

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.4
(606 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
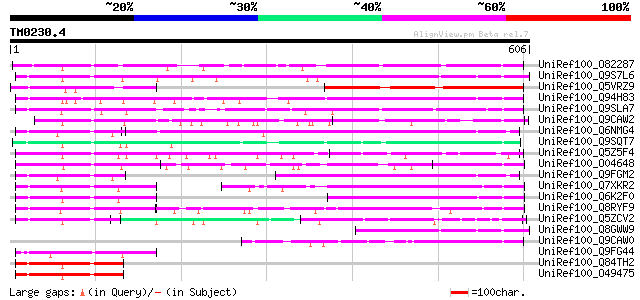
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_O82287 Putative DnaJ protein [Arabidopsis thaliana] 353 8e-96
UniRef100_Q9S7L6 Expressed protein [Arabidopsis thaliana] 250 1e-64
UniRef100_Q5VRZ9 DnaJ protein-like [Oryza sativa] 234 5e-60
UniRef100_Q94H83 Putative heat shock protein [Oryza sativa] 212 2e-53
UniRef100_Q9SLA7 At2g25560/F13B15.22 [Arabidopsis thaliana] 209 1e-52
UniRef100_Q9CAW2 Hypothetical protein T9J14.7 [Arabidopsis thali... 199 3e-49
UniRef100_Q6NMG4 At5g53150 [Arabidopsis thaliana] 182 2e-44
UniRef100_Q9SQT7 Putative DnaJ protein [Arabidopsis thaliana] 181 6e-44
UniRef100_Q5Z5F4 Putative DNAJ heat shock N-terminal domain-cont... 173 1e-41
UniRef100_O04648 A_TM021B04.9 protein [Arabidopsis thaliana] 165 3e-39
UniRef100_Q9FGM2 DnaJ protein-like [Arabidopsis thaliana] 162 3e-38
UniRef100_Q7XKR2 OSJNBa0053B21.9 protein [Oryza sativa] 161 5e-38
UniRef100_Q6K2F0 Heat shock protein-like [Oryza sativa] 161 6e-38
UniRef100_Q8RYF9 Heat shock protein-like [Oryza sativa] 150 1e-34
UniRef100_Q5ZCV2 DNAJ heat shock N-terminal domain-containing pr... 127 1e-27
UniRef100_Q8GWW9 Hypothetical protein At2g05250/F5G3.15 [Arabido... 117 7e-25
UniRef100_Q9CAW0 Hypothetical protein T9J14.9 [Arabidopsis thali... 105 5e-21
UniRef100_Q9FG44 Similarity to DnaJ protein [Arabidopsis thaliana] 100 9e-20
UniRef100_Q84TH2 Hypothetical protein At4g19570 [Arabidopsis tha... 100 1e-19
UniRef100_O49475 Hypothetical protein F24J7.130 [Arabidopsis tha... 100 1e-19
>UniRef100_O82287 Putative DnaJ protein [Arabidopsis thaliana]
Length = 575
Score = 353 bits (906), Expect = 8e-96
Identities = 242/628 (38%), Positives = 341/628 (53%), Gaps = 92/628 (14%)
Query: 4 SKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT--- 60
S+ E+E++ K LAES + SAL +A++A L P EG+S MVT+ I+S+
Sbjct: 7 SEVEKESIHHKALAESSFNCGDL-MSALTHARKALSLSPNTEGLSAMVTAFEIISSAATV 65
Query: 61 -----DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVL 115
+WY VL VEPF++ N +++QY+KL+L+LHPDKN EE FKL+ EAFRV
Sbjct: 66 AGGFPEWYKVLKVEPFSHINTIKQQYRKLALVLHPDKNPY---VGCEEGFKLLNEAFRVF 122
Query: 116 SDSA--LKKGYDAELRKKEAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAV 173
SD + G + E TF CS CR +H+F+R+Y+G +L+CP C SFEA E
Sbjct: 123 SDKGEMVSGGSGGD----ETSTFSAVCSGCRSVHKFDRKYLGQNLMCPTCKNSFEAKEVE 178
Query: 174 LSDGSSDEG---EKVGVRRNEKREVVLGVAAPNGVKGKVGGEKGLKGKVGDGDG---VGD 227
+ + G K+ KR V D DG + +
Sbjct: 179 KEEEGRENGACTSKIITYSRRKRPV-------------------------DSDGESLIKE 213
Query: 228 VGGGGSKKRMRSVGDVFERSKPKGIITGEETMTLAEFQSQVKRKVQQEKVKDKEEEWGRT 287
V G + +VFE S + E MTLAE Q+ +KR + KV K E +
Sbjct: 214 VETGEMSQEAAEAINVFEGSDEED----EGMMTLAEMQAVIKRN--KSKVNSKITEKDSS 267
Query: 288 DKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRGSEVGEERSLKKNVKLA------- 340
+ + + + + + TL+ + NAV +K+ EV + R K K+A
Sbjct: 268 GEENMGRETQKRSLADASMTETLR-EMSTNAVNNKQ--EVLKNRKNIKKKKMANHKDLTE 324
Query: 341 ---IKEKPEASGKRKRLELEECGDANGGELEVMAVVDSDF--YDFDKDRVERSFKKGQVW 395
++ P KR R +L + M + D DF YDFDKDR+ RSFKKGQ+W
Sbjct: 325 IVDLEYVPRVDRKRDRGKLSQ----------EMYMEDEDFELYDFDKDRMPRSFKKGQIW 374
Query: 396 AAYDD-DDGMPRHYALIDETVSANPFEVRISWLDLQSNADGKIVSREKMGF-HIPCGRFK 453
A YD DD MPR Y L+ E VS NPF+V ISWLD +S K++S K+ H+PCGRF+
Sbjct: 375 AIYDGGDDKMPRSYCLVSEVVSLNPFKVWISWLDFESE---KLISWMKISSSHMPCGRFR 431
Query: 454 VARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWALYGEATLDADGRHFADEGKRCN 513
V+ K I V FSH+V+C+R ARE+Y+IYP+KGSVWA+Y E R R
Sbjct: 432 VSEKALIEQVKPFSHLVNCERAAREIYQIYPKKGSVWAVYSETNPGLQRRK-----TRRY 486
Query: 514 DIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNM-WLISHQIPARK 572
+IVV LT Y + GLS+AYLEKV+ Y +FKR+D G +A+R++ K+++ L+SHQIPA+K
Sbjct: 487 EIVVCLTMYTDAYGLSVAYLEKVNDYSNLFKRRDYGYNAVRWVEKEDVAALLSHQIPAKK 546
Query: 573 SPCIDETPELLEDCWELDPASLPSYLLT 600
P DE+ L++ W LD AS+P L++
Sbjct: 547 LP-EDESGADLKESWVLDLASVPPDLVS 573
>UniRef100_Q9S7L6 Expressed protein [Arabidopsis thaliana]
Length = 706
Score = 250 bits (638), Expect = 1e-64
Identities = 202/709 (28%), Positives = 319/709 (44%), Gaps = 118/709 (16%)
Query: 8 EEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT------- 60
EEALR+K +AE + + SA YA +A+ LFP+LEG+S+MV + + A+
Sbjct: 6 EEALRVKQIAERRF-AEKDFTSARSYALKAKSLFPDLEGLSQMVATFEVYLASQTRSGGQ 64
Query: 61 -DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
D+Y VLG++P A V+KQYKK+++LLHPDKN ++ AF L+ EA+ LS+
Sbjct: 65 IDYYAVLGLKPSAGKREVKKQYKKMAVLLHPDKNK---CIGADGAFHLISEAWSFLSNEF 121
Query: 120 LKKGYDAELRK----------------------------------KEAPTFWTACSACRL 145
K + + +K + TFWT C++C++
Sbjct: 122 NKSTFYYKRKKHIDSTEVQKHSTEYMPGTGTGTGTAVFDRFPPSSERLDTFWTVCTSCKV 181
Query: 146 LHQFERRYVGHSLVCPNCNKSFEAVE---------------------------AVLSDGS 178
+++ R+YV L C NC +F AVE A S+G
Sbjct: 182 QYEYLRKYVNKRLSCKNCRGAFIAVETGPAPVSAPFHYTPPSHAPPSHAPPSHAPPSNGY 241
Query: 179 SDEGEKVGVRRNEKREVVLG--VAAPNGVKGKVGGE------KGLKGKVGDGD--GVGDV 228
G R LG +G G G G+ D V V
Sbjct: 242 GAHGYDAMSRMPTNSTYFLGHYPGQGHGYDYSTNGSYEWSSYSGTTTSPGNLDLKRVSSV 301
Query: 229 GGGGSKKRMRSV-------------GDVFERSKPKGII-TGEETMTLAEFQSQVKRKVQQ 274
G K SV G ++S P G+I + +M+ ++V R ++
Sbjct: 302 SNGYPYKHSNSVVSGGINKVKDGSNGTCSKKSTPGGLIYPNQISMSAHASANKVGRPGKK 361
Query: 275 EKVKDKEEEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRGSEVGEERSLK 334
KV + G + V + + KL I+ + + S + + L
Sbjct: 362 SKVFMEAAANGFVENPLRSVSVSKTANTDAKMDQDYKLHIQSSTRRWSAASVLDTRKPLI 421
Query: 335 KNVKLAIKEKPE--------ASGKRKRLELEE-------CGDANGGELE-VMAVVDSDFY 378
+ + IK++ E A+ L+E GD G + + V DSDF+
Sbjct: 422 QKARTDIKQRLEMMRLALEAAAAAEDATPLDEKTVLSCKLGDVTGRKTNGPITVPDSDFH 481
Query: 379 DFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNADGKIV 438
DFDK+R E SF+ Q+WA YD+DDGMPR Y ++ E +S PF++ I++L +++ + +
Sbjct: 482 DFDKNRSEESFEPRQIWAIYDEDDGMPRLYCVVREVLSVQPFKIDIAYLSSKTDIEFGSM 541
Query: 439 SREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAR-EVYKIYPRKGSVWALYGEAT 497
+ GF CG F++ D ++ VNIFSH++ + R +I+P G +WA+Y +
Sbjct: 542 KWVQYGFTKSCGHFRIRNSDIVDHVNIFSHLLKGKKTGRGGCVRIFPTAGEIWAVYKNWS 601
Query: 498 LDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLG 557
L+ DG DE + ++V L Y E G+ + L K++GYKTV+ R T + +++
Sbjct: 602 LNWDG-STPDEVRHQYEMVEILDEYTEQYGVCVTPLVKLEGYKTVYHR-STREDSKKWIP 659
Query: 558 KDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLLTIGGIDN 606
+ M SHQ+P+ D T E+CW+LDPA++P LL IG N
Sbjct: 660 RCEMLRFSHQVPSWFLK--DATSGFPENCWDLDPAAIPEELLHIGAGTN 706
>UniRef100_Q5VRZ9 DnaJ protein-like [Oryza sativa]
Length = 478
Score = 234 bits (597), Expect = 5e-60
Identities = 120/232 (51%), Positives = 154/232 (65%), Gaps = 18/232 (7%)
Query: 368 EVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWL 427
E+M V DSDFY+FD DR E+ FK+GQVWA Y DDDGMPRHYAL++ F +I WL
Sbjct: 259 EMMDVEDSDFYNFDADRCEKCFKRGQVWALYGDDDGMPRHYALVEMITPGGRFRAQIRWL 318
Query: 428 DLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKG 487
DLQ + G PCG FKV R +++SVNIFSH V +RVAREVY+IYP+KG
Sbjct: 319 DLQPDG----------GEGTPCGEFKVGRTVTVHSVNIFSHQVAYERVAREVYRIYPKKG 368
Query: 488 SVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQD 547
SVWAL+G AD G+ + VVFL+ Y+++ G S YLEKV+G++++F RQD
Sbjct: 369 SVWALHGGKD--------ADSGRPKYEFVVFLSGYSDLYGASFGYLEKVEGFRSIFTRQD 420
Query: 548 TGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLL 599
G A++ L K +M +SHQIPAR++P + + DCWELDPASLPS LL
Sbjct: 421 VGRDAVQTLHKGDMGKLSHQIPARRAPKGEGSTLPPTDCWELDPASLPSELL 472
Score = 97.4 bits (241), Expect = 1e-18
Identities = 62/181 (34%), Positives = 103/181 (56%), Gaps = 21/181 (11%)
Query: 2 APSKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSATD 61
A ++E RL LAE+ L+ + ++A K+A+RA RL P+ S ++T++++L+A D
Sbjct: 11 ADDDDDDELSRLLSLAEADLE-AGRLRAAHKHARRAARLDPDSTRASLLLTAVSVLAADD 69
Query: 62 WY----TVLGVEPFANSN-----AVRKQYKKLSLLLH--PDKNNTHVAAASEEAFKLVGE 110
+L P + ++ A+R+ YK LS L P ++ V++A +EA + +
Sbjct: 70 SSHRATLLLPDSPHSQASPLSPSALRRHYKSLSESLRSAPPSSSPAVSSAVKEALRRAAD 129
Query: 111 AFRVLSDSALKKGYDAELRKKEAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAV 170
A+ L++ A PTFWTAC+ CRLLH+F+R+YVG L+CP+C ++F A
Sbjct: 130 AYAALANQAAAP---------VPPTFWTACAGCRLLHEFDRKYVGFRLMCPSCRRTFLAS 180
Query: 171 E 171
E
Sbjct: 181 E 181
>UniRef100_Q94H83 Putative heat shock protein [Oryza sativa]
Length = 748
Score = 212 bits (540), Expect = 2e-53
Identities = 193/748 (25%), Positives = 323/748 (42%), Gaps = 175/748 (23%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT------ 60
++EALR K +AE K + S + + A K+A +AQ LFP LEGI +M+T+L + A+
Sbjct: 5 KDEALRAKEIAERKFE-SKDLQGAKKFALKAQALFPGLEGIVQMITTLDLYLASEVLISG 63
Query: 61 --DWYT--------------------VLGVEPFANSN----------------------- 75
DWY+ VL + P N +
Sbjct: 64 EKDWYSILSVESSADDETLKKQYRKLVLQLHPDKNKSVGAEGAFKMVQEAWTVLSDKTKR 123
Query: 76 AVRKQYKKLSLLLHPDKNNTHVAAASEEA------FKLVGEAFRVLSDSALKKG-YDAEL 128
A+ Q +KL ++L + + T+ A+A+ A F A +V + K G + +
Sbjct: 124 ALYDQKRKL-MVLKRNTSQTNKASAAPGASNGFYNFAANAAASKVTRGNKQKAGPATSSV 182
Query: 129 RKK-----------------EAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSF---- 167
R++ + PTFWT+C+ C++ +++ + Y+ H+L+CP C + F
Sbjct: 183 RQRPPPPPPPPRQAPAPPPAKPPTFWTSCNKCKMNYEYLKVYLNHNLLCPTCREPFLAQE 242
Query: 168 ------EAVEAVLSDGSSDEGEKVGVRRN------------------------------- 190
E+V AV S + RN
Sbjct: 243 VPMPPTESVHAVHDPNISGANQNTNGSRNFQWGPFSRTAGAASATASSAAAAQAANVVHH 302
Query: 191 -------EKREVVLGVAAPNGVKGKVGGEKGLKGKVGDGDGVGDVGGGGSKKRMRSVG-D 242
E+ E ++ K K + + + +G GG S K+MR++G D
Sbjct: 303 TYEKVRREREEAQAAARREEALRRKYNPPK-RQANISENLNLGT--GGNSSKKMRTMGND 359
Query: 243 VFERSK-------------PKGIITGEETMTLAEFQS---------------QVKRKVQQ 274
+ S P G I+ FQ + + Q
Sbjct: 360 IGIGSSSILSGSGANYFGVPGGNISFSTNSGAHHFQGVNGGFSWKPRPPTRISLVKTFTQ 419
Query: 275 EKVKDKEEEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRG---SEVGEER 331
V+ E ++D + K ++ KR + N K+K+ G++
Sbjct: 420 FDVRGILMEKAKSDLKDKLKEMQTKRS-----------QVAANGKKNKKNMFKESGGDDE 468
Query: 332 SLKKNVKLAIKEKPEASGKRKRLELEECGDANGGELEVMAVVDSDFYDFDKDRVERSFKK 391
SL + A + + + D N L V D DF+DFDKDR E F+
Sbjct: 469 SLASDDSTARQAAHVDPEDNASVNSTDADDENDDPLS-YNVPDPDFHDFDKDRTEECFQS 527
Query: 392 GQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNADGKIVSREKMGFHIPCGR 451
Q+WA YDD+DGMPR+YA I + +S PF+++IS+L ++N++ ++ GF CG
Sbjct: 528 DQIWATYDDEDGMPRYYAFIQKVLSLEPFQLKISFLTSRTNSEFGSLNWVSSGFTKTCGD 587
Query: 452 FKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWALYGEATLDADGRHFADEGKR 511
F++ R ++ + +N+FSH + ++ R V KIYP+KG++WA+Y + D D D+
Sbjct: 588 FRICRYETCDILNMFSHQIKWEKGPRGVIKIYPQKGNIWAVYRNWSPDWD-EDTPDKVLH 646
Query: 512 CNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPAR 571
D+V L Y+E G+S+ L KV G++TVF+R + +AI+ + K+ M+ SH++P
Sbjct: 647 AYDVVEVLDEYDEDLGISVIPLVKVAGFRTVFQR-NQDLNAIKKIPKEEMFRFSHEVPFY 705
Query: 572 KSPCIDETPELLEDCWELDPASLPSYLL 599
+ +E P + +D +ELDPA++ LL
Sbjct: 706 RMSG-EEAPNVPKDSYELDPAAISKELL 732
>UniRef100_Q9SLA7 At2g25560/F13B15.22 [Arabidopsis thaliana]
Length = 656
Score = 209 bits (533), Expect = 1e-52
Identities = 201/691 (29%), Positives = 289/691 (41%), Gaps = 144/691 (20%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTI-LSA------ 59
+EEA R + +A+ K +N+ A K+A +AQ L+PEL+GI++MV + + LSA
Sbjct: 5 KEEATRAREIAKRKF-LANDFAGARKFALKAQFLYPELDGIAQMVATFDVHLSAQNIIYG 63
Query: 60 -TDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAF------------- 105
D Y VLG+ P A+ VRK+Y+KL+++LHPD+N + +EEAF
Sbjct: 64 DVDHYGVLGLNPEADDEIVRKRYRKLAVMLHPDRNKS---VGAEEAFKFLSQAWGVFSDK 120
Query: 106 ----------------------------------KLVGEAFRVLSDS-ALKKGYDAELRK 130
K G +V S +K+ DA
Sbjct: 121 AKRADYDLKRNVGLYKGGGASSSRPATNGFQKVTKASGNTTKVKSSKRGIKRASDASAAA 180
Query: 131 KEAP---------TFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAVLSDGSSDE 181
+ TFWT C CR +++ Y+ +L+CPNC K F AVE +D
Sbjct: 181 TTSTSAQKTTADGTFWTVCRTCRTQYEYHSVYLNQNLLCPNCRKPFIAVE-------TDP 233
Query: 182 GEKVGVRRNEKREVVLGVAAPNGVKGKVGGEKGLKGKVGDGDGV-GDVGGGGSKKRMRSV 240
+R+ + G +K + G+ D +GV G+
Sbjct: 234 PGSGSIRKTFHEHQFDSLRHTTD-----GRKKNVPGR--DNNGVYGEY------------ 274
Query: 241 GDVFERSKPKGIITGEETMTLAEFQSQVKRKVQQEKVKDKEEEWGRTDKRSSRKRVENKR 300
D FE G+ TG +T A K +V + + + T RK +EN
Sbjct: 275 -DSFEW----GVFTGTKTSAHATPTGSRKDEVVRREYTKRTAGPSSTIPPKRRKVMEN-- 327
Query: 301 GLEIGEVRTLKLPIKENAVKSKRGSEVGEERSLKKNVKLAIKEK--------PEASGKRK 352
+ G + + P K VK E+ + LKK K I E +
Sbjct: 328 AVAGGNIASCLAP-KSTGVKEVSEDEL--KNLLKKKAKSVISRNLPELCTIVAETETNGR 384
Query: 353 RLELEECGDANGGELEVMAVVDS----------------------DFYDFDKDRVERSFK 390
+E E+ N G ++S DF DFDKDR E+S K
Sbjct: 385 GMETEDLNGFNAGSSVNKNAIESCCMNTDKELNSLGALTLDVTAPDFCDFDKDRTEKSVK 444
Query: 391 KGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNADGKIVSREKMGFHIP-- 448
Q+WA YD +G+PR YALI +S +PF+VR+SWL +N G+ S +GF IP
Sbjct: 445 DNQIWAFYDSHEGLPRSYALIHNVISVDPFKVRMSWLTPVTN--GEPSSTNWLGFGIPKS 502
Query: 449 CGRFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWALYGEATLDADGRHFADE 508
CG F+V + S FSH V+ + + IYPR G VWALY + D +
Sbjct: 503 CGGFRVRKTLIYRSPYSFSHKVNLVKGNHGEFLIYPRTGDVWALYRK--WSPDWNYLTGV 560
Query: 509 GKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQI 568
DIV + Y E G+ + L KV G+K VF RFL +D + SH+I
Sbjct: 561 ETVEYDIVEVVEGYTEEYGVVVVPLVKVAGFKAVFHHHLDSKETKRFL-RDEISRFSHKI 619
Query: 569 PARKSPCIDETPELLEDCWELDPASLPSYLL 599
P+ E P C +LDPA+ PS LL
Sbjct: 620 PSYLLTG-QEAPGAPRGCRQLDPAATPSQLL 649
>UniRef100_Q9CAW2 Hypothetical protein T9J14.7 [Arabidopsis thaliana]
Length = 1165
Score = 199 bits (505), Expect = 3e-49
Identities = 200/714 (28%), Positives = 304/714 (42%), Gaps = 162/714 (22%)
Query: 30 ALKYAKRAQRLFPELEGISEMVTSLTILSAT--------DWYTVLGVEPFANSNAVRKQY 81
A K+ +AQRLFP LE I +M+T + S+ DWY VL V+P+A+++ ++KQY
Sbjct: 9 AHKFVTKAQRLFPNLENIVQMMTICDVHSSAIKKIKGLDDWYGVLQVQPYADADTIKKQY 68
Query: 82 KKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSALKKGYDAELRKKE--------- 132
+KL+LLLHPDKN A +E AFKLVGEA R+LSD + YD R
Sbjct: 69 RKLALLLHPDKNKF---AGAEAAFKLVGEANRLLSDQIKRSQYDNRYRSHSMFANRHVNV 125
Query: 133 ------------------APTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAVL 174
TFWT C C +++ R Y+ S+ C +C KSF A + +
Sbjct: 126 YSGRHCAATNNAAENIAGVFTFWTRCRHCGQCYKYLREYMNTSMHCSSCQKSFVACK-MR 184
Query: 175 SDGSSDEGEKVGVRRNEKREVVL---------------GVAAPNGVKGKVGG-------E 212
DG G R E ++ V+ AA G GKVGG E
Sbjct: 185 CDGVPPSSSTAG--RKEFQDQVMSNTSRQNASTAAESGSSAADMGKNGKVGGKVNKKNQE 242
Query: 213 KGLKGKVGDG----DGVGDVGGGGSKKRMRSVGDVFERSKPKGII------TGEETMTLA 262
K G V G +G + G K + G PK + EE T A
Sbjct: 243 KQKNGAVNRGTKKEEGCTENDAEGRKPQTSETGTNISAEMPKADVLKPQHQVKEEPDTSA 302
Query: 263 EFQ------------SQVKRKVQQEKVKDKEEE-----WGRTD------KRSSRKRVENK 299
E + KRKV +E K E + +TD ++SSRK+ +
Sbjct: 303 EKSIPDLSAPQKNRAPKKKRKVVEESSKSFEVDSSDTAGAKTDTNEHNKRKSSRKKPQVS 362
Query: 300 RGLEIGEVRTLKLPIKENAVKSKRGSEVGEERSLK------KNVKLAIKEKPEAS----- 348
E + + P K K+K G E E + K K+ KLA+ AS
Sbjct: 363 YAKEGSDGDFVSPPNK----KTKSGFEFESEPNKKQTAEDNKSPKLAVSGVSSASSHSYK 418
Query: 349 GKRK-------------RLELEECGDANGGELEVMA-------------------VVDSD 376
GK K + ++ E D NG + +++ + D +
Sbjct: 419 GKAKKNAHSGNEDNLSAKNKVSEGCDGNGEDAALLSKIGRVEKGYKANENSNPLDIPDLE 478
Query: 377 FYDFDKDRVERSFKKGQVWAAYDD-DDGMPRHYALIDETVSANPFEVRISWLDLQSNADG 435
F F +R F QVW+ D DGMPR YA + + ++ F++RI++LD
Sbjct: 479 FSVFKVERKTEDFAVNQVWSTTTDCRDGMPRKYARVKKVLNGE-FKLRITYLD------- 530
Query: 436 KIVSREKMGFHIPCGRFKVARKDSINSVNIFS---HVVDCDRVAREVYKIYPRKGSVWAL 492
++ + + CG+FK + + +IFS H + C+ + IYPRKG +WA+
Sbjct: 531 PVLDKTDESIPVACGKFKNGKTMEVKDSSIFSGQMHHLRCNNIV----SIYPRKGEIWAI 586
Query: 493 YGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFK-RQDTGSH 551
+ E + + + D V +++++++NG+ +AYL K+ G +F G
Sbjct: 587 FREWEEEWNTSLKKHKFPYKYDFVEIVSDFHDLNGVGVAYLGKLKGSVQLFHWEPQHGIC 646
Query: 552 AIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLLTIGGID 605
I+ KD M SH++PA K E + + +ELDPA+LP + + +D
Sbjct: 647 QIQCSPKD-MLRFSHKVPAVKMTG-KEKESVPPNSYELDPAALPKDIFQVDAVD 698
Score = 81.3 bits (199), Expect = 8e-14
Identities = 67/230 (29%), Positives = 106/230 (45%), Gaps = 30/230 (13%)
Query: 378 YDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNADGKI 437
+DF R E F+ Q+WA Y +D+GMP Y I + + F +R + +L
Sbjct: 959 FDFQNLRSEDKFEVNQIWAIYSNDNGMPLEYVKIKKIETKPKFVLRGTPTELYP------ 1012
Query: 438 VSREKMGFHIPCGRFKVAR-KDSINSVNIFSHVV-DCDRVAREVYKIYPRKGSVWALYG- 494
S E + + CG FK+ + + I FSH+V D R +K+YPRKG +WALY
Sbjct: 1013 PSTEPVTRTVSCGEFKLLKGRPKIIPHASFSHLVKPFDSSKRFRFKVYPRKGEIWALYKN 1072
Query: 495 -EATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAI 553
++T + D ++ C+ +V + M + F+R+ +
Sbjct: 1073 CDSTEEPDIVEVVED--NCDGEIVKVVALTAMG--------------SSFQRKQGSDVGL 1116
Query: 554 RFLGKDNMWLISHQIPARKSPCIDETPELLED--CWELDPASLPSYLLTI 601
+ K M SHQIPA + P +T L++ WELDP ++PS + I
Sbjct: 1117 IDISKAEMSRFSHQIPAIRHP--KKTTRLVKGGYYWELDPIAIPSRTIVI 1164
>UniRef100_Q6NMG4 At5g53150 [Arabidopsis thaliana]
Length = 726
Score = 182 bits (463), Expect = 2e-44
Identities = 142/480 (29%), Positives = 229/480 (47%), Gaps = 22/480 (4%)
Query: 131 KEAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAVLSDGSSDEGEKVGVRRN 190
+E+ TFWT C+ C ++++R Y+ +L+CP+C+ F A E + ++ V + N
Sbjct: 199 QESSTFWTMCNKCDTQYEYQRVYLNQTLLCPHCHHGFVAEEK--TPPTNIPKPPVNISSN 256
Query: 191 EKREVVLGVAAPNGVKGK-VGGEKGLKGKVGDGDGVGDVGGGGSKKRMRSVGDVFERSKP 249
+ A+ G E +GG S+ +V ++ +
Sbjct: 257 QHHRSSKNQASNKNSNGSSYRREPATSVNHNFQWDSSRMGGSYSRNATNETANVVQQGQD 316
Query: 250 KGIITGEETMTLAEFQSQVKRKVQQEKVKDKEEEWGRTDKRSSRK-------RVENKRGL 302
K ET + + K + ++ R SR R + K+ L
Sbjct: 317 KLKRVFWETQEREAARGFTNSDLGNFKRQKTDDSHMRGPSAGSRHPYVQALLRSDIKKAL 376
Query: 303 -EIGEVRTLK-LPIKENAVKSKRGSEVGEERSLKK-NVKLAIKEKPEASGKRKRLELE-E 358
+ G+ K LP+ ++ K GE+ S K + K + E+ + S +E E
Sbjct: 377 MDRGQSEIFKRLPMMIAKMEGKVNPTEGEKNSTKAMSSKASEVERSKMSSTANEVERSVE 436
Query: 359 CGDANGGELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSAN 418
E++ + V DSDF++FD DR E +FK Q+WAAYDD DGMPR YA I + +S N
Sbjct: 437 VIPHESDEVKEIVVPDSDFHNFDLDRSESAFKDDQIWAAYDDADGMPRFYARIQKVISVN 496
Query: 419 PFEVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVARE 478
PF+++ISWL+ ++ ++ + GF CG F+ R +S +++N FSH VD + AR
Sbjct: 497 PFKLKISWLNSKTTSEFGPIDWMGAGFAKSCGDFRCGRYESTDTLNAFSHSVDFTKGARG 556
Query: 479 VYKIYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMN-GLSMAYLEKVD 537
+ I P+KG VWALY + + D ++ DE K ++V L +Y E + L++A L K +
Sbjct: 557 LLHILPKKGQVWALYRNWSPEWD-KNTPDEVKHKYEMVEVLDDYTEDDQSLTVALLLKAE 615
Query: 538 GYKTVFKRQDTGSHAIRFLGKDNMWLISHQIP--ARKSPCIDETPELLEDCWELDPASLP 595
G++ VF+R T +R + K+ M SHQ+P D P E ELDPA+ P
Sbjct: 616 GFRVVFRR-CTEKLGVRKIAKEEMLRFSHQVPHYILTGKEADNAP---EGFLELDPAATP 671
Score = 99.4 bits (246), Expect = 3e-19
Identities = 57/143 (39%), Positives = 84/143 (57%), Gaps = 18/143 (12%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSL--------TILS 58
++EA R +AE K+ T + A K+A +AQ LFPEL+G+ ++ ++ T
Sbjct: 5 KDEAKRAMDIAERKM-TEKDYTGAKKFANKAQNLFPELDGLKQLFVAINVYISGEKTFAG 63
Query: 59 ATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
DWY VLGV+PFA+ A++KQY+KL L+LHPDKN +E AF LV EA+ +LSD
Sbjct: 64 EADWYGVLGVDPFASDEALKKQYRKLVLMLHPDKNK---CKGAEGAFNLVAEAWALLSDK 120
Query: 119 ------ALKKGYDAELRKKEAPT 135
+K+G D + ++ PT
Sbjct: 121 DKRILYNVKRGKDVKAAQQRFPT 143
>UniRef100_Q9SQT7 Putative DnaJ protein [Arabidopsis thaliana]
Length = 673
Score = 181 bits (459), Expect = 6e-44
Identities = 168/676 (24%), Positives = 266/676 (38%), Gaps = 113/676 (16%)
Query: 4 SKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT--- 60
S +EALR K LAE L + +A K A +AQ++ LE IS M+ + A
Sbjct: 2 SINRDEALRAKDLAEG-LMKKTDFTAARKLAMKAQKMDSSLENISRMIMVCDVHCAATEK 60
Query: 61 ------DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRV 114
DWY +L VE AN ++KQYK+L+LLLHPDKN +E AFKL+GEA R+
Sbjct: 61 LFGTEMDWYGILQVEQIANDVIIKKQYKRLALLLHPDKNKL---PGAESAFKLIGEAQRI 117
Query: 115 LSDSALKKGYDAEL---RKKEAP------------------------------------- 134
L D + +D + RK AP
Sbjct: 118 LLDREKRTLHDNKRKTWRKPAAPPYKAQQMPNYHTQPHFRASVNTRNIFTELRPEIRHPF 177
Query: 135 --------------TFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAVLSDGSSD 180
TF T+C CR+ ++++R +V + C C K F A E L
Sbjct: 178 QKAQAQPAAFTHLKTFGTSCVFCRVRYEYDRAHVNKEVTCETCKKRFTAFEEPLQSAPQA 237
Query: 181 EGEKVGV-------------------RRNEKREVVLGVAAPNGVKGKVGGEKGLKGKVGD 221
+G +R E V A + G G + +
Sbjct: 238 KGPSQTTYCFPQQSKFPDQRACSEPHKRPENPPTVSSSKASFPMPGSTAKHNGKRKRKNV 297
Query: 222 GDGVGDVGGGGSKKRMRSVGDVFERSKPKGIITGEETMTLAEFQSQVKRKVQQEKVKDKE 281
+ S + V + ++ G GE+ + +V E + D +
Sbjct: 298 AECSESSDSESSSESEDDVNNDTTAAQDSGSNGGEQPRRSVRSKQKVS---YNENLSDDD 354
Query: 282 EEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRGSEVGEERSLKKNVKLAI 341
+ + S K ++ +R KE + + S+ N K+ +
Sbjct: 355 VDLVNDNGEGSGKNIDTERE-------------KETEEEKQTNENHSSTESIDMNGKIEV 401
Query: 342 KEKPEASGKRKRLELEECGDANGGELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDD 401
+ SG E G A L + D DF DFDK R + F+ GQ+WA YD++
Sbjct: 402 DQVETPSGASDSEEDLSSGSAEKPNL--INYDDPDFNDFDKLREKSCFQAGQIWAVYDEE 459
Query: 402 DGMPRHYALIDETVSANPFEVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSIN 461
+GMPR YALI + V+ F +R W ++ + + E + G+F V + N
Sbjct: 460 EGMPRFYALI-KKVTTPDFMLRYVWFEVDQDQE-----NETPNLPVSVGKFVVGNIEETN 513
Query: 462 SVNIFSH-VVDCDRVAREVYKIYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLT 520
+IFSH V ++ + ++P+KG +WAL+ ++ + K + V L+
Sbjct: 514 LCSIFSHFVYSTTKIRTRKFTVFPKKGEIWALFKNWDINCSADSVSPM-KYEYEFVEILS 572
Query: 521 NYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETP 580
++ E +S+ +L KV G+ VF + SH IP+ + E
Sbjct: 573 DHAEGATVSVGFLSKVQGFNCVFCPMPKDESNTCEIPPHEFCRFSHSIPSFRLTG-TEGR 631
Query: 581 ELLEDCWELDPASLPS 596
+ + +ELDPA+LP+
Sbjct: 632 GITKGWYELDPAALPA 647
>UniRef100_Q5Z5F4 Putative DNAJ heat shock N-terminal domain-containing protein
[Oryza sativa]
Length = 1018
Score = 173 bits (439), Expect = 1e-41
Identities = 195/713 (27%), Positives = 294/713 (40%), Gaps = 141/713 (19%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEG-ISEMVTSLTILSAT----- 60
++EA++ K LAE K++ + A + +AQ L +++ IS+M+T I A+
Sbjct: 5 KDEAVKAKALAEKKMREKDFA-GAKRMINKAQNLSKDVDSNISQMLTVCDIHCASATKVN 63
Query: 61 ---DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSD 117
DWY +L V A+ ++KQY+KL+LLLHPDKNN A +E AFKLVGEA L+D
Sbjct: 64 GEIDWYGILQVPVTADDTLIKKQYRKLALLLHPDKNNF---AGAEAAFKLVGEANMTLTD 120
Query: 118 SALKKGYD----------------------AELRKKEAP--------------------- 134
+ + YD A +R P
Sbjct: 121 RSKRSVYDMKRNASVRIGSARVPYQQSRRTAPVRPTTTPVNLHNVHQSQQHKPSNPSDSQ 180
Query: 135 TFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAV---EAVLSDGSSDEGEKV----GV 187
TFWT C C + +Q+ + +L C NC K F A+ E + G++ V G
Sbjct: 181 TFWTICPTCGMRYQYYLSILKKALRCQNCLKPFVALDLNEQAVPSGANQRSAGVWKSSGA 240
Query: 188 RRN-EKREVVLGVAAPNGV-----------------KGKVGGEKG-LKGKVGDGDGVGDV 228
+N + +G A N +G G E G LK K G+
Sbjct: 241 PQNFPGSQANVGQQAQNSANPVHANFGSHNAHVETKRGADGNEAGGLKNKRKFAKATGNS 300
Query: 229 GGG----GSKKRMR--------SVGDVFERSKPKGIITGEETMTLAEFQSQVKRKVQQEK 276
GSKKR + S D S+ + I G + Q + Q+++
Sbjct: 301 SKASSVAGSKKRRKAMFESSESSASDTSTDSEEEIIEDGPAASNVGPDQHPRRSSRQKQE 360
Query: 277 VKDKEEEWGR-TD----------KRSSRKRVENKRGLEIGEVRTLKLPIK---------- 315
VK E+ G TD S KR+ GE KL
Sbjct: 361 VKYNEDSDGDDTDCHGNGDDGFVSSPSLKRLRKGGLFHGGENNETKLNADTTGPGHDGPT 420
Query: 316 ------ENAVKSKRGSEVGEE---RSLKKNVKLAIKEKPEASGKRKRLELEECGDANGGE 366
N +RGS E+ ++ A KEK S LE +++
Sbjct: 421 NGVNNYNNTEDIERGSACAEQIKRETMSGGGNSAEKEKLSHSVSNNGLE----SNSDDAP 476
Query: 367 LEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISW 426
EV+ DS+F+DF++ R FK Q+WA YD MPR+YA I + F + W
Sbjct: 477 NEVICA-DSEFFDFNQLRHINQFKANQIWACYDSQSCMPRYYARITKVKHVPKFMLNFIW 535
Query: 427 L--DLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDR-VAREVYKIY 483
L D ++ A+ S + + CGRFK D+ ++FSH + ++ R Y+IY
Sbjct: 536 LEFDPKNKAEAAWSSGD---LPVSCGRFKHGVSDTAKESSMFSHAIFYEKNKTRNSYEIY 592
Query: 484 PRKGSVWALYGEATLDADGRHFADEGKRCN-DIVVFLTNYNEMNGLSMAYLEKVDGYKTV 542
PRKG VWAL+ D D AD+ K ++V L++ + + L K+ G+ ++
Sbjct: 593 PRKGEVWALF--KGWDIDWSADADKHKNYEYEVVQVLSDLTSSTSIIVMPLVKIKGFVSL 650
Query: 543 FKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLP 595
F + S + + +D+ SH +P R + E + E ELDPA+LP
Sbjct: 651 FIQSKEASPYV--IPQDDTLRFSHCVP-RHTMIGTEKEGIPEGAIELDPAALP 700
Score = 100 bits (249), Expect = 1e-19
Identities = 72/234 (30%), Positives = 112/234 (47%), Gaps = 23/234 (9%)
Query: 374 DSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDL--QS 431
+S+F++F + R F+ GQ+WA Y D D P +YA I +TV E+++ WLD QS
Sbjct: 795 ESEFFEFSEIRSIHKFQPGQIWALYSDVDKFPNYYACI-KTVDVKNNELQVRWLDACPQS 853
Query: 432 NADGKIVSREKMGFHIPCGRFKVARKDSI---NSVNIFSHVVDCDRVAREVYKIYPRKGS 488
+ ++V + + CG FK++ I N SH V R Y+I P +G
Sbjct: 854 EEERRLVRED---LTVACGTFKISSFHGIQTYNGTEYLSHPVQAKPGRRNEYEIVPCQGD 910
Query: 489 VWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDT 548
+WA++ + + K+C+ +V + + + + + + L KVDGY+ VF D
Sbjct: 911 IWAVFKNWRTGWTAKDY----KKCDYELVEIFGHTD-SSIQVQLLRKVDGYRAVF-MPDR 964
Query: 549 GSHAIRFLGKDNMWLISHQIPARKSPCIDETPE---LLEDCWELDPASLPSYLL 599
A++ + KD SHQI PC T E L ELDP S+P L
Sbjct: 965 REGAVKTIRKDEYPKFSHQI-----PCFHLTNERGGKLRGFLELDPLSVPEMFL 1013
>UniRef100_O04648 A_TM021B04.9 protein [Arabidopsis thaliana]
Length = 1609
Score = 165 bits (418), Expect = 3e-39
Identities = 181/717 (25%), Positives = 300/717 (41%), Gaps = 151/717 (21%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSATD----- 61
+EEA R K LAE K++ + A K +AQ LF LE + +M+ + ++ +
Sbjct: 5 KEEACRAKTLAEDKMKEGDFV-GAQKLLLKAQSLFSGLESLPQMLAVCDVHNSAEKKINC 63
Query: 62 ---WYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
WY +L V FA+ ++KQ +KL+LLLHPDKN +E AFKLV +A R L+D
Sbjct: 64 LENWYGILQVMHFADDATIKKQVRKLALLLHPDKNQF---PGAEAAFKLVWDASRFLADK 120
Query: 119 ALKKGYDAELR---------------------KKEAPTFWTACSACRLLHQFERRYVGHS 157
+ YD R TFWT C C +++ R+YV
Sbjct: 121 DKRSQYDIRRRIYLRLATNQLNANSGLQCAATNSATDTFWTCCEHCGYRYKYLRKYVNIL 180
Query: 158 LVCPNCNKSFEAVEAVLSDGSSDE--------------------GEKVGVRRNEKREVVL 197
L C C +S+ A + ++ S GE +G +
Sbjct: 181 LNCNICQRSYMAYDTGFNEAPSKSNTGQKEVQNQGPCNTSLNTNGESIGAQPGS------ 234
Query: 198 GVAAPNGVKG--------KVGGEKGLKGKVGDGDGVGDVGGGGSKKRMRSVGDVFE---- 245
VAA KG + GG + K +V + + +++M+S + +
Sbjct: 235 -VAAEVDKKGTFNKKFNKRNGGGESKKTEVKNSKIETEFVRNDDEEKMKSAAKLHKPHPE 293
Query: 246 ---------RSKP-KGIITGEETMTLAEFQSQVKRKVQQ-----------EKVKDKEEEW 284
+S P + + +E + ++ ++++K+ V++ +K K+
Sbjct: 294 VTEPEIGASKSVPDESVSRSDEAPSTSKDKNKMKKGVEESINVDGSDFIKDKAYSKDNNK 353
Query: 285 GRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENA-VKSKRGSEVGE-------------- 329
++ +RS + + + + K ++ N +KS++ + G
Sbjct: 354 RKSPRRSQQSSYAEEEKISDNSLGPPKKRLRSNVGLKSEQTTRKGLVGVGSSKRFDSGGG 413
Query: 330 --------ERSLKK----------NVKLAIKEKPEASGKRKRLELEECGDANGGE-LEVM 370
+R KK +VK +E +ASG+ +E + + N E L
Sbjct: 414 SSAPSCAFKRKAKKFVDSGYQEISSVKDKEREVRKASGEGVVMEAKIDNNHNPNENLITE 473
Query: 371 AVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLD-L 429
+ D +F +F+ F QVW+ YD DGMPR YA ID+ V F++ I+W+D L
Sbjct: 474 DLPDPEFSNFEL--TTSCFGVNQVWSMYDPIDGMPRLYARIDK-VLVPEFKLWITWIDPL 530
Query: 430 QSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFS----HVVDCDRVAREVYKIYPR 485
Q N D I I CG F+ + N FS H+ + V IYPR
Sbjct: 531 QDNKDNSI--------PIACGIFQGGGSEEENDHLKFSCQMFHLTRNNSVV-----IYPR 577
Query: 486 KGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKR 545
KG +WA++ + + D V L+N+N+ NGL + +L KV+G+ ++F R
Sbjct: 578 KGEIWAIFRGWDISWSASSENHKHPYEYDFVEVLSNFNDENGLGVGFLGKVEGFVSLF-R 636
Query: 546 QDTGSHAIRF-LGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLLTI 601
QD ++ + M SH++P+ K E + C+ELDPA+LP L +
Sbjct: 637 QDAQDGVLQLQIPPSQMLRFSHKVPSFKMTG-KEREGVPPGCFELDPAALPKELFEV 692
Score = 68.6 bits (166), Expect = 5e-10
Identities = 79/326 (24%), Positives = 137/326 (41%), Gaps = 43/326 (13%)
Query: 177 GSSDEGEKVGVRRNEKREVVLGVAAPNGVKG-KVGGEKGLKGKVGDGDGVGDVGGGG--S 233
G E KV ++ N RE ++P K KV G K DG S
Sbjct: 712 GPFPEASKVEMQANTTRE-----SSPFKKKRPKVHDNHGSFSKESDGRRTNHERNSAKES 766
Query: 234 KKRMRSVGDVFERSKPKGIITGEETMTLAEFQSQVKRKVQQEKVKDKEEEWGRTDKRS-S 292
+K +++V + R P+ + + T ++ S+ R++ ++K + E G+ S S
Sbjct: 767 RKSVKAVDALNLRKSPRLL-----SETNSQANSRSGRQIAEKKSANNGESCGKPGGLSLS 821
Query: 293 RKRVENKRGLEIGEVRTLKLPIKENAVKSKRGSEVGEERSLKKNVKLAIKEKPEASGKRK 352
+++ K I +R I + V K+ E S + L + +G K
Sbjct: 822 GEKMSEKT---ITRMRKTPQDIHKPTVNVKKHGRNDESLSQSRGDGLLT----QLNGNMK 874
Query: 353 RLELEECGDANGGELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALID 412
E E ++ V + ++F+ R F+ Q+WA Y +D G PR YA I
Sbjct: 875 SSEPETRVPSS------CKTVKENTFNFENQRSWDKFQIDQIWAIYSNDKGSPRKYAQIK 928
Query: 413 ETVSANPFEVRISWLDLQSNADGKIVSREKMGFHIP----CGRFKV-ARKDSINSVNIFS 467
+ ++ F++ ++ L+L + H+P CGRFK+ K + + FS
Sbjct: 929 KIDTSPEFKLHVAPLELY-----------RPPIHMPRPVCCGRFKLKTGKAEVYVPSSFS 977
Query: 468 HVVDCDRVAREVYKIYPRKGSVWALY 493
H V + + +++YP KG +WALY
Sbjct: 978 HQVKAVKTKKNRFEVYPGKGEIWALY 1003
>UniRef100_Q9FGM2 DnaJ protein-like [Arabidopsis thaliana]
Length = 755
Score = 162 bits (409), Expect = 3e-38
Identities = 105/290 (36%), Positives = 160/290 (54%), Gaps = 10/290 (3%)
Query: 311 KLPIKENAVKSKRGSEVGEERSLKK-NVKLAIKEKPEASGKRKRLELE-ECGDANGGELE 368
+LP+ ++ K GE+ S K + K + E+ + S +E E E++
Sbjct: 416 RLPMMIAKMEGKVNPTEGEKNSTKAMSSKASEVERSKMSSTANEVERSVEVIPHESDEVK 475
Query: 369 VMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLD 428
+ V DSDF++FD DR E +FK Q+WAAYDD DGMPR YA I + +S NPF+++ISWL+
Sbjct: 476 EIVVPDSDFHNFDLDRSESAFKDDQIWAAYDDADGMPRFYARIQKVISVNPFKLKISWLN 535
Query: 429 LQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGS 488
++ ++ + GF CG F+ R +S +++N FSH VD + AR + I P+KG
Sbjct: 536 SKTTSEFGPIDWMGAGFAKSCGDFRCGRYESTDTLNAFSHSVDFTKGARGLLHILPKKGQ 595
Query: 489 VWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMN-GLSMAYLEKVDGYKTVFKRQD 547
VWALY + + D ++ DE K ++V L +Y E + L++A L K +G++ VF+R
Sbjct: 596 VWALYRNWSPEWD-KNTPDEVKHKYEMVEVLDDYTEDDQSLTVALLLKAEGFRVVFRR-C 653
Query: 548 TGSHAIRFLGKDNMWLISHQIP--ARKSPCIDETPELLEDCWELDPASLP 595
T +R + K+ M SHQ+P D P E ELDPA+ P
Sbjct: 654 TEKLGVRKIAKEEMLRFSHQVPHYILTGKEADNAP---EGFLELDPAATP 700
Score = 99.4 bits (246), Expect = 3e-19
Identities = 57/143 (39%), Positives = 84/143 (57%), Gaps = 18/143 (12%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSL--------TILS 58
++EA R +AE K+ T + A K+A +AQ LFPEL+G+ ++ ++ T
Sbjct: 5 KDEAKRAMDIAERKM-TEKDYTGAKKFANKAQNLFPELDGLKQLFVAINVYISGEKTFAG 63
Query: 59 ATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
DWY VLGV+PFA+ A++KQY+KL L+LHPDKN +E AF LV EA+ +LSD
Sbjct: 64 EADWYGVLGVDPFASDEALKKQYRKLVLMLHPDKNK---CKGAEGAFNLVAEAWALLSDK 120
Query: 119 ------ALKKGYDAELRKKEAPT 135
+K+G D + ++ PT
Sbjct: 121 DKRILYNVKRGKDVKAAQQRFPT 143
Score = 45.8 bits (107), Expect = 0.004
Identities = 16/41 (39%), Positives = 28/41 (68%)
Query: 131 KEAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVE 171
+E+ TFWT C+ C ++++R Y+ +L+CP+C+ F A E
Sbjct: 199 QESSTFWTMCNKCDTQYEYQRVYLNQTLLCPHCHHGFVAEE 239
>UniRef100_Q7XKR2 OSJNBa0053B21.9 protein [Oryza sativa]
Length = 729
Score = 161 bits (408), Expect = 5e-38
Identities = 118/361 (32%), Positives = 181/361 (49%), Gaps = 31/361 (8%)
Query: 248 KPKGIITGEETMTLAEFQSQVKRKVQQEKVK----DKEEEWGRTDKRSSRKRVENKRGLE 303
K + ++G L E R + EK K +K +EW T SS + E RG
Sbjct: 379 KLRSAVSGRRANVLREISQIDTRALLIEKAKAAIQEKLQEWNIT---SSSRLAE--RGKS 433
Query: 304 IGEVRTLKLPIKEN---AVKSKRGSEVGEERSLKKNVKLAIKEKPEASGKRKRLELEECG 360
G+V IK+N + K +G + RS+ ++ PE ++R+ +
Sbjct: 434 QGKVYPSDNNIKQNGGLSDKHVKGLKQCSSRSVDTQAPTVDEKNPE----QRRVPVS--- 486
Query: 361 DANGGELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPF 420
+ V D DF+DFDKDR ER+F QVWA YD +DGMPR YA++ + +S PF
Sbjct: 487 ---------IDVPDPDFHDFDKDRTERAFDSDQVWATYDSEDGMPRLYAMVQKVLSMRPF 537
Query: 421 EVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVY 480
+R+S+L+ +SN++ +S GF CG F+V R +VNIFSH V + R +
Sbjct: 538 RIRMSFLNSKSNSELAPISWVASGFQKTCGDFRVGRYQISETVNIFSHKVSWTKGPRGII 597
Query: 481 KIYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYK 540
+I P+KG WALY + D + D+ +IV + ++ + GL++ L KV G+K
Sbjct: 598 RIVPQKGDTWALYRNWSPDWN-ELTPDDVIYKYEIVEIIDDFTDEQGLTVIPLLKVAGFK 656
Query: 541 TVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLLT 600
VF R A R + K+ ++ SH++P+R +E + C ELDPA+ P LL
Sbjct: 657 AVFHRHMDPKEA-RRIPKEELFRFSHRVPSRLLTG-EEGNNAPKGCHELDPAATPVDLLK 714
Query: 601 I 601
+
Sbjct: 715 V 715
Score = 132 bits (332), Expect = 3e-29
Identities = 79/217 (36%), Positives = 116/217 (53%), Gaps = 57/217 (26%)
Query: 8 EEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSA-------- 59
+EA++ +G+AES+ S + + A KYA +AQ L P LEGIS+MV++L + A
Sbjct: 2 DEAIKARGVAESRFH-SRDIRGARKYAIKAQNLCPSLEGISQMVSTLEVHLAAESKIDGE 60
Query: 60 TDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
+DWY +L + FA+ V+KQY+KL+L LHPDKN + +EEAFKL+ EA+ VLSD++
Sbjct: 61 SDWYRILSLTAFADEEEVKKQYRKLALQLHPDKNK---SVGAEEAFKLISEAWSVLSDNS 117
Query: 120 LKKGYD---------------------------------------------AELRKKEAP 134
K YD A R
Sbjct: 118 KKVLYDQKRKDHSVVNVTNGMYTYDKKANKRARKNAAAAAAAAAAAAAAAEATTRPAGVD 177
Query: 135 TFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVE 171
TFWT+C+ CR+ +++ R Y+ H+L+CPNC+ +F AVE
Sbjct: 178 TFWTSCNRCRMQYEYLRIYLNHNLLCPNCHHAFLAVE 214
>UniRef100_Q6K2F0 Heat shock protein-like [Oryza sativa]
Length = 734
Score = 161 bits (407), Expect = 6e-38
Identities = 90/230 (39%), Positives = 134/230 (58%), Gaps = 3/230 (1%)
Query: 372 VVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQS 431
V D DFYDFDKDR ER+F QVWA YD +DGMPR YA++ + +S PF +R+S+L+ +S
Sbjct: 494 VPDPDFYDFDKDRTERTFDNDQVWATYDSEDGMPRLYAMVQKVISRKPFRIRMSFLNSKS 553
Query: 432 NADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWA 491
N + ++ GF CG F+V R +VNIFSH V + R + KI P+KG WA
Sbjct: 554 NIELSPINWVASGFSKTCGDFRVGRYQIFETVNIFSHRVSWSKGPRGIIKIVPKKGDTWA 613
Query: 492 LYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSH 551
LY + D + D+ +IV + ++ + G+++ L KV G+K VF R+ T S
Sbjct: 614 LYRNWSSDWN-ELTPDDVIYKYEIVEVIDDFTDEQGVTVIPLLKVAGFKAVFHRR-TDSD 671
Query: 552 AIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLLTI 601
+R + K+ ++ SH++P+R +E + C ELDPA+ P LL +
Sbjct: 672 VVRRIPKEELFRFSHRVPSRLLTG-EEGNNAPKGCHELDPAATPVDLLKV 720
Score = 120 bits (302), Expect = 9e-26
Identities = 75/218 (34%), Positives = 110/218 (50%), Gaps = 58/218 (26%)
Query: 8 EEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT------- 60
+EAL+ + AE K + + K A + A +AQ L P L+GIS+MV++L +L A+
Sbjct: 12 DEALKARDAAERKFH-ARDVKGARRSAIKAQNLCPSLDGISQMVSTLEVLLASESKVDGE 70
Query: 61 -DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
DWY +L + A+ V+KQY+KL+L LHPDKN + +E AFKL+ EA+ VLSD +
Sbjct: 71 NDWYRILSLSASADEEEVKKQYRKLALQLHPDKNK---SVGAEGAFKLISEAWAVLSDKS 127
Query: 120 LKKGYD----------------------------------------------AELRKKEA 133
K YD A R
Sbjct: 128 RKMQYDQKRKDHPVTNGANGLYTYDKKAHKRARKNAAASAAAAAAAAAAAAEATTRPVGL 187
Query: 134 PTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVE 171
TFWT+C+ CR+ +++ R Y+ H+L+CPNC+ +F AVE
Sbjct: 188 DTFWTSCNRCRMQYEYLRIYLNHNLLCPNCHHAFMAVE 225
>UniRef100_Q8RYF9 Heat shock protein-like [Oryza sativa]
Length = 744
Score = 150 bits (378), Expect = 1e-34
Identities = 138/469 (29%), Positives = 215/469 (45%), Gaps = 69/469 (14%)
Query: 171 EAVLSDGSSDEGEKVGVRRN--------EKREVVLGVAAPNGVKGKVGGEKGLKGKV--- 219
+A S GSS KV RR EKR + K KV G+K KV
Sbjct: 300 QASSSLGSSSSAAKVIHRRKAVTKEMEAEKRRCINN-------KSKVSGQKNNTNKVVGK 352
Query: 220 -----GDGDGVGDVGGGGSKKRMRSVGD--VFERSKPKGIITGEETMTLAEFQSQVKRKV 272
DGD G +K S+G R P +G + + Q++ ++
Sbjct: 353 STSSAADGDS-GPQMHPAKRKSASSIGTSGTKRRKMPSDHNSGNARTSFGKVFLQLETEI 411
Query: 273 ---QQEKVK----DKEEEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRGS 325
+ EK+K DK EE+ +S R VENK + + + K+ A
Sbjct: 412 PGLKMEKMKLQIRDKLEEF-----KSRRANVENKGNVHVSLEKKKTWKWKKPATLF---- 462
Query: 326 EVGEERSLKKNVKLAIKEKPEASGKRKRLE-----LEECGDANGGELEVMAVVDSDFYDF 380
V R+ K++ K + A K L+ L++ ++ G VM V ++DFY F
Sbjct: 463 -VYTRRNRKEHRKEPGVDAIGAGSSHKHLDGKYSCLDQVPSSDEGSC-VMPVPEADFYTF 520
Query: 381 DKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNADGKIVSR 440
D E SF+ GQ+WAAYD++DGMPR+YALI + +S +PF+VR+++L + ++ +
Sbjct: 521 G-DHPETSFQNGQIWAAYDEEDGMPRYYALIQKVLSRHPFKVRLAFLKAKDCSEFVTSNW 579
Query: 441 EKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWALYGEATLDA 500
G+ CG F V + + +N FSHVV ++ + +I+PRKG +WALY
Sbjct: 580 ISYGYSKTCGDFIVGTPKNTDQLNTFSHVVTWEKGPGGIIRIFPRKGDIWALY------- 632
Query: 501 DGRHFADEGKRCN--------DIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHA 552
++++ E C D+V L +YN G+S+ + KV G+ +VF + +
Sbjct: 633 --QNWSPEWNTCTPDDTIYKYDLVQVLDSYNPSAGISVMPIVKVPGFVSVFTPLLDPTKS 690
Query: 553 IRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLLTI 601
R + K+ M SHQ+P +E + C+ELDP S P LL +
Sbjct: 691 -RTIPKEEMLRFSHQVPFHVLTG-EEAKNSPKGCYELDPGSTPKELLQV 737
Score = 95.1 bits (235), Expect = 5e-18
Identities = 63/210 (30%), Positives = 94/210 (44%), Gaps = 50/210 (23%)
Query: 8 EEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTIL--------SA 59
++A+R K +AE K N+ A ++A +A+ LF LEGI M+++L I
Sbjct: 6 DDAIRSKEIAERKFN-ENDIAGAKRFALKAKTLFDSLEGIDNMISALDIHIRAQTKIEGE 64
Query: 60 TDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
D Y +L + + ++KQY+KL+L HPDKN + +E AFKL+ +A+ VLSD
Sbjct: 65 NDLYGILDISASDDDEKIKKQYRKLALQTHPDKNK---FSGAESAFKLIQDAWDVLSDKD 121
Query: 120 LKKGYDAEL--------------------------------------RKKEAPTFWTACS 141
K+ YD + TFWT C
Sbjct: 122 KKRSYDQKRFGGSSRVYQNGFAENANATPGSTMSSMNGFFWQNSGRHPSYATDTFWTYCD 181
Query: 142 ACRLLHQFERRYVGHSLVCPNCNKSFEAVE 171
+C++ Q+ R YV +L C C F AVE
Sbjct: 182 SCQMSFQYSREYVNRNLACSFCQTEFVAVE 211
>UniRef100_Q5ZCV2 DNAJ heat shock N-terminal domain-containing protein-like [Oryza
sativa]
Length = 1008
Score = 127 bits (318), Expect = 1e-27
Identities = 141/537 (26%), Positives = 214/537 (39%), Gaps = 85/537 (15%)
Query: 118 SALKKGYDAELRKKEA-PTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAV------ 170
S K+GY + +R A FWT C C+ +Q+ + H L C NC K F AV
Sbjct: 209 SNAKEGYGSSVRPPSAGEAFWTMCVNCKTKYQYYSNVLNHKLRCQNCKKDFRAVMLNEQD 268
Query: 171 --EAVLSDGSSDEGEKVGVRRNEKREVVLGVAAPNGVKGKVGGEK-------------GL 215
S + G+ V + E AA K V G + G
Sbjct: 269 VPSVFSSSAAKSAGQHCDVPKQEDCSTKFSSAANRDAKPMVNGGQHDEQMKNSASVRAGG 328
Query: 216 KGKVGDGDGV---------------GDVGG-GGSKKRMRSVGDVFERSKPKGII-TGEET 258
+G V + + +VG G K DV R P + T ET
Sbjct: 329 EGTVNHTESIRKGGLEFSTLHVSSAANVGSKAGGKMTSCPTPDVAGRQNPGNRVNTSAET 388
Query: 259 --MTLAEFQSQVKRKVQQEK--VKDKEEEWGRT------DKRSSRKRVENKRGLEIGEVR 308
M + + +RK + ++D + RT + SSRK+V + + + +
Sbjct: 389 GVMNIPNPRRSARRKENADASIIQDTPSKKRRTILDWFSNPDSSRKKVADDNVVR-ADGQ 447
Query: 309 TLKLPIKENAVKSKRGSEVGEERSLKKNVKLAIKEKPEASGKRKRLELEECGDANGGELE 368
+ + A ++G+ E KRK + + G
Sbjct: 448 ACEPHVSSEAHNHQKGTTSNEGNQ----------------EKRKDVAHDTNAQKKSGIPG 491
Query: 369 VMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWL- 427
+ D +F+DFD+ R F Q+WA YDD DGMPR+YA I + N F V+ +WL
Sbjct: 492 NFSYPDPEFFDFDRCRDVSMFAVDQIWALYDDRDGMPRYYARIRRIDTTN-FRVQFTWLE 550
Query: 428 -DLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDR-VAREVYKIYPR 485
D ++ + K E + CG F + + +FSH+V + R Y+IYPR
Sbjct: 551 HDAKNEEEDKWTDEE---LPVACGNFFLGKTVVSQDALMFSHIVSWVKGRKRSSYEIYPR 607
Query: 486 KGSVWALYG----EATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKT 541
KG VWALY + + DAD +H A E + V L+N+ G ++ L K+ G+ +
Sbjct: 608 KGEVWALYKGWSMQWSSDAD-KHRAYE----YEAVEILSNFTVEAGAAVGPLVKIKGFVS 662
Query: 542 VFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYL 598
+F + + + M SH IP ++ DE + ELD ASLPS L
Sbjct: 663 LFAKVKEKPSFV--IPPSEMLRFSHSIPFFRTKG-DEKVGVAGGFLELDTASLPSNL 716
Score = 92.0 bits (227), Expect = 4e-17
Identities = 76/271 (28%), Positives = 133/271 (49%), Gaps = 19/271 (7%)
Query: 340 AIKEKPEASGKRKRLELEECG----DANGGELEVMAVVDSDFYDFDKDRVERSFKKGQVW 395
A+KE GKRK LE A +S+F++F++ R F++GQ+W
Sbjct: 749 ALKENLMNEGKRKDHSLERTPVHQQSAAYSSPSTFDYPNSEFHNFEEYRSYSKFERGQIW 808
Query: 396 AAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVA 455
A Y D D P++Y + + V +PF V ++WL++ + + + E+ + CG FK+
Sbjct: 809 ALYSDLDQFPKYYGWVTK-VDTDPFRVHLTWLEVCPQLEQENMWLEQ-NIPVSCGTFKIR 866
Query: 456 R-KDSINSVNIFSHVVDCDRVA-REVYKIYPRKGSVWALYGEATLDADGRHFADEGKRCN 513
+ +++ + FSH+V+ +V + ++I+P+ G +WA+Y A G + K
Sbjct: 867 NWRIKLDTNDAFSHLVETSQVGWKRYFEIHPQVGEIWAIYNNW---APG--WVPSSKDTF 921
Query: 514 DIVV-FLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARK 572
+ + +T+ E + + L +VDGY+ VFK D+ + +N+ SH IP+ +
Sbjct: 922 EYTIGEITDCTEAS-TKVLLLTRVDGYRAVFK-PDSVRGTLEIPTNENI-RFSHLIPSFR 978
Query: 573 SPCIDETPELLEDCWELDPASLPSYLLTIGG 603
E L +ELDPAS+P L G
Sbjct: 979 --LTKENGGKLCGFYELDPASVPDTFLFRSG 1007
Score = 87.0 bits (214), Expect = 1e-15
Identities = 53/131 (40%), Positives = 78/131 (59%), Gaps = 12/131 (9%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT------ 60
+EEAL+ + +A K++ S + A + A +AQR+FPELE IS+M+T + A
Sbjct: 6 KEEALKARDIAAKKME-SKDFVGAKRIALKAQRIFPELENISQMLTVCEVHCAAEAKMNG 64
Query: 61 --DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
D+Y VL V+ A+ ++KQ++KL+ LHPDKN A +E AFKLV EA LSD
Sbjct: 65 LLDFYGVLQVDVMADEATIKKQFRKLAFSLHPDKNG---FAGAEAAFKLVAEAQSTLSDR 121
Query: 119 ALKKGYDAELR 129
++ YD + R
Sbjct: 122 TKRRAYDIKWR 132
>UniRef100_Q8GWW9 Hypothetical protein At2g05250/F5G3.15 [Arabidopsis thaliana]
Length = 200
Score = 117 bits (294), Expect = 7e-25
Identities = 68/204 (33%), Positives = 113/204 (55%), Gaps = 5/204 (2%)
Query: 404 MPRHYALIDETVSANPFEVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSV 463
MPR Y ++ E +S PF++ I++L +++ + + + GF CG F++ D ++ V
Sbjct: 1 MPRLYCVVREVLSVQPFKIDIAYLSSKTDIEFGSMKWVQYGFTKSCGHFRIRNSDIVDHV 60
Query: 464 NIFSHVVDCDRVAR-EVYKIYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNY 522
NIFSH++ + R +I+P G +WA+Y +L+ DG DE + ++V L Y
Sbjct: 61 NIFSHLLKGKKTGRGGCVRIFPTAGEIWAVYKNWSLNWDG-STPDEVRHQYEMVEILDEY 119
Query: 523 NEMNGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPEL 582
E G+ + L K++GYKTV+ R T + +++ + M SHQ+P+ D T
Sbjct: 120 TEQYGVCVTPLVKLEGYKTVYHR-STREDSKKWIPRCEMLRFSHQVPSWFLK--DATSGF 176
Query: 583 LEDCWELDPASLPSYLLTIGGIDN 606
E+CW+LDPA++P LL IG N
Sbjct: 177 PENCWDLDPAAIPEELLHIGAGTN 200
>UniRef100_Q9CAW0 Hypothetical protein T9J14.9 [Arabidopsis thaliana]
Length = 603
Score = 105 bits (261), Expect = 5e-21
Identities = 95/358 (26%), Positives = 153/358 (42%), Gaps = 56/358 (15%)
Query: 271 KVQQEKVKDKEEEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRGSEVGEE 330
++ E ++ K E +K+ +++++ EV+ L +KE + EE
Sbjct: 257 RLLNETIQGKSME---LEKKEVNFQLKHEAAARETEVKNKFLELKEKKL---------EE 304
Query: 331 RSLKKNVKLAIKEKPEASG---KRKRLELEECGDANGG---------------------- 365
R +K KEKP KR RLE E A G
Sbjct: 305 REQHLELKQRKKEKPAIRAETRKRSRLEYESPLSAEKGRYAETLIRPGKKQVQKREAHEI 364
Query: 366 ----ELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFE 421
E E D DF+DF+ SF GQVWA YD D MPR+YA I + +
Sbjct: 365 IYIDEDEPFNCPDPDFHDFNNTM--SSFAVGQVWALYDPVDDMPRYYAEIRKVLQPQ-LS 421
Query: 422 VRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVYK 481
+R++WL+ + I + CGRF+ + ++ + + +FSH + + +
Sbjct: 422 LRVTWLESLQTTEEPIPA---------CGRFEHGKSETSSHL-MFSHEM-YHTIRGQYVT 470
Query: 482 IYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKT 541
I PRKG WAL+G+ T + D V +T ++ G+ +AYL +V+G+ +
Sbjct: 471 INPRKGETWALFGDWTKTWKSHSEQQKTPYSYDFVEVVTEFDSDRGIGVAYLGRVEGFTS 530
Query: 542 VFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLL 599
V++R + D M SH++P+ K DE + +ELDPA++P L
Sbjct: 531 VYERAAQNGLVEIMISCDEMLRFSHRVPSFKMTG-DEKEGVPAGSFELDPAAVPRVYL 587
>UniRef100_Q9FG44 Similarity to DnaJ protein [Arabidopsis thaliana]
Length = 287
Score = 100 bits (250), Expect = 9e-20
Identities = 63/186 (33%), Positives = 99/186 (52%), Gaps = 29/186 (15%)
Query: 8 EEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEG----ISEMVTSLTILSATDWY 63
EEA ++ E KL + + A + A LFP L+ + ++ S + + +DWY
Sbjct: 21 EEATKI---VERKL-SEKDYVGAKNFINNAFNLFPSLDARWKTMIDVYISGSNVGESDWY 76
Query: 64 TVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSALKKG 123
VLGV+P ++ V+K YK+L+LLLHPDKN + +E AFKLV EA+ +LSD +
Sbjct: 77 GVLGVDPLSDDETVKKHYKQLALLLHPDKNKCY---GAEGAFKLVSEAWCLLSDKLQRSS 133
Query: 124 YDAELRK-----------------KEAPTFWTACSACRLLHQFERRY-VGHSLVCPNCNK 165
YD +K +++ TFWT C +C+ +F R + + +++CPNC +
Sbjct: 134 YDQRRKKSKQGKSSKPKAADSSKQRKSRTFWTMCRSCKTKGEFLRHWNLNKAILCPNCRQ 193
Query: 166 SFEAVE 171
F A E
Sbjct: 194 IFIATE 199
>UniRef100_Q84TH2 Hypothetical protein At4g19570 [Arabidopsis thaliana]
Length = 558
Score = 100 bits (249), Expect = 1e-19
Identities = 56/134 (41%), Positives = 86/134 (63%), Gaps = 12/134 (8%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTI-LSAT----- 60
+EEA R +AE KL + N+ A +YAK+A R++P L G+ +++ + + +SAT
Sbjct: 5 KEEAKRALDIAEKKL-SKNDYNRAKRYAKKAHRMYPNLVGLEQVLIMIDVYISATNKING 63
Query: 61 --DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
DWY VLGV+P A+ AV+K+Y+KL+LLLHPDKN +E AFKL+ EA+ +LSD
Sbjct: 64 EADWYRVLGVDPLADDEAVKKRYRKLALLLHPDKNR---FTGAEGAFKLILEAWDLLSDK 120
Query: 119 ALKKGYDAELRKKE 132
+ + YD + + +
Sbjct: 121 SQRSSYDQKRKSNQ 134
Score = 37.7 bits (86), Expect = 0.99
Identities = 15/41 (36%), Positives = 24/41 (57%), Gaps = 1/41 (2%)
Query: 135 TFWTACSACRLLHQFER-RYVGHSLVCPNCNKSFEAVEAVL 174
TFWT CS C+ + R ++ + CPNC++ F A E ++
Sbjct: 310 TFWTVCSRCKTHCKLVRANHLNKTFPCPNCSQEFVAAEMII 350
Score = 36.6 bits (83), Expect = 2.2
Identities = 17/46 (36%), Positives = 24/46 (51%), Gaps = 1/46 (2%)
Query: 125 DAELRKKEAPTFWTACSACRLLHQFER-RYVGHSLVCPNCNKSFEA 169
D L + TFWT C+ C+ +F R + ++ CPNC K F A
Sbjct: 201 DISLTTVKVGTFWTVCNRCKTYCEFMRASCLNKTVPCPNCGKYFIA 246
>UniRef100_O49475 Hypothetical protein F24J7.130 [Arabidopsis thaliana]
Length = 539
Score = 100 bits (249), Expect = 1e-19
Identities = 56/134 (41%), Positives = 86/134 (63%), Gaps = 12/134 (8%)
Query: 7 EEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTI-LSAT----- 60
+EEA R +AE KL + N+ A +YAK+A R++P L G+ +++ + + +SAT
Sbjct: 5 KEEAKRALDIAEKKL-SKNDYNRAKRYAKKAHRMYPNLVGLEQVLIMIDVYISATNKING 63
Query: 61 --DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDS 118
DWY VLGV+P A+ AV+K+Y+KL+LLLHPDKN +E AFKL+ EA+ +LSD
Sbjct: 64 EADWYRVLGVDPLADDEAVKKRYRKLALLLHPDKNR---FTGAEGAFKLILEAWDLLSDK 120
Query: 119 ALKKGYDAELRKKE 132
+ + YD + + +
Sbjct: 121 SQRSSYDQKRKSNQ 134
Score = 37.7 bits (86), Expect = 0.99
Identities = 15/41 (36%), Positives = 24/41 (57%), Gaps = 1/41 (2%)
Query: 135 TFWTACSACRLLHQFER-RYVGHSLVCPNCNKSFEAVEAVL 174
TFWT CS C+ + R ++ + CPNC++ F A E ++
Sbjct: 291 TFWTVCSRCKTHCKLVRANHLNKTFPCPNCSQEFVAAEMII 331
Score = 35.4 bits (80), Expect = 4.9
Identities = 17/57 (29%), Positives = 26/57 (44%), Gaps = 1/57 (1%)
Query: 125 DAELRKKEAPTFWTACSACRLLHQFER-RYVGHSLVCPNCNKSFEAVEAVLSDGSSD 180
D L + TFWT C+ C+ +F R + ++ CPNC K ++ S D
Sbjct: 201 DISLTTVKVGTFWTVCNRCKTYCEFMRASCLNKTVPCPNCGKLSPCNQSTWKKASDD 257
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.134 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,043,238,022
Number of Sequences: 2790947
Number of extensions: 47047997
Number of successful extensions: 159346
Number of sequences better than 10.0: 2942
Number of HSP's better than 10.0 without gapping: 1473
Number of HSP's successfully gapped in prelim test: 1516
Number of HSP's that attempted gapping in prelim test: 151365
Number of HSP's gapped (non-prelim): 5733
length of query: 606
length of database: 848,049,833
effective HSP length: 133
effective length of query: 473
effective length of database: 476,853,882
effective search space: 225551886186
effective search space used: 225551886186
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0230.4


