
BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33181.1 120 aa
(120 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
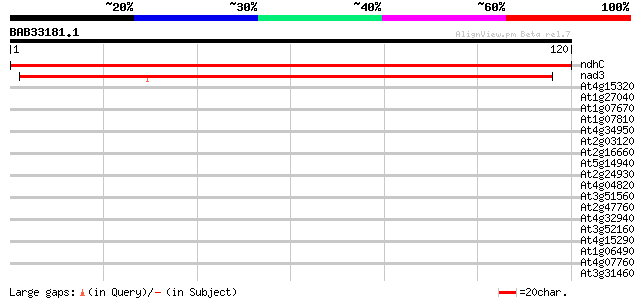
Score E
Sequences producing significant alignments: (bits) Value
ndhC -chloroplast genome- NADH dehydrogenase D3 219 2e-58
nad3 -mitochondrial genome- NADH dehydrogenase subunit 3 82 3e-17
At4g15320 cellulose synthase like protein 29 0.33
At1g27040 nitrate transporter like protein 29 0.43
At1g07670 endoplasmic reticulum-type calcium-transporting ATPase... 28 0.56
At1g07810 ER-type Ca2+-pump protein 28 0.56
At4g34950 unknown protein 27 1.2
At2g03120 unknown protein 27 1.2
At2g16660 nodulin-like protein 27 1.6
At5g14940 oligopeptide transporter -like protein 27 2.1
At2g24930 hypothetical protein 26 2.8
At4g04820 hypothetical protein 26 3.6
At3g51560 disease resistance-like protein 26 3.6
At2g47760 Not56-like protein 26 3.6
At4g32940 gamma-VPE (vacuolar processing enzyme) 25 4.7
At3g52160 beta-ketoacyl-CoA synthase like protein 25 4.7
At4g15290 cellulose synthase like protein 25 6.2
At1g06490 glucan synthase, putative 25 6.2
At4g07760 putative transposon protein 25 8.0
At3g31460 hypothetical protein 25 8.0
>ndhC -chloroplast genome- NADH dehydrogenase D3
Length = 120
Score = 219 bits (558), Expect = 2e-58
Identities = 105/120 (87%), Positives = 112/120 (92%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFWAFL+IS IP+LAF ISG L+PI KGPEKLSSYESGIEP+GDAWLQF+I
Sbjct: 1 MFLLYEYDIFWAFLLISSAIPVLAFLISGVLSPIRKGPEKLSSYESGIEPIGDAWLQFRI 60
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLG+S FIEA IFVLILI+G VYAWRKGALEWS
Sbjct: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGVSAFIEAFIFVLILILGLVYAWRKGALEWS 120
>nad3 -mitochondrial genome- NADH dehydrogenase subunit 3
Length = 119
Score = 82.4 bits (202), Expect = 3e-17
Identities = 37/115 (32%), Positives = 71/115 (61%), Gaps = 1/115 (0%)
Query: 3 LLYEYDIFWAFLIISIFIPILAFTIS-GFLAPINKGPEKLSSYESGIEPMGDAWLQFQIR 61
++ E+ +L+IS+ + ++ + F + + PEKLS+YE G +P GDA +F IR
Sbjct: 1 MMSEFAPISIYLVISLLVSLILLGVPFPFASNSSTYPEKLSAYECGFDPSGDARSRFDIR 60
Query: 62 YYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGA 116
+Y+ +++F++ D+E F +PWA+ + + + F + F+ IL +G +Y W++GA
Sbjct: 61 FYLVSILFLIPDLEVTFFFPWAVPPNKIDLFGFWSMMAFLFILTIGFLYEWKRGA 115
>At4g15320 cellulose synthase like protein
Length = 828
Score = 29.3 bits (64), Expect = 0.33
Identities = 16/50 (32%), Positives = 22/50 (44%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEP 50
M L + W F I I + +L + GF+ PE S YES +P
Sbjct: 676 MILGFSVKSCWLFSIQDIILKLLGISKIGFIVAKKNMPETRSGYESKSKP 725
>At1g27040 nitrate transporter like protein
Length = 563
Score = 28.9 bits (63), Expect = 0.43
Identities = 30/97 (30%), Positives = 43/97 (43%), Gaps = 12/97 (12%)
Query: 12 AFLIISIFIPILAFT-ISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQIRYYMF----- 65
AFL + +++ L I G L + G E+ ++ G P G YY+F
Sbjct: 143 AFLFVGLYLVSLGIGGIKGSLP--SHGAEQ---FDEGT-PKGRKQRSTFFNYYVFCLSCG 196
Query: 66 ALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVL 102
ALV V F V W F V IS+F+ L+F+L
Sbjct: 197 ALVAVTFVVWIEDNKGWEWGFGVSTISIFLSILVFLL 233
>At1g07670 endoplasmic reticulum-type calcium-transporting ATPase 4
(ECA4)
Length = 1061
Score = 28.5 bits (62), Expect = 0.56
Identities = 18/72 (25%), Positives = 33/72 (45%), Gaps = 9/72 (12%)
Query: 38 PEKLSSYESGIEPMGDAWLQFQIRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEA 97
PE S ++ +E D ++ + + + V FD + +GI+ F+E
Sbjct: 66 PEGTSIFKLILEQFNDTLVRILLAAAVISFVLAFFDGD---------EGGEMGITAFVEP 116
Query: 98 LIFVLILIVGSV 109
L+ LILIV ++
Sbjct: 117 LVIFLILIVNAI 128
>At1g07810 ER-type Ca2+-pump protein
Length = 1061
Score = 28.5 bits (62), Expect = 0.56
Identities = 18/72 (25%), Positives = 33/72 (45%), Gaps = 9/72 (12%)
Query: 38 PEKLSSYESGIEPMGDAWLQFQIRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEA 97
PE S ++ +E D ++ + + + V FD + +GI+ F+E
Sbjct: 66 PEGTSIFKLILEQFNDTLVRILLAAAVISFVLAFFDGD---------EGGEMGITAFVEP 116
Query: 98 LIFVLILIVGSV 109
L+ LILIV ++
Sbjct: 117 LVIFLILIVNAI 128
>At4g34950 unknown protein
Length = 567
Score = 27.3 bits (59), Expect = 1.2
Identities = 11/38 (28%), Positives = 22/38 (56%)
Query: 69 FVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIV 106
F VF++ V + + S+D++GI ++ F IL++
Sbjct: 215 FAVFNIVAVVVAVYLQSYDIIGIKTGAFSIAFASILLI 252
>At2g03120 unknown protein
Length = 344
Score = 27.3 bits (59), Expect = 1.2
Identities = 13/31 (41%), Positives = 16/31 (50%), Gaps = 5/31 (16%)
Query: 4 LYEYDIFWAFLIISIFIPILAFTISGFLAPI 34
L+ YDIFW F F P++ F API
Sbjct: 194 LFFYDIFWVF-----FTPVMVSVAKSFDAPI 219
>At2g16660 nodulin-like protein
Length = 546
Score = 26.9 bits (58), Expect = 1.6
Identities = 10/37 (27%), Positives = 22/37 (59%)
Query: 69 FVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILI 105
F +F++ V + + S+D++GI + ++ F IL+
Sbjct: 213 FTIFNIVAVVVAVYLQSYDIIGIKTGVFSVAFASILL 249
>At5g14940 oligopeptide transporter -like protein
Length = 547
Score = 26.6 bits (57), Expect = 2.1
Identities = 24/113 (21%), Positives = 44/113 (38%), Gaps = 12/113 (10%)
Query: 16 ISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQIRYYMFALVFVVFDVE 75
I +F+ I+A I+ + K+ ++P+ WL Q + +F V ++
Sbjct: 394 IGMFLSIIAIVIAALVERKRLKISKMMKTTPNLDPVSILWLLPQYILLGISDIFTVVGMQ 453
Query: 76 TVF---------LYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEW 119
F +A+ V G+ F+ A LI I+ + + R G W
Sbjct: 454 EFFYSEVPVSMRTMGFALYTSVFGVGSFVSA---ALISIIETYTSSRGGKHNW 503
>At2g24930 hypothetical protein
Length = 926
Score = 26.2 bits (56), Expect = 2.8
Identities = 23/87 (26%), Positives = 37/87 (42%), Gaps = 3/87 (3%)
Query: 33 PINKGPEKLSSYESGIEPMGDAWLQFQIRYYMFALVFVVFDVETVFLYPWA-MSFDVLGI 91
P +GP L G+ AW + R+ F VFD E + YPW +++ L +
Sbjct: 136 PTGEGPN-LDELRQGLVTCR-AWPTEKRRWIPFESAKRVFDDEAMKTYPWGRAAYEALVV 193
Query: 92 SVFIEALIFVLILIVGSVYAWRKGALE 118
S+ + I + G V+ + A E
Sbjct: 194 SIKLLRPIGKTYTVSGLVFVLQAWAYE 220
>At4g04820 hypothetical protein
Length = 253
Score = 25.8 bits (55), Expect = 3.6
Identities = 18/66 (27%), Positives = 30/66 (45%), Gaps = 1/66 (1%)
Query: 54 AWLQFQIRYYMFALVFVVFDVETVFLYPWA-MSFDVLGISVFIEALIFVLILIVGSVYAW 112
AW + R+ F VFD E + YPW +++ L +S+ + I + G V+
Sbjct: 155 AWPPEKRRWIPFESAKRVFDDEAMKTYPWGRAAYEALVVSIKLLRPIGKTYTVSGLVFVL 214
Query: 113 RKGALE 118
+ A E
Sbjct: 215 QAWAYE 220
>At3g51560 disease resistance-like protein
Length = 1253
Score = 25.8 bits (55), Expect = 3.6
Identities = 14/35 (40%), Positives = 20/35 (57%), Gaps = 1/35 (2%)
Query: 55 WLQFQIRYYMF-ALVFVVFDVETVFLYPWAMSFDV 88
W I+Y++ V D+E +FL P A+SFDV
Sbjct: 475 WKPLIIKYFLEDRQVLGSEDIEAIFLDPSALSFDV 509
>At2g47760 Not56-like protein
Length = 438
Score = 25.8 bits (55), Expect = 3.6
Identities = 12/43 (27%), Positives = 23/43 (52%)
Query: 63 YMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILI 105
Y + + FD+ VF++ W+++F + VF+ V +LI
Sbjct: 240 YPVSYIANAFDLGRVFIHFWSVNFKFVPERVFVSKEFAVCLLI 282
>At4g32940 gamma-VPE (vacuolar processing enzyme)
Length = 494
Score = 25.4 bits (54), Expect = 4.7
Identities = 13/39 (33%), Positives = 22/39 (56%), Gaps = 2/39 (5%)
Query: 27 ISGFLAPINKGPEKLSSYESGIEPMGDAW--LQFQIRYY 63
+ L I++GPE L+ S +P+ D W L+ Q+R +
Sbjct: 400 VGKILFGISRGPEVLNKVRSAGQPLVDDWNCLKNQVRAF 438
>At3g52160 beta-ketoacyl-CoA synthase like protein
Length = 451
Score = 25.4 bits (54), Expect = 4.7
Identities = 10/18 (55%), Positives = 14/18 (77%)
Query: 13 FLIISIFIPILAFTISGF 30
F+ +S F+PILAF +S F
Sbjct: 57 FIFVSRFLPILAFPLSTF 74
>At4g15290 cellulose synthase like protein
Length = 710
Score = 25.0 bits (53), Expect = 6.2
Identities = 15/46 (32%), Positives = 20/46 (42%)
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYES 46
M L + W F I I + +L + GF+ PE S YES
Sbjct: 555 MSLGFSVQSCWLFSIQDIILKLLGISQIGFVIAKKTIPETKSVYES 600
>At1g06490 glucan synthase, putative
Length = 1933
Score = 25.0 bits (53), Expect = 6.2
Identities = 8/17 (47%), Positives = 12/17 (70%)
Query: 1 MFLLYEYDIFWAFLIIS 17
MF L++Y FW L++S
Sbjct: 674 MFALFKYTFFWVMLLLS 690
>At4g07760 putative transposon protein
Length = 783
Score = 24.6 bits (52), Expect = 8.0
Identities = 15/31 (48%), Positives = 18/31 (57%), Gaps = 2/31 (6%)
Query: 14 LIISIFIPILAFTISGFLAPINKGPEKLSSY 44
LII + P + FT G PI KG KL+SY
Sbjct: 180 LIIWVINPQVEFTNMG--EPIGKGSVKLTSY 208
>At3g31460 hypothetical protein
Length = 172
Score = 24.6 bits (52), Expect = 8.0
Identities = 13/32 (40%), Positives = 19/32 (58%), Gaps = 1/32 (3%)
Query: 63 YMFALVFVVFDVETVFLYPWAMS-FDVLGISV 93
Y ++ VFD E + +YPWA S F+ L S+
Sbjct: 79 YHLRVLKEVFDDEAMKIYPWARSAFEALVDSI 110
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.333 0.149 0.480
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,571,517
Number of Sequences: 26719
Number of extensions: 95590
Number of successful extensions: 316
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 301
Number of HSP's gapped (non-prelim): 23
length of query: 120
length of database: 11,318,596
effective HSP length: 96
effective length of query: 24
effective length of database: 8,753,572
effective search space: 210085728
effective search space used: 210085728
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 52 (24.6 bits)
Description of BAB33181.1

