
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33181.1 120 aa
(120 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
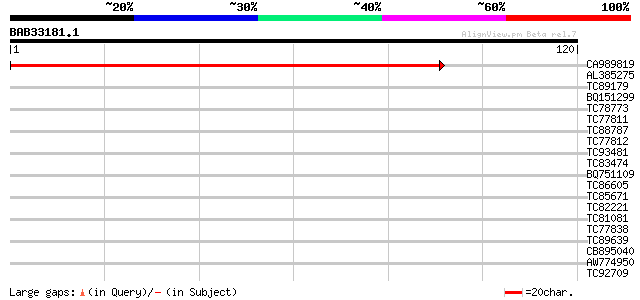
CA989819 homologue to SP|Q9BBT8|NU3C_ NADH-plastoquinone oxidore... 181 6e-47
AL385275 30 0.28
TC89179 similar to GP|10505183|gb|AAG18376.1 lipoxygenase {Zante... 29 0.37
BQ151299 28 0.82
TC78773 similar to GP|17473842|gb|AAL38345.1 unknown protein {Ar... 28 1.1
TC77811 similar to GP|17473842|gb|AAL38345.1 unknown protein {Ar... 27 1.4
TC88787 weakly similar to GP|21593424|gb|AAM65391.1 fatty acid e... 27 1.4
TC77812 similar to GP|17473842|gb|AAL38345.1 unknown protein {Ar... 27 1.4
TC93481 26 4.1
TC83474 similar to PIR|T46028|T46028 hypothetical protein T10K17... 26 4.1
BQ751109 26 4.1
TC86605 similar to PIR|T01853|T01853 probable hexose transport p... 26 4.1
TC85671 similar to SP|Q9XF97|RL4_PRUAR 60S ribosomal protein L4 ... 25 5.3
TC82221 25 6.9
TC81081 lectin (LEC1) 25 6.9
TC77838 similar to GP|10998146|dbj|BAB03117. gene_id:MEC18.18~un... 25 6.9
TC89639 weakly similar to GP|20259609|gb|AAM14161.1 unknown prot... 25 6.9
CB895040 25 6.9
AW774950 25 6.9
TC92709 weakly similar to GP|8978342|dbj|BAA98195.1 short chain ... 25 9.0
>CA989819 homologue to SP|Q9BBT8|NU3C_ NADH-plastoquinone oxidoreductase
chain 3 chloroplast (EC 1.6.5.3). {Lotus japonicus},
partial (76%)
Length = 652
Score = 181 bits (459), Expect = 6e-47
Identities = 87/92 (94%), Positives = 88/92 (95%)
Frame = +2
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFW FLIISIFIPILAF ISG LAPI KGPEKLSSYESGIEPMGDAWLQFQI
Sbjct: 377 MFLLYEYDIFWTFLIISIFIPILAFLISGILAPIRKGPEKLSSYESGIEPMGDAWLQFQI 556
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGIS 92
RYYMFALVFVVFDVETVFLYPWAMSFDVLG+S
Sbjct: 557 RYYMFALVFVVFDVETVFLYPWAMSFDVLGVS 652
>AL385275
Length = 482
Score = 29.6 bits (65), Expect = 0.28
Identities = 16/50 (32%), Positives = 24/50 (48%)
Frame = +3
Query: 14 LIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQIRYY 63
+ ISIF I+ + F+ INK PE L S + + + W Q + Y
Sbjct: 3 MFISIFFVIILIVVINFIRKINKKPEGLESIPN-VSTLEILWANLQQKNY 149
>TC89179 similar to GP|10505183|gb|AAG18376.1 lipoxygenase {Zantedeschia
aethiopica}, partial (22%)
Length = 742
Score = 29.3 bits (64), Expect = 0.37
Identities = 17/42 (40%), Positives = 26/42 (61%), Gaps = 1/42 (2%)
Frame = +3
Query: 70 VVFDVETVFLYPWAMSFD-VLGISVFIEALIFVLILIVGSVY 110
+V + TV+LY W++S D VL ++ I LI IL+ +VY
Sbjct: 537 LVVESRTVYLYDWSISIDLVL*YNLLILYLI*YTILVAETVY 662
>BQ151299
Length = 574
Score = 28.1 bits (61), Expect = 0.82
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = -2
Query: 5 YEYDIFWAFLIISIFIPILAFTISGFL 31
Y YDI+W FL + +F+ F GFL
Sbjct: 390 YIYDIYWVFLYVLVFV----FRFIGFL 322
>TC78773 similar to GP|17473842|gb|AAL38345.1 unknown protein {Arabidopsis
thaliana}, partial (90%)
Length = 1449
Score = 27.7 bits (60), Expect = 1.1
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +1
Query: 4 LYEYDIFWAFLIISIFIPILAFTISGFLAPI 34
L+ YDIFW F F P++ F API
Sbjct: 769 LFVYDIFWVF-----FTPVMVSVAKSFDAPI 846
>TC77811 similar to GP|17473842|gb|AAL38345.1 unknown protein {Arabidopsis
thaliana}, partial (91%)
Length = 1284
Score = 27.3 bits (59), Expect = 1.4
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +3
Query: 4 LYEYDIFWAFLIISIFIPILAFTISGFLAPI 34
L+ YDIFW F F P++ F API
Sbjct: 582 LFFYDIFWVF-----FTPVMVSVAKSFDAPI 659
>TC88787 weakly similar to GP|21593424|gb|AAM65391.1 fatty acid
elongase-like protein (cer2-like) {Arabidopsis
thaliana}, partial (28%)
Length = 741
Score = 27.3 bits (59), Expect = 1.4
Identities = 10/24 (41%), Positives = 15/24 (61%)
Frame = +2
Query: 33 PINKGPEKLSSYESGIEPMGDAWL 56
P+N GPEK + ++P GD W+
Sbjct: 653 PLNFGPEKDPATVKRVDPAGDHWI 724
>TC77812 similar to GP|17473842|gb|AAL38345.1 unknown protein {Arabidopsis
thaliana}, partial (74%)
Length = 809
Score = 27.3 bits (59), Expect = 1.4
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +1
Query: 4 LYEYDIFWAFLIISIFIPILAFTISGFLAPI 34
L+ YDIFW F F P++ F API
Sbjct: 604 LFFYDIFWVF-----FTPVMVSVAKSFDAPI 681
>TC93481
Length = 579
Score = 25.8 bits (55), Expect = 4.1
Identities = 10/23 (43%), Positives = 16/23 (69%)
Frame = +3
Query: 9 IFWAFLIISIFIPILAFTISGFL 31
++W+ + SI+ PI+ FTI FL
Sbjct: 495 LYWSKINKSIYSPIIKFTIKDFL 563
>TC83474 similar to PIR|T46028|T46028 hypothetical protein T10K17.270 -
Arabidopsis thaliana, partial (16%)
Length = 814
Score = 25.8 bits (55), Expect = 4.1
Identities = 16/49 (32%), Positives = 26/49 (52%)
Frame = +2
Query: 29 GFLAPINKGPEKLSSYESGIEPMGDAWLQFQIRYYMFALVFVVFDVETV 77
GF PI P+ S+ + G+ +L F I+Y +F LVF + V ++
Sbjct: 443 GFYFPIPPMPKGSSTL*REVHAQGE-FLNFLIKYMLFFLVFSHYVVGSI 586
>BQ751109
Length = 296
Score = 25.8 bits (55), Expect = 4.1
Identities = 8/19 (42%), Positives = 14/19 (73%)
Frame = -1
Query: 68 VFVVFDVETVFLYPWAMSF 86
+ V++DV T+ LYPW ++
Sbjct: 110 LLVLYDVMTISLYPWTRNY 54
>TC86605 similar to PIR|T01853|T01853 probable hexose transport protein
F9D12.17 - Arabidopsis thaliana, partial (91%)
Length = 3111
Score = 25.8 bits (55), Expect = 4.1
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = +3
Query: 97 ALIFVLILIVGSVYAWRKGALEW 119
AL+ V++ I S +AW G L W
Sbjct: 2445 ALVVVMVCIFVSAFAWSWGPLAW 2513
>TC85671 similar to SP|Q9XF97|RL4_PRUAR 60S ribosomal protein L4 (L1).
[Apricot] {Prunus armeniaca}, complete
Length = 1508
Score = 25.4 bits (54), Expect = 5.3
Identities = 17/50 (34%), Positives = 26/50 (52%), Gaps = 2/50 (4%)
Frame = -2
Query: 60 IRYYMF--ALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVG 107
+ Y+ F AL+ V +ETVFL+ + SF L +S I + +L G
Sbjct: 1207 VLYHAFPAALIAVASSLETVFLFLSSFSFLDLTLSASASKAILLAVLA*G 1058
>TC82221
Length = 546
Score = 25.0 bits (53), Expect = 6.9
Identities = 12/21 (57%), Positives = 16/21 (76%)
Frame = -2
Query: 8 DIFWAFLIISIFIPILAFTIS 28
DIF AFLI++I IP + F +S
Sbjct: 80 DIFSAFLIMTIGIPPILFFLS 18
>TC81081 lectin (LEC1)
Length = 859
Score = 25.0 bits (53), Expect = 6.9
Identities = 17/47 (36%), Positives = 22/47 (46%)
Frame = -2
Query: 63 YMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSV 109
++ ALV F VE V PW + F F L FV++ V SV
Sbjct: 234 FLTALVSCNFPVEVVMALPWKIRF-------FWSWLNFVMVKEVSSV 115
>TC77838 similar to GP|10998146|dbj|BAB03117. gene_id:MEC18.18~unknown
protein {Arabidopsis thaliana}, partial (86%)
Length = 1353
Score = 25.0 bits (53), Expect = 6.9
Identities = 7/20 (35%), Positives = 17/20 (85%)
Frame = +3
Query: 97 ALIFVLILIVGSVYAWRKGA 116
+L+F++ L+ G +++W+KG+
Sbjct: 759 SLVFLMGLLQGRIWSWKKGS 818
>TC89639 weakly similar to GP|20259609|gb|AAM14161.1 unknown protein
{Arabidopsis thaliana}, partial (40%)
Length = 1000
Score = 25.0 bits (53), Expect = 6.9
Identities = 10/42 (23%), Positives = 25/42 (58%)
Frame = +2
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLS 42
+ LL ++D+F + +I ++ IP++ + L + P+K++
Sbjct: 650 LMLLLQWDLFSSLIISAVMIPLMMMQLRSLLCIL---PQKIN 766
>CB895040
Length = 755
Score = 25.0 bits (53), Expect = 6.9
Identities = 9/22 (40%), Positives = 16/22 (71%)
Frame = +1
Query: 56 LQFQIRYYMFALVFVVFDVETV 77
L I++Y+F LV V+FD +++
Sbjct: 37 LSHHIKFYIFELVSVIFDCDSL 102
>AW774950
Length = 550
Score = 25.0 bits (53), Expect = 6.9
Identities = 13/40 (32%), Positives = 21/40 (52%)
Frame = +2
Query: 71 VFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVY 110
VFD +Y + ++ ++ + V L FVL+L V S Y
Sbjct: 50 VFDCWATLIYHYTVTLQLVFLLVQQPLLCFVLLLYVSSSY 169
>TC92709 weakly similar to GP|8978342|dbj|BAA98195.1 short chain alcohol
dehydrogenase-like {Arabidopsis thaliana}, partial (75%)
Length = 733
Score = 24.6 bits (52), Expect = 9.0
Identities = 13/29 (44%), Positives = 18/29 (61%)
Frame = +1
Query: 89 LGISVFIEALIFVLILIVGSVYAWRKGAL 117
+G VFI ++ V+ L GSVYA K A+
Sbjct: 511 MGSIVFISSIAGVVSLGTGSVYAASKAAI 597
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.149 0.480
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,035,939
Number of Sequences: 36976
Number of extensions: 49859
Number of successful extensions: 396
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 396
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 396
length of query: 120
length of database: 9,014,727
effective HSP length: 84
effective length of query: 36
effective length of database: 5,908,743
effective search space: 212714748
effective search space used: 212714748
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 52 (24.6 bits)
Description of BAB33181.1

