GERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDML-KVMAETRGRTIILSIHQPG 238
G+++RV+I ++ +PK+L+LDE TS LD S + D L +VM + RT I+ H+
Sbjct: 361 GQKQRVAIARAIVKNPKILLLDEATSALDVESERVVQDALDQVMVD---RTTIVVAHR-- 525
Query: 239 FRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVV 283
+K +S+ ++ NG + G D L + + LH N+V
Sbjct: 526 LSTIKDADSIAVVQNGVIAEQGKHDTLLNKGGIYASLVGLHTNLV 660
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.140 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,412,704
Number of Sequences: 63676
Number of extensions: 422149
Number of successful extensions: 2597
Number of sequences better than 10.0: 146
Number of HSP's better than 10.0 without gapping: 2527
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2557
length of query: 640
length of database: 12,639,632
effective HSP length: 103
effective length of query: 537
effective length of database: 6,081,004
effective search space: 3265499148
effective search space used: 3265499148
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC122164.8


