

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121244.7 - phase: 0
(418 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
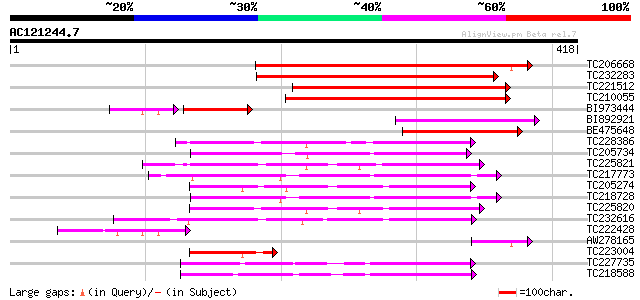
Score E
Sequences producing significant alignments: (bits) Value
TC206668 similar to UP|Q94FN1 (Q94FN1) Phosphatidylinositol tran... 273 1e-73
TC232283 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol tr... 239 2e-63
TC221512 similar to UP|Q94A34 (Q94A34) C7A10_870/C7A10_870 (At4g... 204 6e-53
TC210055 similar to UP|Q9LRX7 (Q9LRX7) Phosphatidylinositol/phos... 189 3e-48
BI973444 homologue to GP|14486705|gb phosphatidylinositol transf... 99 1e-27
BI892921 similar to GP|15215780|gb| C7A10_870/C7A10_870 {Arabido... 95 7e-20
BE475648 89 3e-18
TC228386 78 6e-15
TC205734 similar to UP|O48939 (O48939) Polyphosphoinositide bind... 72 4e-13
TC225821 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8,... 69 4e-12
TC217773 similar to UP|O48939 (O48939) Polyphosphoinositide bind... 69 5e-12
TC205274 similar to GB|AAN73306.1|25141223|BT002309 At3g51670/T1... 67 1e-11
TC218728 UP|O48939 (O48939) Polyphosphoinositide binding protein... 66 3e-11
TC225820 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8,... 65 6e-11
TC232616 similar to GB|AAD25756.1|4587525|T5I8 Contains the PF|0... 64 1e-10
TC222428 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol tr... 63 3e-10
AW278165 similar to GP|14486705|gb phosphatidylinositol transfer... 63 3e-10
TC223004 similar to GB|AAN73306.1|25141223|BT002309 At3g51670/T1... 62 5e-10
TC227735 similar to UP|O48940 (O48940) Polyphosphoinositide bind... 62 6e-10
TC218588 UP|O48940 (O48940) Polyphosphoinositide binding protein... 60 2e-09
>TC206668 similar to UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like
protein III, partial (72%)
Length = 1782
Score = 273 bits (697), Expect = 1e-73
Identities = 140/237 (59%), Positives = 164/237 (69%), Gaps = 33/237 (13%)
Frame = +1
Query: 182 YNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIA 241
Y PHG+HGVDKEGRPV+IER K+D NKLMQVTT+DRYVKYH Q E+ IKFPACTIA
Sbjct: 10 YYPHGHHGVDKEGRPVYIERLGKVDPNKLMQVTTMDRYVKYHVQEFEKAFKIKFPACTIA 189
Query: 242 SKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSL 301
+KRHIDSS TILD+QG+G N ++ R+++ R KI DNYP+T Q FIIN G R L
Sbjct: 190 AKRHIDSSTTILDVQGVGLKNFTKSARDLIMRLQKIDGDNYPETLCQMFIINAGPGFRLL 369
Query: 302 RSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPW 361
+ + F+DPK SK+HV+G++YQ KLL++IDASELP FLGGTCTCA+QGGCLRSDKGPW
Sbjct: 370 WNTVKSFLDPKTTSKIHVLGNKYQSKLLEIIDASELPEFLGGTCTCADQGGCLRSDKGPW 549
Query: 362 NNPEILK---------------------------------VKGSDTLTAESSSEAED 385
NPEILK VKGSDT TAES SEAED
Sbjct: 550 KNPEILKMILSGEARRARPVVKVLNSEGKVIAYARPQYPMVKGSDTSTAESGSEAED 720
>TC232283 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like
protein III, partial (28%)
Length = 535
Score = 239 bits (609), Expect = 2e-63
Identities = 114/178 (64%), Positives = 139/178 (78%)
Frame = +1
Query: 183 NPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIAS 242
+PHG+HG+DKEGRPV+IER K+D NKLMQVTT+DRYVKYH Q E+ AIKFPAC+IA+
Sbjct: 1 DPHGHHGIDKEGRPVYIERLGKVDPNKLMQVTTLDRYVKYHVQEFEKAFAIKFPACSIAA 180
Query: 243 KRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLR 302
KRHIDSS TILD+ G+G N ++ RE++ R KI DNYP+T Q FIIN G R L
Sbjct: 181 KRHIDSSTTILDVHGVGLKNFTKSARELITRLQKIDGDNYPETLCQMFIINAGPGFRLLW 360
Query: 303 SICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGP 360
+ + F+DPK SK+HV+G++YQ KLL+VIDASELP FLGGTCTC + GGCL SDKGP
Sbjct: 361 NTVKSFLDPKTTSKIHVLGNKYQSKLLEVIDASELPEFLGGTCTCEDHGGCLTSDKGP 534
>TC221512 similar to UP|Q94A34 (Q94A34) C7A10_870/C7A10_870
(At4g36490/C7A10_870), partial (43%)
Length = 1158
Score = 204 bits (519), Expect = 6e-53
Identities = 93/161 (57%), Positives = 127/161 (78%)
Frame = +1
Query: 209 KLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADR 268
KLMQVTT++RY+KYH + E A+K PAC+IA+K+HID S TILD+QG+G +L +A R
Sbjct: 7 KLMQVTTMERYLKYHVKEFERTFAVKLPACSIAAKKHIDQSTTILDVQGVGLKSLNKAAR 186
Query: 269 EIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKL 328
++++R KI DNYP++ + FIIN G R L + + F+DPK SK+HV+G++YQ KL
Sbjct: 187 DLLQRLQKIDGDNYPESLNRMFIINAGSGFRLLWNTIKSFLDPKTTSKIHVLGNKYQSKL 366
Query: 329 LKVIDASELPTFLGGTCTCANQGGCLRSDKGPWNNPEILKV 369
L++IDASELP FLGGTCTCA++GGC+ SDKGPWN+P+ILK+
Sbjct: 367 LEIIDASELPEFLGGTCTCADKGGCMLSDKGPWNDPDILKM 489
>TC210055 similar to UP|Q9LRX7 (Q9LRX7)
Phosphatidylinositol/phosphatidylcholine transfer
protein-like, partial (27%)
Length = 615
Score = 189 bits (479), Expect = 3e-48
Identities = 82/166 (49%), Positives = 125/166 (74%)
Frame = +3
Query: 204 KLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNL 263
+++ +KLM VTT+DR++KYH Q E+M KFPAC+IA+KRHID + TILD+ G+ + +
Sbjct: 3 QVEPSKLMNVTTVDRFLKYHVQGFEKMFKEKFPACSIAAKRHIDKTTTILDVHGVNWVSF 182
Query: 264 EEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDR 323
+ +++ R KI DNYP+T Q FI+N G + L + + F+DP+ +K+HV+G++
Sbjct: 183 SKVAHDLVMRMQKIDGDNYPETLNQMFIVNAGSGFKLLWNTAKGFLDPRTTAKIHVLGNK 362
Query: 324 YQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPWNNPEILKV 369
+Q +LL++ID+S+LP FLGG+C+C N GGCLRS+KGPWN+P+ILK+
Sbjct: 363 FQSRLLEIIDSSQLPDFLGGSCSCPNDGGCLRSNKGPWNDPDILKL 500
>BI973444 homologue to GP|14486705|gb phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (18%)
Length = 355
Score = 99.4 bits (246), Expect(2) = 1e-27
Identities = 44/51 (86%), Positives = 48/51 (93%)
Frame = +2
Query: 129 DDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEV 179
DDYHMMLRFLKARKFDI KAKHMW DML+WRKEFGADTI++DFEF EL+EV
Sbjct: 203 DDYHMMLRFLKARKFDIEKAKHMWTDMLQWRKEFGADTIVQDFEFKELDEV 355
Score = 42.4 bits (98), Expect(2) = 1e-27
Identities = 29/63 (46%), Positives = 34/63 (53%), Gaps = 12/63 (19%)
Frame = +3
Query: 74 RRSDFKKKLLNAFSKFKHSFKKK--------RVDELQAVNVVP----AVVDAFRKLLIMD 121
R KKK LNA SKFKH+ +KK RV + +V VDAFR+ LIMD
Sbjct: 3 RIGSLKKKALNASSKFKHTLRKKSSRRKSDGRVSSVSIEDVRDFEELQAVDAFRQSLIMD 182
Query: 122 ELL 124
ELL
Sbjct: 183 ELL 191
>BI892921 similar to GP|15215780|gb| C7A10_870/C7A10_870 {Arabidopsis
thaliana}, partial (14%)
Length = 352
Score = 94.7 bits (234), Expect = 7e-20
Identities = 46/106 (43%), Positives = 64/106 (59%)
Frame = +3
Query: 285 TGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGT 344
T Q FIIN G R L + F+D + +K+HV+G Y LL+ ID+S LPTFL
Sbjct: 3 TLNQLFIINAGSGFRMLWKAVKTFLDVRTVAKIHVLGFNYLSVLLEAIDSSNLPTFLCVN 182
Query: 345 CTCANQGGCLRSDKGPWNNPEILKVKGSDTLTAESSSEAEDNVIAV 390
CTC++ GGCL SD+GPW NPE+L++ L E ++ED + +
Sbjct: 183 CTCSDYGGCLMSDRGPWKNPEVLEMIQVVNLREEIDGKSEDGDVDI 320
>BE475648
Length = 422
Score = 89.4 bits (220), Expect = 3e-18
Identities = 44/90 (48%), Positives = 59/90 (64%), Gaps = 1/90 (1%)
Frame = +1
Query: 290 FIINVGLEL-RSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCA 348
+IIN G R L + F+D K +K+ V+ + KLL +ID+S+LP FLGGTCTC
Sbjct: 19 YIINAGPGFKRMLWPAAQKFLDAKTIAKIQVLEPKSLCKLLDIIDSSQLPDFLGGTCTCP 198
Query: 349 NQGGCLRSDKGPWNNPEILKVKGSDTLTAE 378
+GGCLRS KGPWN+P+I+K+ S T E
Sbjct: 199 GEGGCLRSSKGPWNDPDIMKMVHSVEATFE 288
>TC228386
Length = 1517
Score = 78.2 bits (191), Expect = 6e-15
Identities = 63/228 (27%), Positives = 111/228 (48%), Gaps = 7/228 (3%)
Frame = +1
Query: 123 LLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKY 182
LL + D ++L+FL+AR+F + +A M + ++WRKEFG + +ME+ +EL +V+
Sbjct: 163 LLADERSDV-ILLKFLRAREFRVKEAFTMLKNTIQWRKEFGMEELMEEKLGDELEKVV-- 333
Query: 183 NPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTID-----RYVKYHAQRCEE-MHAIKFP 236
HG DKEG PV +E+ +L + T D +++++ Q E+ + + F
Sbjct: 334 ---FMHGFDKEGHPVCYNIYEEFQNKELYKKTFSDEEKREKFLRWRIQFLEKSIRKLDFN 504
Query: 237 ACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGL 296
I + H++ + + G+ L R+ K L++L DNYP+ + INV
Sbjct: 505 PGGICTIVHVND---LKNSPGLAKWEL----RQATKHALQLLQDNYPEFVAKQVFINVPW 663
Query: 297 ELRSLRSICEYFMDPKVASKVHVIG-DRYQRKLLKVIDASELPTFLGG 343
++ + F+ + SK G + LL+ I +LP GG
Sbjct: 664 WYLAVNRMISPFLTQRTKSKFVFAGPSKSTETLLRYIAPEQLPVKYGG 807
>TC205734 similar to UP|O48939 (O48939) Polyphosphoinositide binding protein
Ssh1p, partial (71%)
Length = 1491
Score = 72.4 bits (176), Expect = 4e-13
Identities = 57/213 (26%), Positives = 96/213 (44%), Gaps = 6/213 (2%)
Frame = +2
Query: 134 MLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNP-HGYHGVDK 192
++RFLKAR ++I KA M D L WR E D ++ +L I+ + G G K
Sbjct: 404 LIRFLKARDWNIAKAHKMLIDCLNWRVENEIDNVLRKPIPMDLYRAIRDSQLIGMSGYSK 583
Query: 193 EGRPVFIERFEKLDRNKLMQVTTIDR-----YVKYHAQRCEEMHAIKFPACTIASKRHID 247
EG PV + ++T D+ Y++ H Q E + P T R+I
Sbjct: 584 EGLPVIAVG---------VGLSTYDKASDKYYIQSHIQLNEYRDQVILPTATRKHGRYIG 736
Query: 248 SSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEY 307
+ + +LD+ G+ F L + ++ I NYP+ +I+NV + + +
Sbjct: 737 TCVKVLDMTGLKFSALNQL--RLLTAISTIDDLNYPEKTDTYYIVNVPYVFSACWKVVKP 910
Query: 308 FMDPKVASKVHVIGDRYQRKLLKVIDASELPTF 340
+ + K+ V+ + +LLKV+D + LP F
Sbjct: 911 LLQERTRRKIQVLQGCGKEELLKVMDYASLPHF 1009
>TC225821 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial
(84%)
Length = 1480
Score = 68.9 bits (167), Expect = 4e-12
Identities = 67/262 (25%), Positives = 121/262 (45%), Gaps = 10/262 (3%)
Frame = +2
Query: 99 DELQAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEW 158
+E + VVP V+ + L+ DE D ++L+FL+AR F + +A +M + + W
Sbjct: 59 EEEKGREVVPEEVEIWGIPLLGDE-----RSDV-ILLKFLRARDFKVKEALNMIRNTVRW 220
Query: 159 RKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTID- 217
RKEFG + ++E+ ++ +V+ + G DKEG PV+ F + + +L T +D
Sbjct: 221 RKEFGIEGLVEEDLGSDWEKVVFKD-----GYDKEGHPVYYNVFGEFEDKELYSKTFLDE 385
Query: 218 ----RYVKYHAQRCEE-MHAIKFPACTIASKRHIDSSITILDLQ---GIGFCNLEEADRE 269
+++++ Q E+ + ++ F S I + + + DL+ G+G L +A +
Sbjct: 386 EKRNKFIRWRIQSLEKSVRSLDF------SPNGISTIVQVNDLKNSPGLGKRELRQATNQ 547
Query: 270 IMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIG-DRYQRKL 328
L++L DNYP+ + INV + + F + SK G + L
Sbjct: 548 A----LQLLQDNYPEFVAKQIFINVPWWYLAFSRMISPFFTQRTKSKFVFAGPSKSADTL 715
Query: 329 LKVIDASELPTFLGGTCTCANQ 350
+ I +P GG A Q
Sbjct: 716 FRYIAPELVPVQYGGLSREAEQ 781
>TC217773 similar to UP|O48939 (O48939) Polyphosphoinositide binding protein
Ssh1p, partial (92%)
Length = 1218
Score = 68.6 bits (166), Expect = 5e-12
Identities = 68/273 (24%), Positives = 121/273 (43%), Gaps = 13/273 (4%)
Frame = +1
Query: 103 AVNVVPAVVDAFRKLLIMDELLPQKHDDYHM------MLRFLKARKFDIGKAKHMWADML 156
A+N + A++D ++L+ +E L + + H + RFLKAR+++ KA M D L
Sbjct: 16 ALNQLQALMD---QVLLEEEPLQRTFQNVHQGCVTETLTRFLKAREWNATKAHKMIVDCL 186
Query: 157 EWRKEFGADTIM-EDFEFNELNEVIKYNP-HGYHGVDKEGRPVF-----IERFEKLDRNK 209
+WR + D I+ + +L I+ + G G +EG PVF + F+K
Sbjct: 187 KWRVQNEIDNILSKPIIPTDLYRGIRDSQLIGLSGYSREGLPVFAIGVGLSTFDK----- 351
Query: 210 LMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADRE 269
++ YV+ H Q E + P+ + +R I + + ILD+ G+ L + +
Sbjct: 352 ----ASVHYYVQSHIQINEYRDRVILPSASKKHERPITTCVKILDMTGLKLSALNQI--K 513
Query: 270 IMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLL 329
++ I NYP+ +I+N + + + + + KV V+ + +LL
Sbjct: 514 LLTIISSIDDLNYPEKTNTYYIVNAPYIFSACWKVVKPLLQERTRRKVQVLQGCGRDELL 693
Query: 330 KVIDASELPTFLGGTCTCANQGGCLRSDKGPWN 362
K++D + LP F C G S G N
Sbjct: 694 KIMDYASLPHF----CRREGSGSSRHSGNGNEN 780
>TC205274 similar to GB|AAN73306.1|25141223|BT002309 At3g51670/T18N14_50
{Arabidopsis thaliana;} , partial (79%)
Length = 1470
Score = 67.4 bits (163), Expect = 1e-11
Identities = 59/220 (26%), Positives = 95/220 (42%), Gaps = 9/220 (4%)
Frame = +3
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMED--FEFNELNEVIKYNPHGYHGV 190
++L+FL+AR F + A M L WR EFGAD I+++ F EL V+ Y HG
Sbjct: 228 ILLKFLRARDFRVHDALSMLLKCLSWRTEFGADNIVDEELGGFKELEGVVAYT----HGY 395
Query: 191 DKEGRPVFIERF-----EKLDRNKLMQVTTIDRYVKYHAQRCEE-MHAIKFPACTIASKR 244
D+EG PV + ++ N + +++++ Q E + + F
Sbjct: 396 DREGHPVCYNAYGVFKDREMYENVFGDEEKLKKFLRWRVQVLERGVRMLHF------KPG 557
Query: 245 HIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSI 304
++S I + DL+ + L A +I+ F DNYP+ + INV L S+
Sbjct: 558 GVNSLIQVTDLKDMPKRELRIASNQILSLFQ----DNYPEMVARKIFINVPWYFSVLYSM 725
Query: 305 CEYFMDPKVASKVHVIGD-RYQRKLLKVIDASELPTFLGG 343
F+ + SK + + L K I +P GG
Sbjct: 726 FSPFLTQRTKSKFVISKEGNAAETLYKFIRPENIPVRYGG 845
>TC218728 UP|O48939 (O48939) Polyphosphoinositide binding protein Ssh1p,
complete
Length = 1416
Score = 66.2 bits (160), Expect = 3e-11
Identities = 60/236 (25%), Positives = 101/236 (42%), Gaps = 7/236 (2%)
Frame = +3
Query: 134 MLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIM-EDFEFNELNEVIKYNP-HGYHGVD 191
++RFLKAR +D KA M D L WR + D I+ + +L ++ + G G
Sbjct: 255 LMRFLKARDWDPCKAHKMLVDCLNWRVQNEIDNILSKPIVPADLYRAVRDSQLIGLSGYS 434
Query: 192 KEGRPVF-----IERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHI 246
+EG PVF + F+K ++ YV+ H Q E I P+ + R I
Sbjct: 435 REGLPVFAIGVGLSTFDK---------ASVHYYVQSHIQINEYRERIILPSASKKQGRPI 587
Query: 247 DSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICE 306
+ I +LD+ G+ L + +++ I NYP+ +I+N + + +
Sbjct: 588 TTCIKVLDMTGLKLSALNQI--KLLTIISSIDDLNYPEKTNTYYIVNAPYIFSACWKVVK 761
Query: 307 YFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPWN 362
+ + K+ V+ + +LL ++D S LP F C G S+ G N
Sbjct: 762 PLLQERTRRKIQVLPGCGRDELLTIMDYSSLPHF----CRREGSGSSRHSESGSEN 917
>TC225820 weakly similar to UP|Q94AG3 (Q94AG3) At1g72160/T9N14_8, partial
(84%)
Length = 1376
Score = 65.1 bits (157), Expect = 6e-11
Identities = 58/228 (25%), Positives = 104/228 (45%), Gaps = 10/228 (4%)
Frame = +2
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDK 192
++L+FL+AR F + A M + + WRKEFG + ++E+ ++ ++V+ HG DK
Sbjct: 98 ILLKFLRARDFKVKDALSMLRNTVRWRKEFGIEGLVEEDLGSDWDKVV-----FSHGHDK 262
Query: 193 EGRPVFIERFEKLDRNKLMQVTTID-----RYVKYHAQRCEE-MHAIKFPACTIASKRHI 246
EG PV+ F + + +L T D + +++ Q E+ + ++ F S I
Sbjct: 263 EGHPVYYNVFGEFEDKELYNKTFWDEEKRNKLIRWMIQSLEKSVRSLDF------SPTGI 424
Query: 247 DSSITILDLQ---GIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRS 303
+ + + DL+ G+G L +A ++++ F DNYP+ + INV +
Sbjct: 425 STIVQVNDLKNSPGLGKRELRQATNQVLQLFQ----DNYPEFVAKQIFINVPWWYLAFSR 592
Query: 304 ICEYFMDPKVASKVHVIG-DRYQRKLLKVIDASELPTFLGGTCTCANQ 350
+ F + SK G + L + I +P GG A Q
Sbjct: 593 MISPFFTQRTKSKFLFAGPSKSAHTLFQYIAPELVPVQYGGLSREAEQ 736
>TC232616 similar to GB|AAD25756.1|4587525|T5I8 Contains the PF|00650
CRAL/TRIO phosphatidyl-inositol-transfer protein domain.
ESTs gb|T76582, gb|N06574 and gb|Z25700 come from this
gene. {Arabidopsis thaliana;} , partial (48%)
Length = 1214
Score = 63.9 bits (154), Expect = 1e-10
Identities = 72/274 (26%), Positives = 120/274 (43%), Gaps = 6/274 (2%)
Frame = +2
Query: 77 DFKKKLLNAFSKFKHSFKKKRVDELQAVNVVPAVVDAFRKLLIMDELLPQKHDD--YHMM 134
D KK+ L + K+ ++K +E + V V V + L LP K + ++
Sbjct: 359 DPKKEALPENEEKKNEGEEKEEEEEKRVEVEENDVSIWGVTL-----LPSKGAEGVEVVL 523
Query: 135 LRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEG 194
L+FL+AR+F + A M L+WRKE D+++++ ++L N GVD EG
Sbjct: 524 LKFLRAREFKVNDAFEMLKKTLKWRKESKIDSVVDEDFGSDLASAAYMN-----GVDHEG 688
Query: 195 RPVFIERFEKLDRNKLMQVT--TIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITI 252
PV F + + Q T T ++ ++ RC+ M K + S + I
Sbjct: 689 HPVCYNIFGAFESEESYQKTFGTEEKRSEFLRWRCQLME--KGIQRLNLKPGGVSSLLQI 862
Query: 253 LDLQGI-GFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDP 311
DL+ G L A ++ + F DNYP+ ++ INV +L ++ F+
Sbjct: 863 NDLKNSPGPSKLRVATKQTLAMFQ----DNYPEMVAKNIFINVPFWYYALNALLSPFLTQ 1030
Query: 312 KVASKVHVI-GDRYQRKLLKVIDASELPTFLGGT 344
+ SK V ++ L K I E+P GG+
Sbjct: 1031RTKSKFVVARPNKGTETLTKYIPIEEIPVHYGGS 1132
>TC222428 homologue to UP|Q94FN1 (Q94FN1) Phosphatidylinositol transfer-like
protein III, partial (18%)
Length = 627
Score = 62.8 bits (151), Expect = 3e-10
Identities = 48/124 (38%), Positives = 64/124 (50%), Gaps = 26/124 (20%)
Frame = +2
Query: 36 LFIHSSEMLRMIFVFGLVIDSSIVMFWFLTGSDENRKERRSDF--------------KKK 81
L++ S+++ + G + + F +GSDE +KERRSDF KKK
Sbjct: 257 LYVFVSQLIIATTMSGPLDRFARPCFEGFSGSDE-KKERRSDFEYSEDERRTRIGSLKKK 433
Query: 82 LLNAFSKFKHSFKKK--------RVDELQAVNVVP----AVVDAFRKLLIMDELLPQKHD 129
LNA SKFKH+ +KK RV + +V VDAFR+ LIMDELLP+
Sbjct: 434 ALNASSKFKHTLRKKSSRRKSDGRVSSVSIEDVRDFEELQAVDAFRQSLIMDELLPEAFA 613
Query: 130 DYHM 133
DYHM
Sbjct: 614 DYHM 625
>AW278165 similar to GP|14486705|gb phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (22%)
Length = 427
Score = 62.8 bits (151), Expect = 3e-10
Identities = 35/78 (44%), Positives = 39/78 (49%), Gaps = 33/78 (42%)
Frame = +2
Query: 341 LGGTCTCANQGGCLRSDKGPWNNPEILK-------------------------------- 368
LGGTCTC +QGGCLRSDKGPW NP+I K
Sbjct: 2 LGGTCTCEDQGGCLRSDKGPWKNPDIFKMVLTGGAWRSKQVVKVLNNERKVIVYAKPVYP 181
Query: 369 -VKGSDTLTAESSSEAED 385
VK +DT TAES SEA+D
Sbjct: 182 MVKCNDTSTAESRSEAKD 235
>TC223004 similar to GB|AAN73306.1|25141223|BT002309 At3g51670/T18N14_50
{Arabidopsis thaliana;} , partial (44%)
Length = 862
Score = 62.0 bits (149), Expect = 5e-10
Identities = 33/67 (49%), Positives = 41/67 (60%), Gaps = 2/67 (2%)
Frame = +3
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMED--FEFNELNEVIKYNPHGYHGV 190
++L+FL+AR F +G A HM L WRKEFGADTI+E+ EL V+ Y G
Sbjct: 354 ILLKFLRARDFRVGDAHHMLMKCLSWRKEFGADTILEEEFLGLKELEGVVAY----MQGY 521
Query: 191 DKEGRPV 197
DKEG PV
Sbjct: 522 DKEGHPV 542
>TC227735 similar to UP|O48940 (O48940) Polyphosphoinositide binding protein
Ssh2p, partial (92%)
Length = 1123
Score = 61.6 bits (148), Expect = 6e-10
Identities = 56/219 (25%), Positives = 98/219 (44%), Gaps = 2/219 (0%)
Frame = +1
Query: 127 KHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHG 186
K ++ MM RFL+AR D+ KA M+ L+W++ F + + +E+ E I +
Sbjct: 316 KEENDLMMRRFLRARSLDVEKASAMFLKYLKWKRSFVPNGYISP---SEIAEDIAQDKVF 486
Query: 187 YHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQR-CEEMHAIKFPACTIASKRH 245
G+DK+GRP+ + K ++K RYV + ++ C M +
Sbjct: 487 TQGLDKKGRPIVVTFAAKHFQSK-NGADGFKRYVVFVLEKLCSRMPPGQ----------- 630
Query: 246 IDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSIC 305
+ + I D++G + N +D L IL D YP+ G+ I++ + +
Sbjct: 631 -EKFLAIADIKGWAYVN---SDLRGYLNSLSILQDCYPERLGKMLIVHAPYMFMKIWKMI 798
Query: 306 EYFMDPKVASK-VHVIGDRYQRKLLKVIDASELPTFLGG 343
F+D K V V + + LL+ I+ S++P GG
Sbjct: 799 YPFIDENTKKKIVFVENKKLKSTLLEEIEESQIPDIYGG 915
>TC218588 UP|O48940 (O48940) Polyphosphoinositide binding protein Ssh2p,
complete
Length = 1030
Score = 60.1 bits (144), Expect = 2e-09
Identities = 56/219 (25%), Positives = 98/219 (44%), Gaps = 1/219 (0%)
Frame = +3
Query: 127 KHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHG 186
K +D M+ RFL+AR D+ KA M L+WR F + +++ + +
Sbjct: 234 KEEDDFMIRRFLRARDLDVEKASAMLLKYLKWRNSFVPN---GSVSVSDVPNELAQDKVF 404
Query: 187 YHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHI 246
G DK GRP+ + + +NK +D + ++ +++ A P
Sbjct: 405 MQGHDKIGRPILMVFGGRHFQNK----DGLDEFKRFVVYVLDKVCASMPPG--------Q 548
Query: 247 DSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICE 306
+ + I +L+G G+ N +D L IL D YP+ G+ FI+N + I
Sbjct: 549 EKFVGIAELKGWGYSN---SDVRGYLSALSILQDYYPERLGKLFIVNAPYIFMKVWQIVY 719
Query: 307 YFMDPKVASK-VHVIGDRYQRKLLKVIDASELPTFLGGT 344
F+D K K V V ++ + LL+ ++ S++P GG+
Sbjct: 720 PFIDNKTKKKIVFVEKNKVKSTLLEEMEESQVPEIFGGS 836
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.140 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,504,285
Number of Sequences: 63676
Number of extensions: 253270
Number of successful extensions: 1693
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 1645
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1663
length of query: 418
length of database: 12,639,632
effective HSP length: 100
effective length of query: 318
effective length of database: 6,272,032
effective search space: 1994506176
effective search space used: 1994506176
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC121244.7


