
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33181.1 120 aa
(120 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
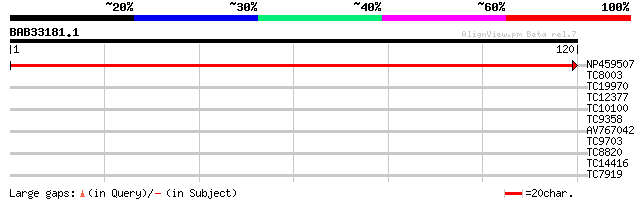
NP459507 NADH dehydrogenase D3 [Lotus japonicus] 244 3e-66
TC8003 homologue to PIR|S52023|S52023 translation initiation fac... 27 0.93
TC19970 similar to UP|Q8W508 (Q8W508) Histone deacetylase, parti... 26 2.1
TC12377 similar to UP|Q9SQF7 (Q9SQF7) Chitinase, partial (7%) 25 2.7
TC10100 25 3.5
TC9358 weakly similar to GB|AAH10781.1|14789753|BC010781 RIO kin... 25 3.5
AV767042 25 4.6
TC9703 similar to UP|Q9XFZ4 (Q9XFZ4) Asparaginyl endopeptidase (... 25 4.6
TC8820 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subuni... 25 4.6
TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, parti... 24 6.0
TC7919 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus cornicula... 24 7.9
>NP459507 NADH dehydrogenase D3 [Lotus japonicus]
Length = 363
Score = 244 bits (623), Expect = 3e-66
Identities = 120/120 (100%), Positives = 120/120 (100%)
Frame = +1
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 60
MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI
Sbjct: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPMGDAWLQFQI 180
Query: 61 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS
Sbjct: 181 RYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 360
>TC8003 homologue to PIR|S52023|S52023 translation initiation factor
eIF-4A.14 - common tobacco {Nicotiana tabacum;} ,
partial (34%)
Length = 708
Score = 26.9 bits (58), Expect = 0.93
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = +2
Query: 88 VLGISVFIEALIFVLILIVGSVYAW 112
VLG+ VF LIF L++++ S Y W
Sbjct: 155 VLGLPVF*SLLIFWLVVLMCSKYLW 229
>TC19970 similar to UP|Q8W508 (Q8W508) Histone deacetylase, partial (17%)
Length = 508
Score = 25.8 bits (55), Expect = 2.1
Identities = 15/51 (29%), Positives = 27/51 (52%)
Frame = -3
Query: 56 LQFQIRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIV 106
L ++I + + F++F + + DVL +SV I ALI VL++ +
Sbjct: 248 LSWRIHLIIVIVTFIIFII----------ALDVLVLSVLINALIRVLLIFI 126
>TC12377 similar to UP|Q9SQF7 (Q9SQF7) Chitinase, partial (7%)
Length = 327
Score = 25.4 bits (54), Expect = 2.7
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +1
Query: 78 FLYPWAMSFDVLGISVFIEALIFVLIL 104
FL+ ++F +L +SV + A FVL+L
Sbjct: 70 FLFSLLLTFSLLSVSVTVTARPFVLVL 150
>TC10100
Length = 572
Score = 25.0 bits (53), Expect = 3.5
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = +3
Query: 13 FLIISIFIPILAFTISGFLAPINKGPEKLSSYE 45
F ++++ P F I FLAP + P L SY+
Sbjct: 90 FPLLTLIAPAPLFLIYPFLAPFHFTPSYLLSYQ 188
>TC9358 weakly similar to GB|AAH10781.1|14789753|BC010781 RIO kinase 2 {Mus
musculus;} , partial (10%)
Length = 1117
Score = 25.0 bits (53), Expect = 3.5
Identities = 14/51 (27%), Positives = 26/51 (50%)
Frame = +1
Query: 53 DAWLQFQIRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLI 103
D WL ++ YY + L + + + + S+ + GI +FI LIF ++
Sbjct: 967 DVWLVIRLMYYQYMLNSGIG--KQIG*VYYKQSYFIYGIVIFILRLIFFIL 1113
>AV767042
Length = 444
Score = 24.6 bits (52), Expect = 4.6
Identities = 13/27 (48%), Positives = 18/27 (66%)
Frame = -2
Query: 13 FLIISIFIPILAFTISGFLAPINKGPE 39
FLII +FIP L ++ L IN+GP+
Sbjct: 86 FLII-VFIPRLTLPVNILLLLINEGPQ 9
>TC9703 similar to UP|Q9XFZ4 (Q9XFZ4) Asparaginyl endopeptidase (VmPE-1),
partial (28%)
Length = 727
Score = 24.6 bits (52), Expect = 4.6
Identities = 11/29 (37%), Positives = 15/29 (50%)
Frame = +3
Query: 27 ISGFLAPINKGPEKLSSYESGIEPMGDAW 55
+ L + KGPE LSS +P+ D W
Sbjct: 126 VGKLLFGMEKGPEVLSSVRPAGQPLVDDW 212
>TC8820 homologue to UP|CAE45585 (CAE45585) Coatomer alpha subunit-like
protein, partial (3%)
Length = 500
Score = 24.6 bits (52), Expect = 4.6
Identities = 10/34 (29%), Positives = 17/34 (49%)
Frame = +1
Query: 2 FLLYEYDIFWAFLIISIFIPILAFTISGFLAPIN 35
F L+ + F F + +F P+L + G P+N
Sbjct: 283 FFLFHFSFFLWFPFLILFAPLLKYKRKGGKKPMN 384
>TC14416 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (28%)
Length = 695
Score = 24.3 bits (51), Expect = 6.0
Identities = 8/14 (57%), Positives = 11/14 (78%)
Frame = -3
Query: 6 EYDIFWAFLIISIF 19
E+ +FW FL+IS F
Sbjct: 507 EFSLFWWFLVISFF 466
>TC7919 GB|AAP47421.1|32331787|AY164847 RTNLB47w {Lotus corniculatus var.
japonicus;} , complete
Length = 1219
Score = 23.9 bits (50), Expect = 7.9
Identities = 15/51 (29%), Positives = 22/51 (42%)
Frame = +3
Query: 1 MFLLYEYDIFWAFLIISIFIPILAFTISGFLAPINKGPEKLSSYESGIEPM 51
+F L EY + ++I L F S + INK P K+ EP+
Sbjct: 393 LFELLEYHLLTLICHLAILALTLLFLWSNGSSLINKSPPKIPQVHIPEEPV 545
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.333 0.149 0.480
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,217,668
Number of Sequences: 28460
Number of extensions: 26070
Number of successful extensions: 202
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 201
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 201
length of query: 120
length of database: 4,897,600
effective HSP length: 79
effective length of query: 41
effective length of database: 2,649,260
effective search space: 108619660
effective search space used: 108619660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 49 (23.5 bits)
Description of BAB33181.1

