

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.8
(334 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
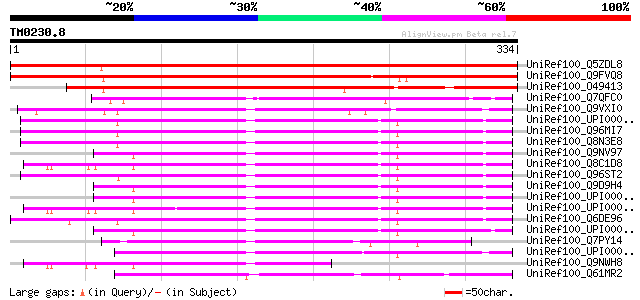
UniRef100_Q5ZDL8 Hypothetical protein P0416D03.37-1 [Oryza sativa] 510 e-143
UniRef100_Q9FVQ8 Hypothetical protein F3C3.8 [Arabidopsis thaliana] 498 e-139
UniRef100_O49413 Hypothetical protein AT4g19000 [Arabidopsis tha... 274 3e-72
UniRef100_Q7QFC0 ENSANGP00000010073 [Anopheles gambiae str. PEST] 150 5e-35
UniRef100_Q9VXI0 CG9915-PA [Drosophila melanogaster] 147 4e-34
UniRef100_UPI00003695DE UPI00003695DE UniRef100 entry 147 5e-34
UniRef100_Q96MI7 Hypothetical protein FLJ32319 [Homo sapiens] 147 5e-34
UniRef100_Q8N3E8 Hypothetical protein DKFZp761G0123 [Homo sapiens] 147 5e-34
UniRef100_Q9NV97 Hypothetical protein FLJ10855 [Homo sapiens] 145 2e-33
UniRef100_Q8C1D8 Mus musculus 12 days embryo head cDNA, RIKEN fu... 145 2e-33
UniRef100_Q96ST2 Hypothetical protein FLJ14655 [Homo sapiens] 145 2e-33
UniRef100_Q9D9H4 Mus musculus adult male testis cDNA, RIKEN full... 143 6e-33
UniRef100_UPI00003AEC5E UPI00003AEC5E UniRef100 entry 142 2e-32
UniRef100_UPI000021D736 UPI000021D736 UniRef100 entry 139 8e-32
UniRef100_Q6DE96 FLJ10006 protein [Xenopus laevis] 135 2e-30
UniRef100_UPI00003AEC5F UPI00003AEC5F UniRef100 entry 134 3e-30
UniRef100_Q7PY14 ENSANGP00000012213 [Anopheles gambiae str. PEST] 131 3e-29
UniRef100_UPI00002BB61E UPI00002BB61E UniRef100 entry 130 5e-29
UniRef100_Q9NWH8 Hypothetical protein FLJ10006 [Homo sapiens] 117 4e-25
UniRef100_Q61MR2 Hypothetical protein CBG08373 [Caenorhabditis b... 113 8e-24
>UniRef100_Q5ZDL8 Hypothetical protein P0416D03.37-1 [Oryza sativa]
Length = 533
Score = 510 bits (1313), Expect = e-143
Identities = 256/337 (75%), Positives = 287/337 (84%), Gaps = 3/337 (0%)
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGF-YGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKK 59
DDQEGVR LDDDNFIDDTGV+P YG+ N+ SP PQAEEGEEDDEI LFK GKKK
Sbjct: 196 DDQEGVRTLDDDNFIDDTGVDPADRYGSDNDGHSPRHYPQAEEGEEDDEIERLFKGGKKK 255
Query: 60 --NERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLE 117
N+R A+I L+VE +AE EV AEEDA LNRQ KPAINKL KLPLL +VLSKK LQ E
Sbjct: 256 KKNDRPRADIGLIVEQFIAEFEVAAEEDANLNRQSKPAINKLMKLPLLIDVLSKKNLQQE 315
Query: 118 FLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVI 177
FLDHGVLTLLKNWLEPLPDGSLPN+NIRTA+LK+L DFPIDLEQ DRREQLK+SGLGKVI
Sbjct: 316 FLDHGVLTLLKNWLEPLPDGSLPNMNIRTAVLKLLTDFPIDLEQYDRREQLKKSGLGKVI 375
Query: 178 MFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKA 237
MFLSKSDEE NRKL KELVDKWSRPIFNKSTRFEDMR +DERAP+RRP +KKP++ +
Sbjct: 376 MFLSKSDEETTSNRKLAKELVDKWSRPIFNKSTRFEDMRRYDDERAPYRRPQMKKPSSSS 435
Query: 238 PGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASH 297
GM+SRD DLD D Q +SGQ +RQHASRPEA+P+DFVIRPQSK+DPE++RARAKQ
Sbjct: 436 SGMESRDDDLDADFSQRKSGQGGARQHASRPEASPLDFVIRPQSKIDPEQIRARAKQVVQ 495
Query: 298 DQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
DQ+R+KMNKKLQQL+APKK+ LQA+KLSVEGRGMIKY
Sbjct: 496 DQRRLKMNKKLQQLKAPKKKNLQASKLSVEGRGMIKY 532
>UniRef100_Q9FVQ8 Hypothetical protein F3C3.8 [Arabidopsis thaliana]
Length = 404
Score = 498 bits (1281), Expect = e-139
Identities = 255/343 (74%), Positives = 295/343 (85%), Gaps = 10/343 (2%)
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGF-YGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKK 59
+D+EGVR +DDDNFIDDTG++P YG SP PQAEEGE++DE+N+LFKMGKKK
Sbjct: 62 NDEEGVRTMDDDNFIDDTGLDPSERYGGDAGDRSPTHYPQAEEGEDEDEVNNLFKMGKKK 121
Query: 60 N--ERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLE 117
ER+PAEIALLVENV+AELEVTAEEDAELNRQGKPAINKLKKL LLT+VL KKQLQ E
Sbjct: 122 KKTERNPAEIALLVENVMAELEVTAEEDAELNRQGKPAINKLKKLSLLTDVLGKKQLQTE 181
Query: 118 FLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVI 177
FLDHGVLTLLKNWLEPLPDGSLPNINIR AIL++L DFPIDL+Q DRREQLK+SGLGKVI
Sbjct: 182 FLDHGVLTLLKNWLEPLPDGSLPNINIRAAILRVLTDFPIDLDQYDRREQLKKSGLGKVI 241
Query: 178 MFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKA 237
MFLSKSDEE N NR+L K+LVDKWSRPIFNKSTRFEDMRN++++R P+RRP VKKP+NKA
Sbjct: 242 MFLSKSDEETNSNRRLAKDLVDKWSRPIFNKSTRFEDMRNLDEDRVPYRRPPVKKPSNKA 301
Query: 238 PGMQSRDSDLDLDLPQPR----SGQSS--SRQHASRPEATPMDFVIRPQSKVDPEEVRAR 291
M+SRD D DL++ + + SGQSS RQ RPEATP+DF+IRPQSK+DP+E+ AR
Sbjct: 302 T-MESRDGDFDLEIRERKTGLTSGQSSRGDRQMTMRPEATPLDFLIRPQSKIDPDEIIAR 360
Query: 292 AKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
AKQ S DQ+R+KMNKKLQQL+ KK++LQATK+SVEGRGMIKY
Sbjct: 361 AKQVSQDQRRVKMNKKLQQLKGTKKKRLQATKVSVEGRGMIKY 403
>UniRef100_O49413 Hypothetical protein AT4g19000 [Arabidopsis thaliana]
Length = 406
Score = 274 bits (700), Expect = 3e-72
Identities = 155/301 (51%), Positives = 201/301 (66%), Gaps = 11/301 (3%)
Query: 38 PQAEEGEEDDEINDLFKMGKKKN--ERSPAEIALLVENVVAELEVTAEEDAELNRQGKPA 95
P ++ E+ +EI LF + KKK+ +++ EI + VE V+A LE+ E+D NR+GKPA
Sbjct: 112 PPKKKDEDAEEIKKLFSLRKKKSKCDKTSMEIGMQVEQVMANLEIAVEDDVICNREGKPA 171
Query: 96 INKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDF 155
INKL KLPLL E LSKK LQ EFLDHGVL LLKNWLEPLPDGSLPNINIR+A+L ILNDF
Sbjct: 172 INKLMKLPLLNETLSKKPLQGEFLDHGVLNLLKNWLEPLPDGSLPNINIRSAVLMILNDF 231
Query: 156 PIDLEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDM 215
IDL+Q RREQL +SGLGKVIMFLSKSDEE NR+L ++++KW R I+NKSTR+++M
Sbjct: 232 RIDLDQDSRREQLIKSGLGKVIMFLSKSDEETTPNRRLANDIINKWGRIIYNKSTRYDNM 291
Query: 216 RNIE--DERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPM 273
E DE+ K A K G ++RD + D+DL + G + R A P M
Sbjct: 292 FTQEELDEQRQILLRRQTKTAPKVSGTRARDFNTDIDLYE--LGTWTGRARAKIPTTMSM 349
Query: 274 DFVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIK 333
DF IRP SKVD + ++ + +M+ K +Q + +K +QA KLSV+GR M+K
Sbjct: 350 DFKIRPPSKVDINQ-----EEEPCSKWQMEKRHKNKQQKNIRKGGMQALKLSVDGRTMLK 404
Query: 334 Y 334
Y
Sbjct: 405 Y 405
>UniRef100_Q7QFC0 ENSANGP00000010073 [Anopheles gambiae str. PEST]
Length = 298
Score = 150 bits (379), Expect = 5e-35
Identities = 101/292 (34%), Positives = 155/292 (52%), Gaps = 24/292 (8%)
Query: 55 MGKKKNERSPA----EIALLVEN------VVAELEVTAEEDAELNRQGKPAINKLKKLPL 104
M KKK E+S +I L+ +N ++ ++ AE D +LN +GKPA K+ L
Sbjct: 14 MAKKKEEKSRRRKRNDIDLINDNDDQIAQLLQQMRQAAEHDRQLNVEGKPATKKIAMLKH 73
Query: 105 LTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDR 164
L KK LQL FL+H VL +L +W+ PLP+ +LP + IR +ILK+L+DFP ID
Sbjct: 74 AMSQLIKKDLQLAFLEHNVLNVLTDWIAPLPNKALPCLQIRESILKLLSDFP----TID- 128
Query: 165 REQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAP 224
R L++SG+GK +M+L K E VNR L+ W RPIFN ST F+ M E ++
Sbjct: 129 RSYLRQSGIGKAVMYLYKHPNETKVNRDRAGRLIADWCRPIFNLSTDFKAMTREERQQRD 188
Query: 225 FRR-PSVKKPANKAPGMQSRDSD----LDLDLPQPRSGQSSSRQHASRPEATPMDFVIRP 279
++ P +KP+ + S+ + D R G Q A P + ++++RP
Sbjct: 189 LQQMPRKRKPSPDSAASTSKSQKGLPFTEADKGTLRPGDKGWIQRARVPMPSDKEYIVRP 248
Query: 280 QSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
+SK+D + K+ + ++ +K K + Q R R +A +S+EGR M
Sbjct: 249 KSKIDVDMSSIAKKKLNRYEKHLK--KFIDQKRLNSTR--RAVDISIEGRKM 296
>UniRef100_Q9VXI0 CG9915-PA [Drosophila melanogaster]
Length = 355
Score = 147 bits (371), Expect = 4e-34
Identities = 112/349 (32%), Positives = 172/349 (49%), Gaps = 35/349 (10%)
Query: 6 VRNLDDDNFID--DTGV-EPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKNE- 61
V+ D DN D D G+ + E S P + E E + I+D M +K E
Sbjct: 17 VKITDTDNEEDEQDKGIGKEKSQDKITESSDDENTPISNEPENSNFISDFDAMLMRKKEE 76
Query: 62 ----RSPAEIAL------LVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSK 111
R +I L L++ ++ ++ +++D +LN G+PA K+ L + L K
Sbjct: 77 KRVRRRKRDIDLINDNDDLIDQLIVSMKNASDDDRQLNMIGQPATKKISMLKQVMSQLIK 136
Query: 112 KQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRS 171
K LQL FL+H +L +L +WL PLP+ SLP + IR +ILK+L+DFP + LK+S
Sbjct: 137 KHLQLAFLEHNILNVLTDWLAPLPNKSLPCLQIRESILKLLSDFP-----TIEKGLLKQS 191
Query: 172 GLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDER---APFRRP 228
G+GK +M+L K +E NR L+ +W+RPIFN S F M E + A R
Sbjct: 192 GIGKAVMYLYKHPQETKSNRDRAGRLISEWARPIFNVSCNFSAMSKEERQERDLAQMSRH 251
Query: 229 SVKKP------ANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSK 282
K P ++KAP + S+ D L R G A P + D+VIRP+S
Sbjct: 252 RHKSPDTEPSSSSKAPTLNQTLSNEDKML---RPGDPGWVARARVPMPSNKDYVIRPKST 308
Query: 283 VDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
V+ E + ++ + ++ MK ++L K +A ++S+EGR M
Sbjct: 309 VEGEVSASTKRKPNRYEKHMKRFLDSKRL----KESRRAVEISIEGRKM 353
>UniRef100_UPI00003695DE UPI00003695DE UniRef100 entry
Length = 350
Score = 147 bits (370), Expect = 5e-34
Identities = 106/354 (29%), Positives = 165/354 (45%), Gaps = 37/354 (10%)
Query: 8 NLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGE-EDDEINDLFKMGKKKNERSPAE 66
NL D F + E + +N+ E + + E ED + +D K GK + S E
Sbjct: 2 NLIADIFGESGDEEEEEFTGFNQEDLEEEKGETQVKEAEDSDSDDNIKRGKHMDFLSDFE 61
Query: 67 IAL-------------------------LVENVVAELEVTAEEDAELNRQGKPAINKLKK 101
+ L +V ++ ++ AEED +LN Q KPA+ KL
Sbjct: 62 MMLQRKKSMSGKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKKLTL 121
Query: 102 LPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQ 161
LP + L K+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P
Sbjct: 122 LPTVVMHLKKQDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP----- 176
Query: 162 IDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDE 221
+E LK SG+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E
Sbjct: 177 SVSQETLKHSGIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTREERE 236
Query: 222 RAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVI 277
+ + ++ N G R DL+ L R G A P + D+V+
Sbjct: 237 QRDLEQMPQRRRMNSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVV 295
Query: 278 RPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
RP+ V+ E R +A + K +K +R K R A K+S+EG M
Sbjct: 296 RPKWNVEMESSRFQATSKKGISRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 348
>UniRef100_Q96MI7 Hypothetical protein FLJ32319 [Homo sapiens]
Length = 494
Score = 147 bits (370), Expect = 5e-34
Identities = 106/354 (29%), Positives = 165/354 (45%), Gaps = 37/354 (10%)
Query: 8 NLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGE-EDDEINDLFKMGKKKNERSPAE 66
NL D F + E + +N+ E + + E ED + +D K GK + S E
Sbjct: 146 NLIADIFGESGDEEEEEFTGFNQEDLEEEKGETQVKEAEDSDSDDNIKRGKHMDFLSDFE 205
Query: 67 IAL-------------------------LVENVVAELEVTAEEDAELNRQGKPAINKLKK 101
+ L +V ++ ++ AEED +LN Q KPA+ KL
Sbjct: 206 MMLQRKKSMSGKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKKLTL 265
Query: 102 LPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQ 161
LP + L K+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P
Sbjct: 266 LPAVVMHLKKQDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP----- 320
Query: 162 IDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDE 221
+E LK SG+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E
Sbjct: 321 SVSQETLKHSGIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTREERE 380
Query: 222 RAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVI 277
+ + ++ N G R DL+ L R G A P + D+V+
Sbjct: 381 QRDLEQMPQRRRMNSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVV 439
Query: 278 RPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
RP+ V+ E R +A + K +K +R K R A K+S+EG M
Sbjct: 440 RPKWNVEMESSRFQATSKKGISRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 492
>UniRef100_Q8N3E8 Hypothetical protein DKFZp761G0123 [Homo sapiens]
Length = 819
Score = 147 bits (370), Expect = 5e-34
Identities = 106/354 (29%), Positives = 165/354 (45%), Gaps = 37/354 (10%)
Query: 8 NLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGE-EDDEINDLFKMGKKKNERSPAE 66
NL D F + E + +N+ E + + E ED + +D K GK + S E
Sbjct: 471 NLIADIFGESGDEEEEEFTGFNQEDLEEEKGETQVKEAEDSDSDDNIKRGKHMDFLSDFE 530
Query: 67 IAL-------------------------LVENVVAELEVTAEEDAELNRQGKPAINKLKK 101
+ L +V ++ ++ AEED +LN Q KPA+ KL
Sbjct: 531 MMLQRKKSMSGKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKKLTL 590
Query: 102 LPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQ 161
LP + L K+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P
Sbjct: 591 LPAVVMHLKKQDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP----- 645
Query: 162 IDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDE 221
+E LK SG+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E
Sbjct: 646 SVSQETLKHSGIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTREERE 705
Query: 222 RAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVI 277
+ + ++ N G R DL+ L R G A P + D+V+
Sbjct: 706 QRDLEQMPQRRRMNSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVV 764
Query: 278 RPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
RP+ V+ E R +A + K +K +R K R A K+S+EG M
Sbjct: 765 RPKWNVEMESSRFQATSKKGISRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 817
>UniRef100_Q9NV97 Hypothetical protein FLJ10855 [Homo sapiens]
Length = 297
Score = 145 bits (366), Expect = 2e-33
Identities = 94/284 (33%), Positives = 144/284 (50%), Gaps = 15/284 (5%)
Query: 56 GKKKNERSPAEIALLVENVVAELEV----TAEEDAELNRQGKPAINKLKKLPLLTEVLSK 111
GK++ R ++VV+ + V AEED +LN Q KPA+ KL LP + L K
Sbjct: 19 GKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKKLTLLPAVVMHLKK 78
Query: 112 KQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRS 171
+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P +E LK S
Sbjct: 79 QDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP-----SVSQETLKHS 133
Query: 172 GLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVK 231
G+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E+ + +
Sbjct: 134 GIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTREEREQRDLEQMPQR 193
Query: 232 KPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVIRPQSKVDPEE 287
+ N G R DL+ L R G A P + D+V+RP+ V+ E
Sbjct: 194 RRMNSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVVRPKWNVEMES 252
Query: 288 VRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
R +A + K +K +R K R A K+S+EG M
Sbjct: 253 SRFQATSKKGISRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 295
>UniRef100_Q8C1D8 Mus musculus 12 days embryo head cDNA, RIKEN full-length enriched
library, clone:3000008H23 product:hypothetical Acyl-CoA
dehydrogenase/Glutamic acid-rich region containing
protein, full insert sequence [Mus musculus]
Length = 766
Score = 145 bits (365), Expect = 2e-33
Identities = 108/357 (30%), Positives = 169/357 (47%), Gaps = 42/357 (11%)
Query: 10 DDDNFIDDTGVEPG------FYG----NYNEPSSPGEAPQAEEGEEDDEIN--------D 51
+++N I D E G F G + E + + +AE+ + DD I
Sbjct: 415 EEENLIADIFGESGDEEEEEFTGFNQEDLEEEKNETQLKEAEDSDSDDNIKRGKHMDFLS 474
Query: 52 LFKM---------GKKKNERSPAEIALLVENVVAELEV----TAEEDAELNRQGKPAINK 98
F+M GK++ R ++VV+ + V AEED +LN Q KPA+ K
Sbjct: 475 DFEMMLQRKKSMCGKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKK 534
Query: 99 LKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPID 158
L LP + L K+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P
Sbjct: 535 LTLLPTVVMHLKKQDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP-- 592
Query: 159 LEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNI 218
+E LK SG+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M
Sbjct: 593 ---SVSQETLKHSGIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTRE 649
Query: 219 EDERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMD 274
E E+ + ++ + G R DL+ L R G A P + D
Sbjct: 650 EREQRDLEQMPQRRRLSSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKD 708
Query: 275 FVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
+V+RP+ V+ E R +A + K +K +R K R A K+S+EG M
Sbjct: 709 YVVRPKWNVEMESSRFQASSKKGISRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 764
>UniRef100_Q96ST2 Hypothetical protein FLJ14655 [Homo sapiens]
Length = 819
Score = 145 bits (365), Expect = 2e-33
Identities = 106/354 (29%), Positives = 164/354 (45%), Gaps = 37/354 (10%)
Query: 8 NLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGE-EDDEINDLFKMGKKKNERSPAE 66
NL D F + E + +N+ E + + E ED + +D K GK + S E
Sbjct: 471 NLIADIFGESGDEEEEEFTGFNQEDLEEEKGETQVKEAEDSDSDDNIKRGKHMDFLSDFE 530
Query: 67 IALL-------------------------VENVVAELEVTAEEDAELNRQGKPAINKLKK 101
+ L V ++ ++ AEED +LN Q KPA+ KL
Sbjct: 531 MMLQRKKSMSGKRRRNRDGGTFISDADDDVSAMIVKMNEAAEEDRQLNNQKKPALKKLTL 590
Query: 102 LPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQ 161
LP + L K+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P
Sbjct: 591 LPAVVMHLKKQDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP----- 645
Query: 162 IDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDE 221
+E LK SG+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E
Sbjct: 646 SVSQETLKHSGIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTREERE 705
Query: 222 RAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVI 277
+ + ++ N G R DL+ L R G A P + D+V+
Sbjct: 706 QRDLEQMPQRRRMNSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVV 764
Query: 278 RPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
RP+ V+ E R +A + K +K +R K R A K+S+EG M
Sbjct: 765 RPKWNVEMESSRFQATSKKGIGRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 817
>UniRef100_Q9D9H4 Mus musculus adult male testis cDNA, RIKEN full-length enriched
library, clone:1700069O15 product:hypothetical Acyl-CoA
dehydrogenase/Glutamic acid-rich region containing
protein, full insert sequence [Mus musculus]
Length = 281
Score = 143 bits (361), Expect = 6e-33
Identities = 93/284 (32%), Positives = 144/284 (49%), Gaps = 15/284 (5%)
Query: 56 GKKKNERSPAEIALLVENVVAELEV----TAEEDAELNRQGKPAINKLKKLPLLTEVLSK 111
GK++ R ++VV+ + V AEED +LN Q KPA+ KL LP + L K
Sbjct: 3 GKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKKLTLLPTVVMHLKK 62
Query: 112 KQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRS 171
+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P +E LK S
Sbjct: 63 QDLKETFIDSGVMSAIKEWLSPLPDRSLPAMKIREELLKILQELP-----SVSQETLKHS 117
Query: 172 GLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVK 231
G+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E+ + +
Sbjct: 118 GIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTREEREQRDLEQMPQR 177
Query: 232 KPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVIRPQSKVDPEE 287
+ + G R DL+ L R G A P + D+V+RP+ V+ E
Sbjct: 178 RRLSSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVVRPKWNVEMES 236
Query: 288 VRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
R +A + K +K +R K R A K+S+EG M
Sbjct: 237 SRFQASSKKGISRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 279
>UniRef100_UPI00003AEC5E UPI00003AEC5E UniRef100 entry
Length = 450
Score = 142 bits (357), Expect = 2e-32
Identities = 92/284 (32%), Positives = 144/284 (50%), Gaps = 15/284 (5%)
Query: 56 GKKKNERSPAEIALLVENVVAELEV----TAEEDAELNRQGKPAINKLKKLPLLTEVLSK 111
GK++ R ++VV+ + V AEED +LN Q KPA+ KL LP + L K
Sbjct: 172 GKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNTQKKPALKKLTLLPTVVMHLKK 231
Query: 112 KQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRS 171
+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P +E LK S
Sbjct: 232 QDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP-----SVSQETLKHS 286
Query: 172 GLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVK 231
G+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E+ + +
Sbjct: 287 GIGRAVMYLYKHPKESRPNKDMAGKLINEWSRPIFGLTSNYKGMTREEREQRDLEQMPQR 346
Query: 232 KPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVIRPQSKVDPEE 287
+ + + G R DL+ L R G A P + D+V+RP+ V+ E
Sbjct: 347 RRLSSSGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVVRPKWNVEMES 405
Query: 288 VRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
R + + K +K +R K R A K+S+EG M
Sbjct: 406 SRFQGTSKKGVSRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 448
>UniRef100_UPI000021D736 UPI000021D736 UniRef100 entry
Length = 715
Score = 139 bits (351), Expect = 8e-32
Identities = 108/357 (30%), Positives = 168/357 (46%), Gaps = 43/357 (12%)
Query: 10 DDDNFIDDTGVEPG------FYG----NYNEPSSPGEAPQAEEGEEDDEIN--------D 51
+++N I D E G F G + E + + +AE+ + DD I
Sbjct: 365 EEENLIADIFGESGDEEEEEFTGFNQEDLEEEKNETQLKEAEDSDSDDNIKRGKHMDFLS 424
Query: 52 LFKM---------GKKKNERSPAEIALLVENVVAELEV----TAEEDAELNRQGKPAINK 98
F+M GK++ R ++VV+ + V AEED +LN Q KPA+ K
Sbjct: 425 DFEMMLQRKKSMCGKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKK 484
Query: 99 LKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPID 158
L LP + L K L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P
Sbjct: 485 LTLLPTVVMHL-KNDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP-- 541
Query: 159 LEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNI 218
+E LK SG+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M
Sbjct: 542 ---SVSQETLKHSGIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNYKGMTRE 598
Query: 219 EDERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMD 274
E E+ + ++ + G R DL+ L R G A P + D
Sbjct: 599 EREQRDLEQMPQRRRMSSTGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKD 657
Query: 275 FVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
+V+RP+ V+ E R +A + K +K +R K R A K+S+EG M
Sbjct: 658 YVVRPKWNVEMESSRFQATSKKGISRLDKQMRKFTDIR-KKSRSAHAVKISIEGNKM 713
>UniRef100_Q6DE96 FLJ10006 protein [Xenopus laevis]
Length = 836
Score = 135 bits (340), Expect = 2e-30
Identities = 102/365 (27%), Positives = 166/365 (44%), Gaps = 41/365 (11%)
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAP-----QAEEGEEDDEINDLFKM 55
D + NL D F + E + +N+ GE + + D + +D K
Sbjct: 477 DSENEEENLIADIFGESGDEEEEEFTGFNQEDLEGEKTVTSVNRQQAAAADSDSDDDMKS 536
Query: 56 GKKKNERSPAEIAL-------------------------LVENVVAELEVTAEEDAELNR 90
GK + +S ++ L +V ++ ++ AEED LN
Sbjct: 537 GKMGDYKSDFDLMLERKKSMSGKQRRNRDGGTFISDADDVVNAMIMKMTEAAEEDRNLNS 596
Query: 91 QGKPAINKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILK 150
KPA+ KL LP + L K+ L+ F+D GV++ +K WL PLPD SLP + IR +LK
Sbjct: 597 SKKPALKKLTLLPTVVMHLKKQDLKETFIDSGVMSAIKEWLTPLPDRSLPALKIREELLK 656
Query: 151 ILNDFPIDLEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKST 210
IL + P +E LK SG+G+ IM+L K +E N+ + +L+++WSRPIF ++
Sbjct: 657 ILQELP-----SVSQETLKHSGIGRAIMYLYKHPKESRPNKDIAGKLINEWSRPIFGLTS 711
Query: 211 RFEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHAS 266
++ M E E+ + ++ + + G R DLD L R G+ A
Sbjct: 712 NYKGMTREEREQRDIEQMPQRRRMSSSGGQTPR-RDLDKVLTGEEKALRPGEPGFCARAR 770
Query: 267 RPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSV 326
P + D+V+RP+ V+ E + + + K +K ++ K R A K+S+
Sbjct: 771 VPLPSNKDYVVRPKWNVEMETSQYQGSAKKGVSRIDKQMRKFVDIK-KKNRFGHAVKISI 829
Query: 327 EGRGM 331
EG M
Sbjct: 830 EGNKM 834
>UniRef100_UPI00003AEC5F UPI00003AEC5F UniRef100 entry
Length = 295
Score = 134 bits (338), Expect = 3e-30
Identities = 90/284 (31%), Positives = 142/284 (49%), Gaps = 16/284 (5%)
Query: 56 GKKKNERSPAEIALLVENVVAELEV----TAEEDAELNRQGKPAINKLKKLPLLTEVLSK 111
GK++ R ++VV+ + V AEED +LN Q KPA+ KL LP + L K
Sbjct: 18 GKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNTQKKPALKKLTLLPTVVMHLKK 77
Query: 112 KQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRS 171
+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P +E LK S
Sbjct: 78 QDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP-----SVSQETLKHS 132
Query: 172 GLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVK 231
G+G+ +M+L K +E N+ + +L+++WSRPIF ++ ++ M E E+ + +
Sbjct: 133 GIGRAVMYLYKHPKESRPNKDMAGKLINEWSRPIFGLTSNYKGMTREEREQRDLEQMPQR 192
Query: 232 KPANKAPGMQSRDSDLDLDLPQP----RSGQSSSRQHASRPEATPMDFVIRPQSKVDPEE 287
+ + + G R DL+ L R G A P + D+V+RP+ V+ E
Sbjct: 193 RRLSSSGGQTPR-RDLEKVLTGEEKALRPGDPGFCARARVPMPSNKDYVVRPKWNVEMES 251
Query: 288 VRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
R + + + +Q R K +A K S+EG M
Sbjct: 252 SRPGTIKRGISRLEKHKRRFAEQKRLSKVH--RAIKFSIEGNRM 293
>UniRef100_Q7PY14 ENSANGP00000012213 [Anopheles gambiae str. PEST]
Length = 339
Score = 131 bits (329), Expect = 3e-29
Identities = 84/255 (32%), Positives = 130/255 (50%), Gaps = 24/255 (9%)
Query: 61 ERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEFLD 120
+R+ +I +L+E ++ A++D +LNR KPA K+ L L KK LQL L+
Sbjct: 73 KRNDDQILVLLET----MQQAADQDRQLNRANKPATKKIALLKHFLSQLIKKDLQLPLLE 128
Query: 121 HGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVIMFL 180
H VL +L +W+ PLP +LP + IR ILK+L +FP + LK+SG+GK +M+L
Sbjct: 129 HNVLNVLADWISPLPSKALPCLQIREGILKMLGEFP-----TIEKSYLKQSGIGKAVMYL 183
Query: 181 SKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANK---- 236
K +E N NRK EL+ W R I+N + + M E ++ + + PANK
Sbjct: 184 YKHPDETNANRKRAGELIHGWFRTIYNVNIELKAMSREERQQLEMK----QMPANKRKTM 239
Query: 237 ----APGMQSRDSDLDLDLPQPRSGQSSSRQHASR---PEATPMDFVIRPQSKVDPEEVR 289
+ G D L LD + + R R P + +V+RP+SK+D +
Sbjct: 240 DSLPSAGCDQLDEALWLDKGERDTRHPGDRDWVGRARVPMPSDKAYVVRPKSKIDVDICT 299
Query: 290 ARAKQASHDQQRMKM 304
A KQ + ++ +KM
Sbjct: 300 ASRKQLNRYEKHLKM 314
>UniRef100_UPI00002BB61E UPI00002BB61E UniRef100 entry
Length = 613
Score = 130 bits (327), Expect = 5e-29
Identities = 85/267 (31%), Positives = 141/267 (51%), Gaps = 15/267 (5%)
Query: 70 LVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKN 129
+V +++++ AEED LN KPA+ KL LP + L K+ L+ F+D GV++ +K
Sbjct: 355 VVSAMISKMNEAAEEDRVLNSHKKPALKKLTLLPQVVMHLKKQDLKESFIDSGVMSAIKE 414
Query: 130 WLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRREQLKRSGLGKVIMFLSKSDEEINV 189
W+ PLPD SLP + IR +L+IL + P +E LK SG+G+ +M+L K +E
Sbjct: 415 WISPLPDKSLPALRIREELLRILMELP-----SVSQETLKHSGIGRAVMYLYKHPKESRS 469
Query: 190 NRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSDLDL 249
N+ L+ +L+++WSRPIF S+ ++ M E ++ + +K + G Q+ DL+
Sbjct: 470 NKDLSLKLINEWSRPIFGLSSNYKGMTREERQQRDLDQQMPQK-RRLSSGGQTPRRDLEK 528
Query: 250 DLPQP----RSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQA-SHDQQRMKM 304
L R G A P + D+V+RP+ V+ + R K+ S ++M+
Sbjct: 529 QLTGEEKALRPGDPGFCARARVPMPSNKDYVVRPKWNVEMDSSRGPMKKGLSRVDKQMRR 588
Query: 305 NKKLQQLRAPKKRQLQATKLSVEGRGM 331
+++L + A K+SVEG M
Sbjct: 589 FADIRRL----TKSGHAVKISVEGNRM 611
>UniRef100_Q9NWH8 Hypothetical protein FLJ10006 [Homo sapiens]
Length = 489
Score = 117 bits (293), Expect = 4e-25
Identities = 77/234 (32%), Positives = 119/234 (49%), Gaps = 36/234 (15%)
Query: 10 DDDNFIDDTGVEPG------FYG----NYNEPSSPGEAPQAEEGEEDDEI---------N 50
+++N I D E G F G + E + +AE+ + DD I +
Sbjct: 261 EEENLIADIFGESGDEEEEEFTGFNQEDLEEEKGETQVKEAEDSDSDDNIKRGKHMDFLS 320
Query: 51 DLFKM--------GKKKNERSPAEIALLVENVVAELEV----TAEEDAELNRQGKPAINK 98
D M GK++ R ++VV+ + V AEED +LN Q KPA+ K
Sbjct: 321 DFGMMLQRKKSMSGKRRRNRDGGTFISDADDVVSAMIVKMNEAAEEDRQLNNQKKPALKK 380
Query: 99 LKKLPLLTEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPID 158
L LP + L K+ L+ F+D GV++ +K WL PLPD SLP + IR +LKIL + P
Sbjct: 381 LTLLPAVVMHLKKQDLKETFIDSGVMSAIKEWLSPLPDRSLPALKIREELLKILQELP-- 438
Query: 159 LEQIDRREQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRF 212
+E LK SG+G+ +M+L K +E N+ + +L+++WSRPIF ++ +
Sbjct: 439 ---SVSQETLKHSGIGRAVMYLYKHPKESRSNKDMAGKLINEWSRPIFGLTSNY 489
>UniRef100_Q61MR2 Hypothetical protein CBG08373 [Caenorhabditis briggsae]
Length = 500
Score = 113 bits (282), Expect = 8e-24
Identities = 76/283 (26%), Positives = 134/283 (46%), Gaps = 34/283 (12%)
Query: 70 LVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEVLSKKQLQLEFLDHGVLTLLKN 129
+V +V +++ A+ D N + KPA K+K LP + ++ + + +++G ++ L
Sbjct: 229 MVAKLVEKMKHAAKSDRNANVERKPAFQKIKMLPEVKAIMLRAGIVEVLIENGFMSALSE 288
Query: 130 WLEPLPDGSLPNINIRTAILKILND---FPIDLEQIDRREQLKRSGLGKVIMFLSKSDEE 186
WL PLPD LP ++IR +LK+L++ + +D R LK+SGLGK +M L K E
Sbjct: 289 WLAPLPDKCLPALDIRITVLKLLHNPRFWKLD------RSTLKQSGLGKAVMMLYKHPNE 342
Query: 187 INVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPFRRPSVKKPANKAPGMQSRDSD 246
N+ + +L+ +W+RPI++ T + + E E + R P + + SR+ +
Sbjct: 343 TKENKAIANKLIGEWARPIYHLDTDYSTVSRQEREERDYSR----MPEKRKKKLHSREEE 398
Query: 247 LDLDLPQPR------------------SGQSSSRQHASRPEATPMDFVIRPQSKVDPEEV 288
D D R G A P+ + D+VIRP+ +V E
Sbjct: 399 PDEDEAPKRPRIRDAEGLGPTKSLDLKPGDKGYINRARVPKPSTKDYVIRPEWRVSGE-- 456
Query: 289 RARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGM 331
+ ++ + +R + Q R K + + K+S+EGR M
Sbjct: 457 -FKGEKKASGSKRYDQTLRDFQERTRKSKANRLVKVSLEGRNM 498
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.312 0.133 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 568,982,434
Number of Sequences: 2790947
Number of extensions: 25523964
Number of successful extensions: 81116
Number of sequences better than 10.0: 314
Number of HSP's better than 10.0 without gapping: 63
Number of HSP's successfully gapped in prelim test: 254
Number of HSP's that attempted gapping in prelim test: 80749
Number of HSP's gapped (non-prelim): 494
length of query: 334
length of database: 848,049,833
effective HSP length: 128
effective length of query: 206
effective length of database: 490,808,617
effective search space: 101106575102
effective search space used: 101106575102
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 75 (33.5 bits)
Lotus: description of TM0230.8


