
BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= BAB33181.1 120 aa
(120 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
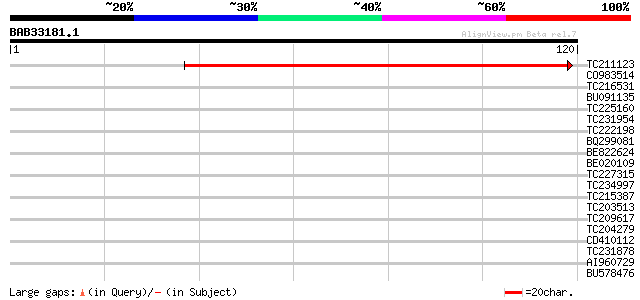
TC211123 homologue to UP|RT12_HELAN (Q96033) Mitochondrial ribos... 95 5e-21
CO983514 29 0.27
TC216531 similar to UP|Q9VPQ7 (Q9VPQ7) CG11840-PA (SD07518p), pa... 28 0.79
BU091135 homologue to GP|10241626|emb Contains similarity to F1N... 28 0.79
TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, comp... 27 1.0
TC231954 similar to GB|AAO42802.1|28416543|BT004556 At1g54520/F2... 27 1.0
TC222198 weakly similar to UP|Q6NLH0 (Q6NLH0) At3g16980, partial... 27 1.8
BQ299081 similar to GP|2827699|emb predicted protein {Arabidopsi... 26 3.0
BE822624 similar to GP|13374872|emb mannosyltransferase-like pro... 26 3.0
BE020109 26 3.0
TC227315 similar to UP|Q9ZRP8 (Q9ZRP8) Delta-8 sphingolipid desa... 26 3.0
TC234997 similar to GB|AAQ56804.1|34098843|BT010361 At1g63770 {A... 26 3.0
TC215387 similar to UP|Q940S9 (Q940S9) AT3g57890/T10K17_100, par... 25 3.9
TC203513 similar to UP|Q8L5P7 (Q8L5P7) LHY protein, partial (42%) 25 3.9
TC209617 25 3.9
TC204279 similar to UP|Q9XFZ4 (Q9XFZ4) Asparaginyl endopeptidase... 25 3.9
CD410112 25 3.9
TC231878 similar to UP|Q9SY30 (Q9SY30) T17H7.16, partial (57%) 25 3.9
AI960729 similar to GP|21593313|gb| protein disulfide isomerase ... 25 3.9
BU578476 25 5.1
>TC211123 homologue to UP|RT12_HELAN (Q96033) Mitochondrial ribosomal protein
S12, partial (92%)
Length = 689
Score = 94.7 bits (234), Expect = 5e-21
Identities = 37/82 (45%), Positives = 60/82 (73%)
Frame = +2
Query: 38 PEKLSSYESGIEPMGDAWLQFQIRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEA 97
PEKLS+YE G +P GDA +F IR+Y+ +++F++FD+E F +PWA+S + + + F
Sbjct: 44 PEKLSAYECGFDPFGDARSRFDIRFYLVSILFIIFDLEVTFFFPWAVSLNKIDLFGFWSM 223
Query: 98 LIFVLILIVGSVYAWRKGALEW 119
+ F+LIL +G +Y W++GAL+W
Sbjct: 224 MAFLLILTIGFLYEWKRGALDW 289
>CO983514
Length = 575
Score = 29.3 bits (64), Expect = 0.27
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 3/43 (6%)
Frame = -2
Query: 81 PWAMSFDVLGISV---FIEALIFVLILIVGSVYAWRKGALEWS 120
PWA SF + I V FI+A+++V+I Y W + WS
Sbjct: 475 PWAYSFAQVLIEVPYIFIQAVVYVIITYPMLSYDWSAYKIFWS 347
>TC216531 similar to UP|Q9VPQ7 (Q9VPQ7) CG11840-PA (SD07518p), partial (11%)
Length = 942
Score = 27.7 bits (60), Expect = 0.79
Identities = 13/31 (41%), Positives = 16/31 (50%)
Frame = +3
Query: 4 LYEYDIFWAFLIISIFIPILAFTISGFLAPI 34
L+ YDIFW F F P++ F API
Sbjct: 450 LFVYDIFWVF-----FTPVMVSVAKSFDAPI 527
>BU091135 homologue to GP|10241626|emb Contains similarity to F1N 19.7 {Oryza
sativa (indica cultivar-group)}, partial (3%)
Length = 421
Score = 27.7 bits (60), Expect = 0.79
Identities = 12/30 (40%), Positives = 20/30 (66%), Gaps = 1/30 (3%)
Frame = -1
Query: 91 ISVFIEALIFVLILIVG-SVYAWRKGALEW 119
+ +F + + +LIL+V SV+ WRK +EW
Sbjct: 247 VGIFTGSEVVLLILLVNMSVHRWRK*GIEW 158
>TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, complete
Length = 2152
Score = 27.3 bits (59), Expect = 1.0
Identities = 15/55 (27%), Positives = 26/55 (47%)
Frame = -3
Query: 58 FQIRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAW 112
F + + L+F++ + T L WA +++ I LILI+ V+AW
Sbjct: 656 FALSLLLLILIFLLLSLFTFLLMRWAWEWEL--------TFILFLILIMRLVWAW 516
>TC231954 similar to GB|AAO42802.1|28416543|BT004556 At1g54520/F20D21_34
{Arabidopsis thaliana;} , partial (6%)
Length = 615
Score = 27.3 bits (59), Expect = 1.0
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 31 LAPINKGPEKLSSYESGIEPMGDAWLQFQIRYY 63
L ++K KLSS E+ + MG+AW +F + Y+
Sbjct: 327 LPSVSKRHGKLSS*ET*MIEMGNAWSRFVLNYF 425
>TC222198 weakly similar to UP|Q6NLH0 (Q6NLH0) At3g16980, partial (69%)
Length = 810
Score = 26.6 bits (57), Expect = 1.8
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = +1
Query: 69 FVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIV 106
F+V D E VFLY VFI + +VL+LIV
Sbjct: 64 FLVIDYEFVFLY------------VFINCICYVLLLIV 141
>BQ299081 similar to GP|2827699|emb predicted protein {Arabidopsis thaliana},
partial (5%)
Length = 422
Score = 25.8 bits (55), Expect = 3.0
Identities = 13/58 (22%), Positives = 31/58 (53%)
Frame = +1
Query: 63 YMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVYAWRKGALEWS 120
Y F + +FD+ +++Y + + + I ++I +L+L+ +++ + LEWS
Sbjct: 73 YQF*FMNFIFDMYFIYIYIYIYIYIYIYIYIYISTYRNMLLLL--NIWHQARQLLEWS 240
>BE822624 similar to GP|13374872|emb mannosyltransferase-like protein
{Arabidopsis thaliana}, partial (15%)
Length = 563
Score = 25.8 bits (55), Expect = 3.0
Identities = 15/52 (28%), Positives = 27/52 (51%)
Frame = -1
Query: 55 WLQFQIRYYMFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIV 106
W+ Q + M+ + + F + VFL WA +F+ A IF+L++I+
Sbjct: 503 WIGAQTHWLMWGYL-LEFKGKNVFLQLWAAGL------LFLAANIFILVMII 369
>BE020109
Length = 422
Score = 25.8 bits (55), Expect = 3.0
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = -1
Query: 82 WAMSFDVLGISVFIEALIFVLILIVGSVYAWR 113
W M + LG + +E I + +LI+G WR
Sbjct: 356 WMMPYS*LGPGLKLEIKILISLLIIGLPTLWR 261
>TC227315 similar to UP|Q9ZRP8 (Q9ZRP8) Delta-8 sphingolipid desaturase,
partial (26%)
Length = 732
Score = 25.8 bits (55), Expect = 3.0
Identities = 8/20 (40%), Positives = 16/20 (80%)
Frame = +2
Query: 1 MFLLYEYDIFWAFLIISIFI 20
+ +LY YDIF++FL+ + ++
Sbjct: 656 LLILYFYDIFYSFLVKNFYV 715
>TC234997 similar to GB|AAQ56804.1|34098843|BT010361 At1g63770 {Arabidopsis
thaliana;} , partial (4%)
Length = 781
Score = 25.8 bits (55), Expect = 3.0
Identities = 12/46 (26%), Positives = 24/46 (52%), Gaps = 5/46 (10%)
Frame = -3
Query: 55 WLQFQIRYYMFALVFVVF-----DVETVFLYPWAMSFDVLGISVFI 95
W + +R + L F++F + + ++ W SFD+ +S+FI
Sbjct: 764 WKKR*VRVLLLLLQFILF*CIIITMSCLHIHSWNYSFDIGHVSIFI 627
>TC215387 similar to UP|Q940S9 (Q940S9) AT3g57890/T10K17_100, partial (44%)
Length = 1105
Score = 25.4 bits (54), Expect = 3.9
Identities = 15/57 (26%), Positives = 26/57 (45%)
Frame = -3
Query: 34 INKGPEKLSSYESGIEPMGDAWLQFQIRYYMFALVFVVFDVETVFLYPWAMSFDVLG 90
+N G + E+ P+G W QFQ+ Y F + ++ E +Y + +V G
Sbjct: 425 MNHGDQPCPMLEA--HPIG*LWAQFQLHSYAFPIGSKMYYREPPAVYDYHQE*EVAG 261
>TC203513 similar to UP|Q8L5P7 (Q8L5P7) LHY protein, partial (42%)
Length = 1163
Score = 25.4 bits (54), Expect = 3.9
Identities = 14/41 (34%), Positives = 26/41 (63%)
Frame = -3
Query: 70 VVFDVETVFLYPWAMSFDVLGISVFIEALIFVLILIVGSVY 110
V+F VFL+ + + L + +FI +I+VL++I+G V+
Sbjct: 1143 VIFYK*QVFLHRFIIRTAFLAVKIFI-FVIYVLLVIIGVVH 1024
>TC209617
Length = 1092
Score = 25.4 bits (54), Expect = 3.9
Identities = 15/39 (38%), Positives = 21/39 (53%), Gaps = 1/39 (2%)
Frame = +1
Query: 77 VFLYPWAMSFDVL-GISVFIEALIFVLILIVGSVYAWRK 114
+FLY AMS DVL +F+ + ++ VYA RK
Sbjct: 862 LFLYLLAMSLDVLTSKGIFVFVFVHT*VMTTCLVYAQRK 978
>TC204279 similar to UP|Q9XFZ4 (Q9XFZ4) Asparaginyl endopeptidase (VmPE-1),
complete
Length = 1868
Score = 25.4 bits (54), Expect = 3.9
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +1
Query: 34 INKGPEKLSSYESGIEPMGDAW 55
I KGPE LSS +P+ D W
Sbjct: 1288 IEKGPELLSSVRPAGQPLVDDW 1353
>CD410112
Length = 661
Score = 25.4 bits (54), Expect = 3.9
Identities = 15/35 (42%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Frame = +2
Query: 67 LVFVVFDVETVFLYPWAMS-FDVLGISVFIEALIF 100
++FVVF +V L PW +S F + GIS F+ + F
Sbjct: 185 VIFVVFVFMSVEL-PWKLSNFTIPGISKFLSKICF 286
>TC231878 similar to UP|Q9SY30 (Q9SY30) T17H7.16, partial (57%)
Length = 662
Score = 25.4 bits (54), Expect = 3.9
Identities = 15/35 (42%), Positives = 22/35 (62%), Gaps = 1/35 (2%)
Frame = -3
Query: 67 LVFVVFDVETVFLYPWAMS-FDVLGISVFIEALIF 100
++FVVF +V L PW +S F + GIS F+ + F
Sbjct: 576 VIFVVFVFMSVEL-PWKLSNFTIPGISKFLSKICF 475
>AI960729 similar to GP|21593313|gb| protein disulfide isomerase
precursor-like {Arabidopsis thaliana}, partial (6%)
Length = 438
Score = 25.4 bits (54), Expect = 3.9
Identities = 18/50 (36%), Positives = 27/50 (54%), Gaps = 1/50 (2%)
Frame = -3
Query: 67 LVFVVFDVETVFLYPWAMSFDVLGISVFI-EALIFVLILIVGSVYAWRKG 115
LV V VE V ++ WA+ + F+ EA IFV++++V V R G
Sbjct: 289 LVVAVVLVELVGIWKWALVRSGGSVIGFLEEAEIFVVVVVVFFVVVLRGG 140
>BU578476
Length = 407
Score = 25.0 bits (53), Expect = 5.1
Identities = 12/41 (29%), Positives = 25/41 (60%)
Frame = +1
Query: 64 MFALVFVVFDVETVFLYPWAMSFDVLGISVFIEALIFVLIL 104
+F +F++F++E +F + + V+ +F + IF+LIL
Sbjct: 133 LF*FLFILFEMEFLFSTDFLGNKRVVKYCLFFNSEIFILIL 255
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.333 0.149 0.480
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,710,307
Number of Sequences: 63676
Number of extensions: 67977
Number of successful extensions: 557
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 554
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 556
length of query: 120
length of database: 12,639,632
effective HSP length: 96
effective length of query: 24
effective length of database: 6,526,736
effective search space: 156641664
effective search space used: 156641664
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 51 (24.3 bits)
Description of BAB33181.1

