
BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33181.1 120 aa
(120 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
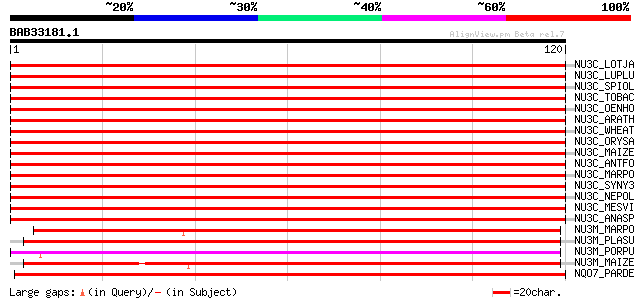
Score E
Sequences producing significant alignments: (bits) Value
NU3C_LOTJA (Q9BBT8) NAD(P)H-quinone oxidoreductase chain 3, chlo... 244 2e-65
NU3C_LUPLU (P52765) NAD(P)H-quinone oxidoreductase chain 3, chlo... 229 1e-60
NU3C_SPIOL (Q9M3L9) NAD(P)H-quinone oxidoreductase chain 3, chlo... 226 5e-60
NU3C_TOBAC (P06258) NAD(P)H-quinone oxidoreductase chain 3, chlo... 226 9e-60
NU3C_OENHO (Q9MTP5) NAD(P)H-quinone oxidoreductase chain 3, chlo... 220 5e-58
NU3C_ARATH (P56751) NAD(P)H-quinone oxidoreductase chain 3, chlo... 219 9e-58
NU3C_WHEAT (P26303) NAD(P)H-quinone oxidoreductase chain 3, chlo... 219 1e-57
NU3C_ORYSA (P12126) NAD(P)H-quinone oxidoreductase chain 3, chlo... 216 6e-57
NU3C_MAIZE (P19044) NAD(P)H-quinone oxidoreductase chain 3, chlo... 216 1e-56
NU3C_ANTFO (Q31792) NAD(P)H-quinone oxidoreductase chain 3, chlo... 191 3e-49
NU3C_MARPO (P06259) NAD(P)H-quinone oxidoreductase chain 3, chlo... 189 1e-48
NU3C_SYNY3 (P19045) NAD(P)H-quinone oxidoreductase chain 3 (EC 1... 166 9e-42
NU3C_NEPOL (Q9TKX9) NAD(P)H-quinone oxidoreductase chain 3, chlo... 163 6e-41
NU3C_MESVI (Q9MUQ9) NAD(P)H-quinone oxidoreductase chain 3, chlo... 161 3e-40
NU3C_ANASP (Q44239) NAD(P)H-quinone oxidoreductase chain 3 (EC 1... 156 7e-39
NU3M_MARPO (P26847) NADH-ubiquinone oxidoreductase chain 3 (EC 1... 104 3e-23
NU3M_PLASU (Q36518) NADH-ubiquinone oxidoreductase chain 3 (EC 1... 103 7e-23
NU3M_PORPU (O99978) NADH-ubiquinone oxidoreductase chain 3 (EC 1... 103 9e-23
NU3M_MAIZE (P16265) NADH-ubiquinone oxidoreductase chain 3 (EC 1... 102 2e-22
NQO7_PARDE (P29919) NADH-quinone oxidoreductase chain 7 (EC 1.6.... 102 2e-22
>NU3C_LOTJA (Q9BBT8) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 244 bits (623), Expect = 2e-65
Identities = 120/120 (100%), Positives = 120/120 (100%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI
Sbjct: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
>NU3C_LUPLU (P52765) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 229 bits (583), Expect = 1e-60
Identities = 112/120 (93%), Positives = 115/120 (95%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFWAFLIIS IPILAF ISG LAPI+KGPEKLSSYESGIEPMGDAWLQF+I
Sbjct: 1 MFLLYEYDIFWAFLIISSLIPILAFLISGILAPISKGPEKLSSYESGIEPMGDAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLG+SVFIEALIFVLILIVG VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGVSVFIEALIFVLILIVGLVYAWRKGALEWS 120
>NU3C_SPIOL (Q9M3L9) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 226 bits (577), Expect = 5e-60
Identities = 110/120 (91%), Positives = 114/120 (94%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFWAFLIIS IPILAF SG LAPI+KGPEKLSSYESGIEPMGDAWLQF+I
Sbjct: 1 MFLLYEYDIFWAFLIISSVIPILAFLFSGILAPISKGPEKLSSYESGIEPMGDAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFD+LG+SVFIEALIFVLILIVG VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDILGVSVFIEALIFVLILIVGLVYAWRKGALEWS 120
>NU3C_TOBAC (P06258) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 226 bits (575), Expect = 9e-60
Identities = 108/120 (90%), Positives = 114/120 (95%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYD FWAFLIISI +PILAF ISG LAPI+KGPEKLS+YESGIEPMGDAWLQF+I
Sbjct: 1 MFLLYEYDFFWAFLIISILVPILAFLISGVLAPISKGPEKLSTYESGIEPMGDAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLG+SVFIEA IFVLILI+G VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGVSVFIEAFIFVLILIIGLVYAWRKGALEWS 120
>NU3C_OENHO (Q9MTP5) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 220 bits (560), Expect = 5e-58
Identities = 105/120 (87%), Positives = 113/120 (93%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFWAFLIIS IPILAF ISG LAP + GPEKLSSYESGIEPMGDAWLQF+I
Sbjct: 1 MFLLYEYDIFWAFLIISSVIPILAFRISGLLAPTSIGPEKLSSYESGIEPMGDAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVET+FLYPWA+SFD+LG+SVFIEALIFVLIL++G VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETIFLYPWALSFDILGVSVFIEALIFVLILVLGLVYAWRKGALEWS 120
>NU3C_ARATH (P56751) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 219 bits (558), Expect = 9e-58
Identities = 105/120 (87%), Positives = 112/120 (92%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFWAFL+IS IP+LAF ISG L+PI KGPEKLSSYESGIEP+GDAWLQF+I
Sbjct: 1 MFLLYEYDIFWAFLLISSAIPVLAFLISGVLSPIRKGPEKLSSYESGIEPIGDAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLG+S FIEA IFVLILI+G VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGVSAFIEAFIFVLILILGLVYAWRKGALEWS 120
>NU3C_WHEAT (P26303) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 219 bits (557), Expect = 1e-57
Identities = 104/120 (86%), Positives = 114/120 (94%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLL+EYDIFW FLII+ IPILAF+ISG LAP+++GPEKLSSYESGIEPMG AW+QF+I
Sbjct: 1 MFLLHEYDIFWTFLIIASLIPILAFSISGLLAPVSEGPEKLSSYESGIEPMGGAWVQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLG+SVFIEALIFVLIL+VG VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGVSVFIEALIFVLILVVGLVYAWRKGALEWS 120
>NU3C_ORYSA (P12126) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 216 bits (551), Expect = 6e-57
Identities = 105/120 (87%), Positives = 111/120 (92%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLL+EYDIFWAFLII+ IPILAF IS LAP+ +GPEKLSSYESGIEPMG AWLQF+I
Sbjct: 1 MFLLHEYDIFWAFLIIASLIPILAFWISALLAPVREGPEKLSSYESGIEPMGGAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEA IFVLIL+VG VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEAFIFVLILVVGLVYAWRKGALEWS 120
>NU3C_MAIZE (P19044) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 216 bits (549), Expect = 1e-56
Identities = 103/120 (85%), Positives = 111/120 (91%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLL+EYDIFW FLII+ IPIL F ISG LAP+++GPEKLSSYESGIEPMG AWLQF+I
Sbjct: 1 MFLLHEYDIFWTFLIIASLIPILVFWISGLLAPVSEGPEKLSSYESGIEPMGGAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLG+SVFIEA IFVLIL+VG VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGVSVFIEAFIFVLILVVGLVYAWRKGALEWS 120
>NU3C_ANTFO (Q31792) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 191 bits (485), Expect = 3e-49
Identities = 88/120 (73%), Positives = 102/120 (84%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFL+ +Y+ FW FL+I+ IP +AF+IS +API+KGPEK +SYE GIEPMGDAW+QFQI
Sbjct: 1 MFLVSKYNYFWIFLLIASLIPTIAFSISRVIAPISKGPEKFTSYECGIEPMGDAWIQFQI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFV+FDVETVFLYPWAMSF LGI FIE IFV ILI+G +YAWRKGALEWS
Sbjct: 61 RYYMFALVFVIFDVETVFLYPWAMSFKQLGIPAFIEVFIFVFILIIGLIYAWRKGALEWS 120
>NU3C_MARPO (P06259) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 189 bits (479), Expect = 1e-48
Identities = 91/120 (75%), Positives = 104/120 (85%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLL +YD F+ FL+I F IL F++S ++APINKGPEK +SYESGIEPMG+A +QFQI
Sbjct: 1 MFLLQKYDYFFVFLLIISFFSILIFSLSKWIAPINKGPEKFTSYESGIEPMGEACIQFQI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFV+FDVETVFLYPWAMSF GIS FIEALIF+LILI+G VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVIFDVETVFLYPWAMSFYNFGISSFIEALIFILILIIGLVYAWRKGALEWS 120
>NU3C_SYNY3 (P19045) NAD(P)H-quinone oxidoreductase chain 3 (EC
1.6.5.-) (NAD(P)H dehydrogenase I, chain 3) (NDH-1,
chain 3)
Length = 120
Score = 166 bits (420), Expect = 9e-42
Identities = 77/120 (64%), Positives = 95/120 (79%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MF+L Y+ F FL I +P+LA T S L P + GPE+ ++YESG+EP+G AW+QF I
Sbjct: 1 MFVLTGYEYFLGFLFICSLVPVLALTASKLLRPRDGGPERQTTYESGMEPIGGAWIQFNI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWA++F+ LG+ F+EALIF+ IL+V VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAVAFNQLGLLAFVEALIFIAILVVALVYAWRKGALEWS 120
>NU3C_NEPOL (Q9TKX9) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 163 bits (413), Expect = 6e-41
Identities = 74/120 (61%), Positives = 94/120 (77%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
M +L Y AF +I+ IP++A + S L P GPE+ ++YESGIEPMG+AW+QF I
Sbjct: 1 MMILEGYGSVLAFFVIASLIPVIALSASKLLRPRGGGPERRTTYESGIEPMGEAWIQFNI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVET+FLYPWA++F LG+S F E LIF+++L++G VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETLFLYPWAVTFQRLGLSAFFEVLIFIIVLLIGLVYAWRKGALEWS 120
>NU3C_MESVI (Q9MUQ9) NAD(P)H-quinone oxidoreductase chain 3,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
3) (NADH-plastoquinone oxidoreductase chain 3)
Length = 120
Score = 161 bits (407), Expect = 3e-40
Identities = 76/120 (63%), Positives = 93/120 (77%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MF+L YD F F I++ +PILA + S + P G EK +YESGIEPMG+AW+QF I
Sbjct: 1 MFILEGYDSFVVFFIVACLVPILALSGSKLIRPKLSGIEKKMTYESGIEPMGEAWVQFNI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFAL+FV+FDVET+FLYPWA+ F LGI+ F+E LIF+ ILI+G VYAWRKGALEWS
Sbjct: 61 RYYMFALIFVIFDVETLFLYPWAIVFKDLGITAFLETLIFLSILIIGLVYAWRKGALEWS 120
>NU3C_ANASP (Q44239) NAD(P)H-quinone oxidoreductase chain 3 (EC
1.6.5.-) (NAD(P)H dehydrogenase I, chain 3) (NDH-1,
chain 3) (NDH-C)
Length = 120
Score = 156 bits (395), Expect = 7e-39
Identities = 75/120 (62%), Positives = 90/120 (74%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MF+L Y+ FLII +P LA + S L P E+ ++YESG+EP+G AW+QF I
Sbjct: 1 MFVLSGYEYLLGFLIICSLVPALALSASKLLRPTGNSLERRTTYESGMEPIGGAWIQFNI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWA++F LG+ FIEALIF+ IL+V VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAVAFHRLGLLAFIEALIFIAILVVALVYAWRKGALEWS 120
>NU3M_MARPO (P26847) NADH-ubiquinone oxidoreductase chain 3 (EC
1.6.5.3)
Length = 118
Score = 104 bits (260), Expect = 3e-23
Identities = 45/116 (38%), Positives = 77/116 (65%), Gaps = 2/116 (1%)
Query: 6 EYDIFWAFLIISIFIPILAFTISGFLAPINK--GPEKLSSYESGIEPMGDAWLQFQIRYY 63
E+ + +L+IS+ + ++ +S A + PEKLS+YE G +P DA +F IR+Y
Sbjct: 2 EFAPIFVYLVISLLLSLILIGVSFLFASSSSLAYPEKLSAYECGFDPFDDARSRFDIRFY 61
Query: 64 MFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEW 119
+ +++F++FD+E FL+PWA+S + +G+ F ++F+ IL +G VY W+KGAL+W
Sbjct: 62 LVSILFIIFDLEVTFLFPWAVSLNKIGLFGFWSMMVFLFILTIGFVYEWKKGALDW 117
>NU3M_PLASU (Q36518) NADH-ubiquinone oxidoreductase chain 3 (EC
1.6.5.3)
Length = 117
Score = 103 bits (257), Expect = 7e-23
Identities = 45/116 (38%), Positives = 72/116 (61%)
Query: 4 LYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQIRYY 63
+ EY + + S+ + L +S AP PEK+S+YE G +P DA +F IR+Y
Sbjct: 1 MIEYLAVLIYFLFSLALASLIIFLSFIFAPQKPDPEKISAYECGFDPFDDARGKFDIRFY 60
Query: 64 MFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEW 119
+ A++F++FD+E FL+PWA++ +G F + F++IL +G +Y W+KGALEW
Sbjct: 61 LVAILFIIFDLEVTFLFPWAVTLGKIGFFGFWTMMAFLIILTIGFIYEWKKGALEW 116
>NU3M_PORPU (O99978) NADH-ubiquinone oxidoreductase chain 3 (EC
1.6.5.3)
Length = 121
Score = 103 bits (256), Expect = 9e-23
Identities = 50/120 (41%), Positives = 72/120 (59%), Gaps = 1/120 (0%)
Query: 1 MFLLY-EYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQ 59
M +LY EY F IS I L +S FL P EK+S+YE G P DA F
Sbjct: 1 MNVLYNEYSAILTFFAISFSISTLILALSYFLNPQPSDQEKVSAYECGFNPFDDARATFD 60
Query: 60 IRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEW 119
+R+Y+ A++F++FD+E FL+PW++ L I F ++F++IL +G +Y W+KGALEW
Sbjct: 61 VRFYLVAILFLIFDLEISFLFPWSLVLGQLSIFGFWSMIVFLIILTLGFIYEWKKGALEW 120
>NU3M_MAIZE (P16265) NADH-ubiquinone oxidoreductase chain 3 (EC
1.6.5.3)
Length = 118
Score = 102 bits (253), Expect = 2e-22
Identities = 45/118 (38%), Positives = 77/118 (65%), Gaps = 3/118 (2%)
Query: 4 LYEYDIFWAFLIISIFIPILAFTISGFLAPINKG--PEKLSSYESGIEPMGDAWLQFQIR 61
+ E+ +L+IS+ + ++ + FL N PEKLS+YE G +P GDA +F IR
Sbjct: 1 MLEFAPICIYLVISLLVSLILLGVP-FLFASNSSTYPEKLSAYECGFDPFGDARSRFDIR 59
Query: 62 YYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEW 119
+Y+ +++F++FD+E F +PWA+S + + + F + F+LIL +GS+Y W++GAL+W
Sbjct: 60 FYLVSILFIIFDLEVTFFFPWAVSLNKIDLFGFWSMMAFLLILFIGSLYEWKRGALDW 117
>NQO7_PARDE (P29919) NADH-quinone oxidoreductase chain 7 (EC
1.6.99.5) (NADH dehydrogenase I, chain 7) (NDH-1, chain
7)
Length = 121
Score = 102 bits (253), Expect = 2e-22
Identities = 46/119 (38%), Positives = 74/119 (61%)
Query: 2 FLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQIR 61
+LL EY FL ++ + I+ + +A N PEK+S+YE G DA ++F +R
Sbjct: 3 YLLQEYLPILVFLGMASALAIVLILAAAVIAVRNPDPEKVSAYECGFNAFDDARMKFDVR 62
Query: 62 YYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
+Y+ +++F++FD+E FL+PWA+SF L F ++F+ +L VG Y W+KGALEW+
Sbjct: 63 FYLVSILFIIFDLEVAFLFPWAVSFASLSDVAFWGLMVFLAVLTVGFAYEWKKGALEWA 121
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.333 0.149 0.480
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,801,213
Number of Sequences: 164201
Number of extensions: 514406
Number of successful extensions: 2045
Number of sequences better than 10.0: 164
Number of HSP's better than 10.0 without gapping: 144
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 1814
Number of HSP's gapped (non-prelim): 166
length of query: 120
length of database: 59,974,054
effective HSP length: 96
effective length of query: 24
effective length of database: 44,210,758
effective search space: 1061058192
effective search space used: 1061058192
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 58 (26.9 bits)
Description of BAB33181.1

