

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0246c.3
(286 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
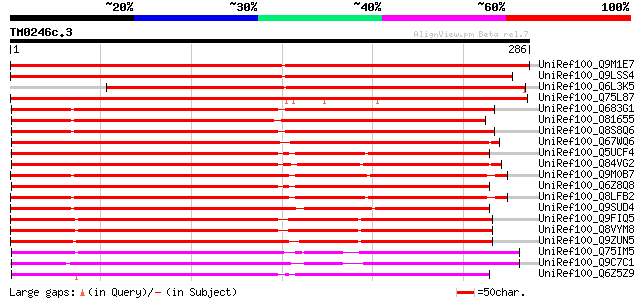
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9M1E7 Hypothetical protein F9K21.180 [Arabidopsis tha... 472 e-132
UniRef100_Q9LSS4 Similarity to senescence-associated protein [Ar... 424 e-117
UniRef100_Q6L3K5 Putative senescence-associated protein [Solanum... 363 2e-99
UniRef100_Q75L87 Hypothetical protein OJ1729_E02.7 [Oryza sativa] 354 1e-96
UniRef100_Q683G1 Similar to senescence-associated protein [Arabi... 276 3e-73
UniRef100_O81655 Senescence-associated protein 5 [Hemerocallis sp.] 276 3e-73
UniRef100_Q8S8Q6 Hypothetical protein At2g23810 [Arabidopsis tha... 276 4e-73
UniRef100_Q67WQ6 Putative senescence-associated protein 5 [Oryza... 273 3e-72
UniRef100_Q5UCF4 Senescence-associated protein DH [Zea mays] 273 4e-72
UniRef100_Q84VG2 Putative senescence-associated protein 5 [Oryza... 263 4e-69
UniRef100_Q9M0B7 Senescence-associated protein homolog [Arabidop... 254 1e-66
UniRef100_Q6Z8Q8 Putative senescence-associated protein [Oryza s... 254 2e-66
UniRef100_Q8LFB2 Senescence-associated protein-like protein [Ara... 252 9e-66
UniRef100_Q9SUD4 Senescence-associated protein-like [Arabidopsis... 249 4e-65
UniRef100_Q9FIQ5 Senescence-associated protein 5-like protein [A... 243 5e-63
UniRef100_Q8VYM8 Putative senescence-associated protein 5 [Arabi... 242 7e-63
UniRef100_Q9ZUN5 Putative senescence-associated protein 5 [Arabi... 241 1e-62
UniRef100_Q75IM5 Putative senescence-associated protein [Oryza s... 232 1e-59
UniRef100_Q9C7C1 Senescence-assocated protein, putative; 28418-2... 231 2e-59
UniRef100_Q6Z5Z9 Putative senescence-associated protein [Oryza s... 219 8e-56
>UniRef100_Q9M1E7 Hypothetical protein F9K21.180 [Arabidopsis thaliana]
Length = 285
Score = 472 bits (1215), Expect = e-132
Identities = 222/286 (77%), Positives = 252/286 (87%), Gaps = 1/286 (0%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
MRTSNHLIG++NFLTFLLSIPILGGGIWLSSRAN+TDCL+FLQWPLI+IG+SIMVVSLAG
Sbjct: 1 MRTSNHLIGLVNFLTFLLSIPILGGGIWLSSRANSTDCLRFLQWPLIVIGISIMVVSLAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
FAGACYRN FLM LYLVVM L+IA LIGFIIFAY VTDKGSGR VLNRGY+DYYL+DYSG
Sbjct: 61 FAGACYRNKFLMWLYLVVMLLIIAALIGFIIFAYAVTDKGSGRTVLNRGYLDYYLEDYSG 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL++RV+ SYWGKI+SC+RDS C K+GR + NG+PE AD+F+LR+L+ V+SGCCKPPT
Sbjct: 121 WLKDRVSDDSYWGKISSCLRDSGACRKIGR-NFNGVPETADMFFLRRLSPVESGCCKPPT 179
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DCG+ Y NET W+ G++ N DC WSNDQ LCY+C SCKAGVL SLKKSWRKVSVI
Sbjct: 180 DCGFSYVNETGWDTRGGMIGPNQDCMVWSNDQSMLCYQCSSCKAGVLGSLKKSWRKVSVI 239
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPSHFQL 286
NIVV+IILVI Y+IAYAAYRN KR+DNDEP GEARMTKSHPSHF L
Sbjct: 240 NIVVLIILVIFYVIAYAAYRNVKRIDNDEPAGEARMTKSHPSHFHL 285
>UniRef100_Q9LSS4 Similarity to senescence-associated protein [Arabidopsis thaliana]
Length = 327
Score = 424 bits (1090), Expect = e-117
Identities = 203/277 (73%), Positives = 230/277 (82%), Gaps = 1/277 (0%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
MR+ ++LIG++NF TFLLSIPILGGGIWLSSRAN+TDCL+FLQWPLIIIG+SIMV+SLAG
Sbjct: 1 MRSRSNLIGLINFFTFLLSIPILGGGIWLSSRANSTDCLRFLQWPLIIIGISIMVISLAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
AGACY+N FLM LYL MF VIA LIGF IFAYVVTDKGSGR V+NR Y+DYYL DYSG
Sbjct: 61 IAGACYQNKFLMWLYLFTMFFVIAALIGFTIFAYVVTDKGSGRFVMNRRYLDYYLNDYSG 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL++RV + YW I SCVRDS VC K+GR D NG+PE A +FY R L+ V+SGCCKPPT
Sbjct: 121 WLKDRVTDNGYWRDIGSCVRDSGVCKKIGR-DLNGVPETAHMFYFRNLSPVESGCCKPPT 179
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DCGY Y NETVW G ++ NPDC W+NDQ LCY+C SCKAGVL SLKKSWRKVSVI
Sbjct: 180 DCGYTYVNETVWIPGGEMVGPNPDCMLWNNDQRLLCYQCSSCKAGVLGSLKKSWRKVSVI 239
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMT 277
NIVV+IILVI Y+IA AAY+N KRM NDEP GEARMT
Sbjct: 240 NIVVVIILVIFYVIACAAYQNVKRMYNDEPVGEARMT 276
>UniRef100_Q6L3K5 Putative senescence-associated protein [Solanum demissum]
Length = 232
Score = 363 bits (933), Expect = 2e-99
Identities = 169/233 (72%), Positives = 198/233 (84%), Gaps = 3/233 (1%)
Query: 54 MVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDY 113
MVVSLAGF GACYRNTFLM +YL MF +IA LIGF+IFA+ VTDKGSGR V+NR Y +Y
Sbjct: 1 MVVSLAGFVGACYRNTFLMYIYLWAMFFIIAALIGFVIFAFAVTDKGSGRPVMNRAYSEY 60
Query: 114 YLQDYSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQS 173
YLQDYSGWLEERV+S SYW KI+SC+RDS C KM R+ NGIPE ++FYLRKL+ ++S
Sbjct: 61 YLQDYSGWLEERVSSQSYWEKISSCIRDSHSCGKMRRI-YNGIPESVEMFYLRKLSPIES 119
Query: 174 GCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKS 233
GCCKPPT+CGY+YQNETVW G GL+ A+PDC KWSNDQ LCY CDSCKAGVLASLKKS
Sbjct: 120 GCCKPPTECGYVYQNETVWIPGGGLVGADPDCVKWSNDQEQLCYNCDSCKAGVLASLKKS 179
Query: 234 WRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPS--HF 284
WRKVSVINIV++I+LVI+Y++A AA+R+NKR+DNDEPYGE RM K+ PS HF
Sbjct: 180 WRKVSVINIVILILLVIMYMVAIAAFRHNKRIDNDEPYGETRMEKTQPSRIHF 232
>UniRef100_Q75L87 Hypothetical protein OJ1729_E02.7 [Oryza sativa]
Length = 339
Score = 354 bits (909), Expect = 1e-96
Identities = 171/337 (50%), Positives = 226/337 (66%), Gaps = 52/337 (15%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
+R L+G++NF+TFL+SIPILGGGIWL+SRAN+TDC++FLQWP+I IG+++MVVSL G
Sbjct: 2 LRGGTSLLGIVNFVTFLISIPILGGGIWLASRANSTDCIRFLQWPIIAIGLAVMVVSLMG 61
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
FAGACYR T+L+RLYL MF ++ L+ FI+FA+ VTD+G G+ V+NR +++Y L DY+G
Sbjct: 62 FAGACYRQTWLLRLYLFAMFFIVVALLFFIVFAFAVTDRGDGQVVMNRRFLEYQLSDYNG 121
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRV--DSNG---IPEPADVFYLRKLTSVQ--- 172
WL +RVA +YW I++C+RD + CA M R D N +PE +FY R L+ +Q
Sbjct: 122 WLRDRVADPAYWATISACLRDGRACAAMRRFARDPNTGMLVPETPSMFYARDLSPIQVTL 181
Query: 173 ------------------------------------------SGCCKPPTDCGYIYQNET 190
SGCCKPPT C Y Y NET
Sbjct: 182 LVQLLRPNNTKTFQSCHYCSGYLFMYSVIFVFVAVDKTWQSKSGCCKPPTSCAYNYVNET 241
Query: 191 VWNLGSGLMSA--NPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIIL 248
W G+ + + DCSKWSNDQ LC++CDSCKAGVLA +KKSWRKV+++NIVV+IIL
Sbjct: 242 FWTANPGVPTVVNDVDCSKWSNDQQTLCFQCDSCKAGVLAGIKKSWRKVAILNIVVLIIL 301
Query: 249 VIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPSHFQ 285
VIVY+ AA+RN +R++NDEP+G ARMTK+ PS FQ
Sbjct: 302 VIVYVAGCAAFRNARRIENDEPFGMARMTKTQPSRFQ 338
>UniRef100_Q683G1 Similar to senescence-associated protein [Arabidopsis thaliana]
Length = 273
Score = 276 bits (707), Expect = 3e-73
Identities = 128/266 (48%), Positives = 186/266 (69%), Gaps = 4/266 (1%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN+L+G+LNFL FLLSIPIL GGIWLS + + T+C +FL P+I +GV +MVV++AG
Sbjct: 3 RCSNNLVGILNFLVFLLSIPILAGGIWLSQKGS-TECERFLDKPVIALGVFLMVVAIAGL 61
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
G+C R T+L+ +YL VMFL+I ++ +FA+VVT+KG+G + +GY +Y L DYS W
Sbjct: 62 VGSCCRVTWLLWVYLFVMFLLILLVFCITVFAFVVTNKGAGEAIEGKGYKEYKLGDYSTW 121
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L++RV + W KI SC+ +SKVC+K+ ++ + P + FY LT++QSGCCKP +
Sbjct: 122 LQKRVENGKNWNKIRSCLVESKVCSKL---EAKFVNVPVNSFYKEHLTALQSGCCKPSDE 178
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
CG+ Y N T W + NPDC W N + LC+ C SCKAG+L ++K +W+KV+++N
Sbjct: 179 CGFEYVNPTTWTKNTTGTHTNPDCQTWDNAKEKLCFDCQSCKAGLLDNVKSAWKKVAIVN 238
Query: 242 IVVMIILVIVYIIAYAAYRNNKRMDN 267
IV ++ L+IVY + A+RNNKR D+
Sbjct: 239 IVFLVFLIIVYSVGCCAFRNNKRDDS 264
>UniRef100_O81655 Senescence-associated protein 5 [Hemerocallis sp.]
Length = 275
Score = 276 bits (707), Expect = 3e-73
Identities = 130/262 (49%), Positives = 181/262 (68%), Gaps = 5/262 (1%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN+L+G+LNF+T LL+IPIL G++LS ++ TDC FLQ P++ +G+ +++VSLAG
Sbjct: 5 RLSNNLVGILNFVTLLLTIPILSTGVYLSHHSS-TDCAHFLQGPILGLGIVLLLVSLAGL 63
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC +N+ L+ +YL VMF++I V+ F IF +VVT+KG+G V RGY DY L DYS W
Sbjct: 64 VGACCKNSLLLWIYLFVMFVLIIVIFCFTIFVFVVTNKGAGEVVSGRGYKDYRLGDYSHW 123
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L++R+ + W KI SC+ D KVC R ++G D F L+ +QSGCCKPPT+
Sbjct: 124 LQKRMENGKNWRKIRSCLVDGKVCE---RFSTDGTNRTLDQFVKENLSPIQSGCCKPPTE 180
Query: 182 CGYIYQNETVWNL-GSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
CG+ YQ+ TVWN +G S N DC+ W ND LCY C SCK GV+A+LK W+KV+V+
Sbjct: 181 CGFTYQSPTVWNKPATGFTSNNTDCATWENDPTILCYDCQSCKGGVIANLKSKWKKVAVV 240
Query: 241 NIVVMIILVIVYIIAYAAYRNN 262
NI+ +I ++IVY + A+RNN
Sbjct: 241 NIIFLIFIIIVYSVGCCAFRNN 262
>UniRef100_Q8S8Q6 Hypothetical protein At2g23810 [Arabidopsis thaliana]
Length = 273
Score = 276 bits (706), Expect = 4e-73
Identities = 128/266 (48%), Positives = 186/266 (69%), Gaps = 4/266 (1%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN+L+G+LNFL FLLSIPIL GGIWLS + + T+C +FL P+I +GV +MVV++AG
Sbjct: 3 RCSNNLVGILNFLVFLLSIPILAGGIWLSQKGS-TECERFLDKPVIALGVFLMVVAIAGL 61
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
G+C R T+L+ +YL VMFL+I ++ +FA+VVT+KG+G + +GY +Y L DYS W
Sbjct: 62 IGSCCRVTWLLWVYLFVMFLLILLVFCITVFAFVVTNKGAGEAIEGKGYKEYKLGDYSTW 121
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L++RV + W KI SC+ +SKVC+K+ ++ + P + FY LT++QSGCCKP +
Sbjct: 122 LQKRVENGKNWNKIRSCLVESKVCSKL---EAKFVNVPVNSFYKEHLTALQSGCCKPSDE 178
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
CG+ Y N T W + NPDC W N + LC+ C SCKAG+L ++K +W+KV+++N
Sbjct: 179 CGFEYVNPTTWTKNTTGTHTNPDCQTWDNAKEKLCFDCQSCKAGLLDNVKSAWKKVAIVN 238
Query: 242 IVVMIILVIVYIIAYAAYRNNKRMDN 267
IV ++ L+IVY + A+RNNKR D+
Sbjct: 239 IVFLVFLIIVYSVGCCAFRNNKRDDS 264
>UniRef100_Q67WQ6 Putative senescence-associated protein 5 [Oryza sativa]
Length = 269
Score = 273 bits (699), Expect = 3e-72
Identities = 133/270 (49%), Positives = 183/270 (67%), Gaps = 7/270 (2%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN L+G+LN +L++ +LGGGIWLS+RA TDC +F++ P++ +GV ++ +SLAG
Sbjct: 3 RCSNGLLGLLNAGVLVLAVVVLGGGIWLSNRAATTDCERFMERPVVALGVLLLALSLAGL 62
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
AGA + L+ LYL+ +FL+I L F +FA+VVT++G+G V RGY +Y L DYS W
Sbjct: 63 AGALCGASCLLWLYLLALFLLILALFVFTVFAFVVTNRGAGWVVSGRGYREYRLGDYSTW 122
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L+ RV + + W KI SC++D KVC K+G E D F L+ +QSGCCKPPT
Sbjct: 123 LQRRVENSANWAKIRSCLQDGKVCEKLG-----ARRETMDQFVGSNLSPIQSGCCKPPTG 177
Query: 182 CGYIYQNETVWNLGSGLMSA-NPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
C + Y +ETVW SG S +PDC+ WSNDQ LCY C SCKAGVLA+LK W+K++ +
Sbjct: 178 CNFAYVSETVWTKPSGFNSTDDPDCTTWSNDQTALCYDCQSCKAGVLANLKNDWKKIATV 237
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEP 270
NI+ +I L+IVY + A+RNN+R DN P
Sbjct: 238 NIIFLIFLIIVYSVGCCAFRNNRR-DNSYP 266
>UniRef100_Q5UCF4 Senescence-associated protein DH [Zea mays]
Length = 273
Score = 273 bits (698), Expect = 4e-72
Identities = 130/263 (49%), Positives = 180/263 (68%), Gaps = 6/263 (2%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN+LIGVLNF+TFLLS+PILG GIWL RA+ T+C ++L P+I +G ++ VSLAG
Sbjct: 4 RLSNNLIGVLNFVTFLLSVPILGAGIWLGHRADGTECERYLSAPVIALGAFLLAVSLAGL 63
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC R T+L+ YL+ MF++I VL F +FA+VVT++G+G V RGY +Y L DYS W
Sbjct: 64 VGACCRVTWLLWAYLLAMFVLILVLFCFTVFAFVVTNRGAGEAVSGRGYKEYRLGDYSNW 123
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L++RV + W +I SC++DSKVC + D N E F L+ ++SGCCKPPT
Sbjct: 124 LQKRVENTRNWDRIRSCLQDSKVCKSL--QDKN---ETVAQFMSSSLSPIESGCCKPPTS 178
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
CGY Y T W + S +PDC WSND LCY C SCKAGV+A+ +++W++V+V+
Sbjct: 179 CGYTYVGGTDWTPVT-TNSTDPDCKTWSNDASALCYNCQSCKAGVVATFQRNWKRVAVVC 237
Query: 242 IVVMIILVIVYIIAYAAYRNNKR 264
IV ++ ++IVY + A+RNN+R
Sbjct: 238 IVFLVFIIIVYSVGCCAFRNNRR 260
>UniRef100_Q84VG2 Putative senescence-associated protein 5 [Oryza sativa]
Length = 276
Score = 263 bits (672), Expect = 4e-69
Identities = 127/270 (47%), Positives = 183/270 (67%), Gaps = 7/270 (2%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN L+G+LN +TFLLS+P+LGGGIWL++RA+ T+C ++ P+I GV +++VSLAG
Sbjct: 4 RLSNSLLGILNAVTFLLSVPVLGGGIWLATRADGTECERYFSAPVIAFGVFLLLVSLAGL 63
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC R L+ YLV MF++I VL F +FA+VVT+KG+G V RGY +Y L DYS W
Sbjct: 64 VGACCRVNCLLWFYLVAMFVLIVVLFCFTVFAFVVTNKGAGEAVSGRGYKEYRLGDYSNW 123
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L++R+ + W +I SC++DSKVC K+ D N F+ L+ ++SGCCKPP+
Sbjct: 124 LQKRMENSKNWNRIRSCLQDSKVCKKL--QDKNW---DRTQFFKADLSPLESGCCKPPSS 178
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
C ++Y + T W S++PDC+ W +D LCY C SCKAG +A+LK+ W++V+V+
Sbjct: 179 CNFLYVSGTNWT-KVPTNSSDPDCNTWVDDGTQLCYNCQSCKAGAVATLKRDWKRVAVVC 237
Query: 242 IVVMIILVIVYIIAYAAYRNNKRMDNDEPY 271
IV ++ +VIVY + A+RNN+R DN Y
Sbjct: 238 IVFLVFIVIVYSLGCCAFRNNRR-DNRGAY 266
>UniRef100_Q9M0B7 Senescence-associated protein homolog [Arabidopsis thaliana]
Length = 272
Score = 254 bits (650), Expect = 1e-66
Identities = 121/274 (44%), Positives = 176/274 (64%), Gaps = 9/274 (3%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
+R SN L+G+LNF FLLS+PIL GIWLS +A T C +FL P+I +GV +M++++AG
Sbjct: 2 VRFSNSLVGILNFFVFLLSVPILSTGIWLSLKAT-TQCERFLDKPMIALGVFLMIIAIAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
G+C R T+L+ YL VMF +I +++ F IFA+VVT KGSG + + Y +Y L+ YS
Sbjct: 61 VVGSCCRVTWLLWSYLFVMFFLILIVLCFTIFAFVVTSKGSGETIQGKAYKEYRLEAYSD 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL+ RV + +W I SC+ +SK C + V +N FY LT+ +SGCCKP
Sbjct: 121 WLQRRVNNAKHWNSIRSCLYESKFCYNLELVTAN---HTVSDFYKEDLTAFESGCCKPSN 177
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DC + Y T WN SG N DC W N++ LCY C +CKAG L +LK +W++V+++
Sbjct: 178 DCDFTYITSTTWNKTSG-THKNSDCQLWDNEKHKLCYNCKACKAGFLDNLKAAWKRVAIV 236
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEA 274
NI+ +++LV+VY + A+RNNK ++ YG +
Sbjct: 237 NIIFLVLLVVVYAMGCCAFRNNK----EDRYGRS 266
>UniRef100_Q6Z8Q8 Putative senescence-associated protein [Oryza sativa]
Length = 277
Score = 254 bits (648), Expect = 2e-66
Identities = 121/263 (46%), Positives = 169/263 (64%), Gaps = 5/263 (1%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN++IG LN +T LLS PIL GGIW+++R + +C + L P I +G +M VSLAG
Sbjct: 4 RLSNNVIGALNLVTLLLSAPILSGGIWMATRGDGGECDRHLSSPAIALGAVLMAVSLAGL 63
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC R T+L+ +YL+ MF +I L+GF FA+ VT++G+G V RGY +Y L DYS W
Sbjct: 64 VGACCRVTWLLWVYLLAMFALIVALLGFTAFAFAVTNRGAGEAVSGRGYREYRLGDYSTW 123
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L V S W KI SC+ + VC + D N E F L+ VQSGCCKPPT
Sbjct: 124 LRRHVGSSKNWDKIRSCLAGADVCRSL--QDRN---ETWAQFVADDLSPVQSGCCKPPTS 178
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
C + Y T W + L SA+PDC +WSND +CY C SCKAGV+A+LK+ W++V+++N
Sbjct: 179 CNFTYGGGTRWGKTARLSSADPDCDEWSNDADEVCYGCRSCKAGVVAALKRDWKRVAIVN 238
Query: 242 IVVMIILVIVYIIAYAAYRNNKR 264
+V + +V+VY + A++N++R
Sbjct: 239 VVFLAFIVVVYSVGCCAFKNSRR 261
>UniRef100_Q8LFB2 Senescence-associated protein-like protein [Arabidopsis thaliana]
Length = 272
Score = 252 bits (643), Expect = 9e-66
Identities = 120/274 (43%), Positives = 175/274 (63%), Gaps = 9/274 (3%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
+R SN L+G+LNF FLLS+PIL GIWLS +A T C +FL P+I +GV +M++++AG
Sbjct: 2 VRFSNSLVGILNFFVFLLSVPILSTGIWLSLKAT-TQCARFLDKPMIALGVFLMIIAIAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
G+C R T+L+ YL VMF +I +++ F IFA+VVT KGSG + + Y +Y L+ YS
Sbjct: 61 VVGSCSRVTWLLWSYLFVMFFLILIVLCFTIFAFVVTSKGSGETIQGKAYKEYRLEAYSD 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL+ RV + +W I SC+ +SK C + V +N FY LT+ +SGCCKP
Sbjct: 121 WLQRRVNNAKHWNSIRSCLYESKFCYNLELVTAN---HTVSDFYKEDLTAFESGCCKPSN 177
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DC + Y T WN S N DC W N++ LCY C +CKAG L +LK +W++V+++
Sbjct: 178 DCDFTYITSTTWNKTS-RTHKNSDCQLWDNEKHKLCYNCKACKAGFLDNLKAAWKRVAIV 236
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEA 274
NI+ +++LV+VY + A+RNNK ++ YG +
Sbjct: 237 NIIFLVLLVVVYAMRCCAFRNNK----EDRYGRS 266
>UniRef100_Q9SUD4 Senescence-associated protein-like [Arabidopsis thaliana]
Length = 263
Score = 249 bits (637), Expect = 4e-65
Identities = 121/265 (45%), Positives = 174/265 (65%), Gaps = 7/265 (2%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
++ SN+L+G+LNF TFLLSIPIL GIWL A T+C +FL P++++G+ +M VS+AG
Sbjct: 2 VQCSNNLLGILNFFTFLLSIPILSAGIWLGKNAA-TECERFLDKPMVVLGIFLMFVSIAG 60
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
GAC R + L+ LYL MFL+I + F IFA+ VT++G+G + +RGY +Y++ DYS
Sbjct: 61 LVGACCRVSCLLWLYLFAMFLLILLGFCFTIFAFAVTNRGAGEVISDRGYKEYHVADYSN 120
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAK-MGRVDSNGIPEPADVFYLRKLTSVQSGCCKPP 179
WL++RV + W +I SC+ S VC+ R S + + FY L ++QSGCCKP
Sbjct: 121 WLQKRVNNAKNWERIRSCLMYSDVCSTYRTRYASINVED----FYKSNLNALQSGCCKPS 176
Query: 180 TDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSV 239
DC + Y N T W G N DC+ W N G LCY C++CKAG+L ++K SW+KV+
Sbjct: 177 NDCNFTYVNPTTWTKTPGPYK-NEDCNVWDNKPGTLCYDCEACKAGLLDNIKNSWKKVAK 235
Query: 240 INIVVMIILVIVYIIAYAAYRNNKR 264
+NIV +I L+IVY + A+RNN++
Sbjct: 236 VNIVFLIFLIIVYSVGCCAFRNNRK 260
>UniRef100_Q9FIQ5 Senescence-associated protein 5-like protein [Arabidopsis thaliana]
Length = 269
Score = 243 bits (619), Expect = 5e-63
Identities = 121/266 (45%), Positives = 172/266 (64%), Gaps = 7/266 (2%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M SN++IG +NF+T LLSIP++G GIWL+ N+ C+K LQWP+II+GV I++V LAG
Sbjct: 1 MPLSNNVIGCINFITVLLSIPVIGAGIWLAIGTVNS-CVKLLQWPVIILGVLILLVGLAG 59
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F G +R T+L+ +YL+ M ++I +L + F Y+VT +GSG +R Y++Y LQD+SG
Sbjct: 60 FIGGFWRITWLLVVYLIAMLILIVLLGCLVGFIYMVTIRGSGHPEPSRAYLEYSLQDFSG 119
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL RV W +I +C+ + +C ++ N A F+ L +QSGCCKPPT
Sbjct: 120 WLRRRVQRSYKWERIRTCLSTTTICPEL-----NQRYTLAQDFFNAHLDPIQSGCCKPPT 174
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
CG+ + N T W + MSA+ DC WSNDQ LCY CDSCKAG+LA++K W K +
Sbjct: 175 KCGFTFVNPTYW-ISPIDMSADMDCLNWSNDQNTLCYTCDSCKAGLLANIKVDWLKADIF 233
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMD 266
++ +I L+IVYII A+RN + D
Sbjct: 234 LLLALIGLIIVYIIGCCAFRNAETED 259
>UniRef100_Q8VYM8 Putative senescence-associated protein 5 [Arabidopsis thaliana]
Length = 269
Score = 242 bits (618), Expect = 7e-63
Identities = 121/266 (45%), Positives = 172/266 (64%), Gaps = 7/266 (2%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M SN++IG +NF+T LLSIP++G GIWL+ N+ C+K LQWP+II+GV I++V LAG
Sbjct: 1 MPLSNNVIGCINFITVLLSIPVIGAGIWLAIGTVNS-CVKLLQWPVIILGVLILLVGLAG 59
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F G +R T+L+ +YL+ M ++I +L + F Y+VT +GSG +R Y++Y LQD+SG
Sbjct: 60 FIGGFWRITWLLVVYLIAMLILIVLLGCLVGFIYMVTIRGSGHPEPSRAYLEYSLQDFSG 119
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL RV W +I +C+ + +C ++ N A F+ L +QSGCCKPPT
Sbjct: 120 WLRRRVQRSYKWERIRTCLSTTTICPEL-----NQRYTLAQDFFNAHLDPIQSGCCKPPT 174
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
CG+ + N T W + MSA+ DC WSNDQ LCY CDSCKAG+LA++K W K +
Sbjct: 175 KCGFTFVNPTYW-VSPIDMSADMDCLNWSNDQNTLCYTCDSCKAGLLANIKVDWLKADIF 233
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMD 266
++ +I L+IVYII A+RN + D
Sbjct: 234 LLLALIGLIIVYIIGCCAFRNAETED 259
>UniRef100_Q9ZUN5 Putative senescence-associated protein 5 [Arabidopsis thaliana]
Length = 270
Score = 241 bits (616), Expect = 1e-62
Identities = 118/267 (44%), Positives = 173/267 (64%), Gaps = 8/267 (2%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M +N+L +LN L L SIPI GIWL+S+ +N +C+ L+WP++++GV I+VVS G
Sbjct: 1 MALANNLTAILNLLALLCSIPITASGIWLASKPDN-ECVNLLRWPVVVLGVLILVVSATG 59
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F GA L+ +YL M ++I +L+ +IFA+VVT RV RGY +Y L+ +S
Sbjct: 60 FIGAYKYKETLLAVYLCCMAILIGLLLVVLIFAFVVTRPDGSYRVPGRGYKEYRLEGFSN 119
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFY-LRKLTSVQSGCCKPP 179
WL+E V WG++ +C+ D+ VC K+ + AD F+ K+T +QSGCCKPP
Sbjct: 120 WLKENVVDSKNWGRLRACLADTNVCPKLNQEFIT-----ADQFFSSSKITPLQSGCCKPP 174
Query: 180 TDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSV 239
T CGY + N T+W L M+A+ DC WSNDQ LCY C+SCKAG+L +L+K WRK ++
Sbjct: 175 TACGYNFVNPTLW-LNPTNMAADADCYLWSNDQSQLCYNCNSCKAGLLGNLRKEWRKANL 233
Query: 240 INIVVMIILVIVYIIAYAAYRNNKRMD 266
I I+ +++L+ VY+IA +A+RN + D
Sbjct: 234 ILIITVVVLIWVYVIACSAFRNAQTED 260
>UniRef100_Q75IM5 Putative senescence-associated protein [Oryza sativa]
Length = 286
Score = 232 bits (591), Expect = 1e-59
Identities = 112/280 (40%), Positives = 166/280 (59%), Gaps = 16/280 (5%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN +IG LN T L SIP++G G+W++ + T C LQ PL++IG +++VSLAGF
Sbjct: 5 RFSNVMIGYLNLATLLASIPVIGAGLWMAKGSTAT-CSSMLQTPLLVIGFVVLLVSLAGF 63
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC+ + + LYL+ M L+IA L+G F + VT G G +V R Y +Y+ DYS W
Sbjct: 64 VGACFHVAWALWLYLLAMMLLIAFLLGLTAFGFAVTAGGGGTQVPGRPYREYHTSDYSSW 123
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L++ + YW +CV SK C K+ P D + LT +QSGCCKPPT
Sbjct: 124 LQKHIQDAKYWRPALACVVGSKACPKIANW------SPMD-YLQHDLTPIQSGCCKPPTA 176
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
C Y + G + + + DC +W+N G LCY C+SC+AGV+ +++ W K+SV+N
Sbjct: 177 CAY--------SGGVAVGAQDEDCFRWNNAAGILCYGCESCRAGVMEKVREDWHKISVLN 228
Query: 242 IVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHP 281
++V+++L+ + A+RN +R ++ PYG RM K HP
Sbjct: 229 VMVLVVLICICACGCCAFRNARRSVSEYPYGVNRMHKIHP 268
>UniRef100_Q9C7C1 Senescence-assocated protein, putative; 28418-29806 [Arabidopsis
thaliana]
Length = 282
Score = 231 bits (589), Expect = 2e-59
Identities = 120/281 (42%), Positives = 166/281 (58%), Gaps = 20/281 (7%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN +IGVLN LT L SIPI+G ++ + ++T C FLQ PL++IG I++VSLAGF
Sbjct: 3 RFSNTVIGVLNLLTLLASIPIIGTALYKAR--SSTTCENFLQTPLLVIGFIILIVSLAGF 60
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC+ + + +YLVVM +IA L+G +F VVT +G G V R Y +Y L DY W
Sbjct: 61 IGACFNVAWALWVYLVVMIFLIATLMGLTLFGLVVTSQGGGVEVPGRIYKEYRLGDYHPW 120
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L ERV YW I SC+ SK C K+ + ++ R +TSVQSGCCKPPT
Sbjct: 121 LRERVRDPEYWNSIRSCILSSKTCTKIESWTTLD-------YFQRDMTSVQSGCCKPPTA 173
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
C Y +G++ DC +W+N LCY CD+CKAGVL ++ WRK+SV+N
Sbjct: 174 CTY----------EAGVVDGGGDCFRWNNGVEMLCYECDACKAGVLEEIRLDWRKLSVVN 223
Query: 242 IVVMIILVIVYIIAYAAYRNNKRMDND-EPYGEARMTKSHP 281
I+V+++L+ VY A+ N + + P + RMT+ P
Sbjct: 224 ILVLVLLIAVYAAGCCAFHNTRHAAHPYHPSDDNRMTRVRP 264
>UniRef100_Q6Z5Z9 Putative senescence-associated protein [Oryza sativa]
Length = 273
Score = 219 bits (557), Expect = 8e-56
Identities = 113/268 (42%), Positives = 161/268 (59%), Gaps = 11/268 (4%)
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANN----TDCLKFLQWPLIIIGVSIMVVS 57
R SN + +N +T LL +L GI+ + T+C +FL+ P + +G +I+ VS
Sbjct: 4 RCSNAVFAAINVVTLLLGAAVLAAGIYYGAPHRGGGGVTECERFLRAPALALGGAIVAVS 63
Query: 58 LAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQD 117
LAG AGAC R T L+ YL++ L+I F +FA VVT+ G+GR V RG+ +Y+L D
Sbjct: 64 LAGLAGACCRATPLLWAYLLLTGLLILAAACFGVFALVVTNAGAGRAVSGRGFREYHLGD 123
Query: 118 YSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCK 177
YS WL V +W +I SC+ D+ VC + SN + D F L+ +QSGCCK
Sbjct: 124 YSTWLRRSVEDGGHWARIRSCLVDTGVCRSL---KSN---QTLDEFVNSNLSPLQSGCCK 177
Query: 178 PPTDCGYIYQNETVW-NLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRK 236
PPT C + YQNET W + ++PDC+ WSNDQ LCY C SCKAGVL +L+ SW+K
Sbjct: 178 PPTACNFTYQNETYWIKPPTPSNYSDPDCNSWSNDQSELCYGCQSCKAGVLGNLRSSWKK 237
Query: 237 VSVINIVVMIILVIVYIIAYAAYRNNKR 264
++ +N + +L++VY + A RNN+R
Sbjct: 238 IAFVNAAFVALLLVVYSLGCCALRNNRR 265
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.140 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 499,596,021
Number of Sequences: 2790947
Number of extensions: 20818054
Number of successful extensions: 86193
Number of sequences better than 10.0: 254
Number of HSP's better than 10.0 without gapping: 104
Number of HSP's successfully gapped in prelim test: 150
Number of HSP's that attempted gapping in prelim test: 85942
Number of HSP's gapped (non-prelim): 270
length of query: 286
length of database: 848,049,833
effective HSP length: 126
effective length of query: 160
effective length of database: 496,390,511
effective search space: 79422481760
effective search space used: 79422481760
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 74 (33.1 bits)
Lotus: description of TM0246c.3


