

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.5
(471 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
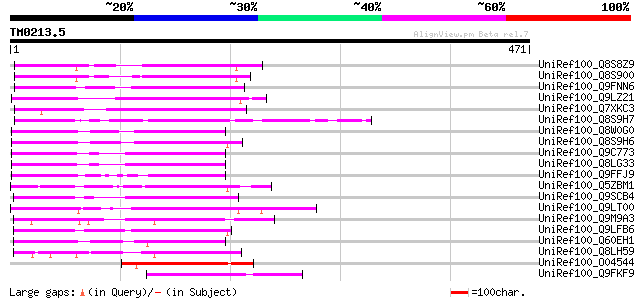
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q8S8Z9 Syringolide-induced protein 1-3-1B [Glycine max] 145 3e-33
UniRef100_Q8S900 Syringolide-induced protein 1-3-1A [Glycine max] 138 4e-31
UniRef100_Q9FNN6 Similarity to I-box binding factor [Arabidopsis... 134 6e-30
UniRef100_Q9LZ21 Putative I-box binding factor [Arabidopsis thal... 133 1e-29
UniRef100_Q7XKC3 OSJNBa0064G10.8 protein [Oryza sativa] 132 3e-29
UniRef100_Q8S9H7 MYB-like transcription factor DIVARICATA [Antir... 131 4e-29
UniRef100_Q8W0G0 P0529E05.19 protein [Oryza sativa] 124 4e-27
UniRef100_Q8S9H6 MYB-like transcription factor DVL1 [Antirrhinum... 124 8e-27
UniRef100_Q9C773 MYB-family transcription factor, putative; alte... 122 2e-26
UniRef100_Q8LG33 MYB-family transcription factor, putative [Arab... 122 2e-26
UniRef100_Q9FFJ9 Similarity to I-box binding factor [Arabidopsis... 122 3e-26
UniRef100_Q5ZBM1 Putative MYB-like transcription factor [Oryza s... 121 5e-26
UniRef100_Q9SCB4 I-box binding factor [Lycopersicon esculentum] 116 1e-24
UniRef100_Q9LT00 Similarity to I-box binding factor [Arabidopsis... 115 2e-24
UniRef100_Q9M9A3 F27J15.20 [Arabidopsis thaliana] 114 8e-24
UniRef100_Q9LFB6 Hypothetical protein F7J8_180 [Arabidopsis thal... 112 2e-23
UniRef100_Q60EH1 Hypothetical protein B1122D01.8 [Oryza sativa] 111 5e-23
UniRef100_Q8LH59 Putative syringolide-induced protein 1-3-1B [Or... 106 2e-21
UniRef100_O04544 F20P5.26 protein [Arabidopsis thaliana] 100 7e-20
UniRef100_Q9FKF9 Similarity to Myb-related transcription activat... 100 1e-19
>UniRef100_Q8S8Z9 Syringolide-induced protein 1-3-1B [Glycine max]
Length = 236
Score = 145 bits (365), Expect = 3e-33
Identities = 91/232 (39%), Positives = 122/232 (52%), Gaps = 33/232 (14%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG--PA 61
WT D+ E AL VP+D PDRWE+IA V G+ E+ E Y LV V I+ G
Sbjct: 17 WTRYHDKLFERALLVVPEDLPDRWEKIADQVPGKSAVEVREHYEALVHDVFEIDSGRVEV 76
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPN-IEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P ++ D V P PS +I+ + N I G P K
Sbjct: 77 PSYVDDSVATP----PSGGAEISTWDNANQISFGSKP----------------------K 110
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
+ N+ +K WT++EHR FL GL K+GKG W++ISR + ++TPTQ+ASHAQKY+LR
Sbjct: 111 QQGDNERKKGTPWTEEEHRLFLIGLSKFGKGDWRSISRNVVVTRTPTQVASHAQKYFLRQ 170
Query: 181 NSTPKRRKRASIHDLTIDDTDLVP---QQNQVPQLNAPMQELSPQQNGVQLD 229
NS K RKR+SIHD+T D++ VP QN VP +Q+ + LD
Sbjct: 171 NSVKKERKRSSIHDITTVDSNSVPVPIDQNWVPPPGGSVQQSLQYPSSNMLD 222
>UniRef100_Q8S900 Syringolide-induced protein 1-3-1A [Glycine max]
Length = 233
Score = 138 bits (347), Expect = 4e-31
Identities = 86/220 (39%), Positives = 116/220 (52%), Gaps = 31/220 (14%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG--PA 61
WT D+ E AL VP+D PDRWE+IA V G+ E+ E Y LV V I+ G
Sbjct: 14 WTRYHDKLFERALLVVPEDLPDRWEKIADQVPGKSAVEVREHYEALVHDVFEIDSGRVEV 73
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKK 121
P ++ D V P PS I+ + N S K+ K+
Sbjct: 74 PSYVDDSVAMP----PSGGAGISTWDNAN--------------------QISFGSKL-KQ 108
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
N+ +K WT++EHR FL GL K+GKG W++ISR + ++TPTQ+ASHAQKY+LR N
Sbjct: 109 QGENERKKGTPWTEEEHRLFLIGLSKFGKGDWRSISRNVVVTRTPTQVASHAQKYFLRQN 168
Query: 182 STPKRRKRASIHDLTIDDTDLVP---QQNQVPQLNAPMQE 218
S K RKR+SIHD+T D++ P Q VP +Q+
Sbjct: 169 SVKKERKRSSIHDITTVDSNSAPMPIDQTWVPPPGGSLQQ 208
>UniRef100_Q9FNN6 Similarity to I-box binding factor [Arabidopsis thaliana]
Length = 298
Score = 134 bits (337), Expect = 6e-30
Identities = 82/211 (38%), Positives = 112/211 (52%), Gaps = 23/211 (10%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAPD 63
W+ D A E ALA D++ +RWE+IAA V G+ +I E Y LLV V IE G
Sbjct: 12 WSREDDIAFERALANNTDESEERWEKIAADVPGKSVEQIKEHYELLVEDVTRIESGC--- 68
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
VP P P S + ++G + S + K S
Sbjct: 69 -----VPLPAYGSPEGSNGHAGDEGASSKKGGN-------------SHAGESNQAGKSKS 110
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
+ K + WT+DEHR FL GL KYGKG W++ISR F+ ++TPTQ+ASHAQKY++RLNS
Sbjct: 111 DQERRKGIAWTEDEHRLFLLGLDKYGKGDWRSISRNFVVTRTPTQVASHAQKYFIRLNSM 170
Query: 184 PKRRKRASIHDLT-IDDTDLVPQQNQVPQLN 213
K R+R+SIHD+T + + D+ Q + N
Sbjct: 171 NKDRRRSSIHDITSVGNADVSTPQGPITGQN 201
>UniRef100_Q9LZ21 Putative I-box binding factor [Arabidopsis thaliana]
Length = 215
Score = 133 bits (334), Expect = 1e-29
Identities = 79/235 (33%), Positives = 118/235 (49%), Gaps = 38/235 (16%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAVGRLPAEIMERYRLLVLHVDNIEEGPA 61
S WT ++D+ E AL P+ +P+RWE+IA + + E+ E Y +LV V I+ G
Sbjct: 3 SSQWTRSEDKMFEQALVLFPEGSPNRWERIADQLHKSAGEVREHYEVLVHDVFEIDSGRV 62
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKK 121
PD++ D D + K
Sbjct: 63 D---------------------------------VPDYMDDSAAAAAGWDSAGQISFGSK 89
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
++ ++ WT++EH+ FL GLK+YGKG W++ISR + ++TPTQ+ASHAQKY+LR N
Sbjct: 90 HGESERKRGTPWTENEHKLFLIGLKRYGKGDWRSISRNVVVTRTPTQVASHAQKYFLRQN 149
Query: 182 STPKRRKRASIHDL-TIDDTDLVPQQNQ--VPQLNAPMQELSPQQNGVQLDAPMQ 233
S K RKR+SIHD+ T+D T +P N Q +P+Q +PQQ + + Q
Sbjct: 150 SVKKERKRSSIHDITTVDATLAMPGSNMDWTGQHGSPVQ--APQQQQIMSEFGQQ 202
>UniRef100_Q7XKC3 OSJNBa0064G10.8 protein [Oryza sativa]
Length = 318
Score = 132 bits (331), Expect = 3e-29
Identities = 84/217 (38%), Positives = 115/217 (52%), Gaps = 25/217 (11%)
Query: 5 WTPAQDRALELALATVPDDAPDR---WEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
WT +++A E ALATV DD + WE++A AV G+ E+ Y LLV VD IE G
Sbjct: 33 WTREREKAFENALATVGDDEEEGDGLWEKLAEAVEGKTADEVRRHYELLVEDVDGIEAGR 92
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P + +EEG A +K +
Sbjct: 93 VPLLVYAG-------------------DGGVEEGSAGGGKKGGGGGGGGGGGGHGEKGSA 133
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
K S + K + WT+DEHR FL GL+KYGKG W++ISR F+ S+TPTQ+ASHAQKY++RL
Sbjct: 134 KSSEQERRKGIAWTEDEHRLFLLGLEKYGKGDWRSISRNFVISRTPTQVASHAQKYFIRL 193
Query: 181 NSTPKRRKRASIHDLT-IDDTDLVPQQNQVP-QLNAP 215
NS + R+R+SIHD+T +++ D Q + Q N P
Sbjct: 194 NSMNRERRRSSIHDITSVNNGDTSAAQGPITGQPNGP 230
>UniRef100_Q8S9H7 MYB-like transcription factor DIVARICATA [Antirrhinum majus]
Length = 307
Score = 131 bits (330), Expect = 4e-29
Identities = 106/326 (32%), Positives = 157/326 (47%), Gaps = 47/326 (14%)
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAPD 63
WT A+++A E ALA ++ P+RWE++A V G+ ++M +Y+ L V +IE G
Sbjct: 26 WTAAENKAFENALAVFDENTPNRWERVAERVPGKTVGDVMRQYKELEDDVSSIEAG---- 81
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
F+ PVP +SS +E G F + SSS + +++
Sbjct: 82 FV------PVPGYSTSSPF-------TLEWGSGHGFDGFKQSYGTGGRKSSSGRPSEQ-- 126
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
+ +K V WT++EH+ FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R S
Sbjct: 127 --ERKKGVPWTEEEHKLFLMGLKKYGKGDWRNISRNFVITRTPTQVASHAQKYFIRQLSG 184
Query: 184 PKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVD 243
K ++RASIHD+T + NQ P + SP D M Q + +
Sbjct: 185 GKDKRRASIHDITTVNL----SDNQTPSPDNKKPPSSP-------DHSMAQQQTSSTSIH 233
Query: 244 ALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQ 303
L F Q S + + H Q N P + F+ G Q+
Sbjct: 234 KLPFQWDQTSNETIMGFASSGHHGNMFQSN---PFGMNSYGFKMQG-------QQMQRGG 283
Query: 304 NCATPLNVQQLSFQ-QNGDPPLNFPH 328
C T L Q ++FQ Q+G L+FP+
Sbjct: 284 FCDTYLGSQNMAFQMQSG---LHFPN 306
>UniRef100_Q8W0G0 P0529E05.19 protein [Oryza sativa]
Length = 508
Score = 124 bits (312), Expect = 4e-27
Identities = 73/197 (37%), Positives = 111/197 (56%), Gaps = 25/197 (12%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAVGRLPA-EIMERYRLLVLHVDNIEEGP 60
+++W+ +++ E ALA V D+P+RWE +AA + R ++M YR L V +IE G
Sbjct: 29 AEAWSAEENKVFERALAQVDLDSPNRWEMVAAMLPRKTVIDVMNHYRDLENDVGSIEAGL 88
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P P+ SS + + +G F + +
Sbjct: 89 VP----------FPHYSSSLSPASSGFTLQDWDGSDGGF-------------RRGCYLKR 125
Query: 121 KISPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLR 179
+P+Q K+ V WT++EH++FL GLKKYG+G W+ ISR F+ S+TPTQ+ASHAQKY++R
Sbjct: 126 GRAPDQERKKGVPWTEEEHKSFLMGLKKYGRGDWRNISRYFVTSRTPTQVASHAQKYFIR 185
Query: 180 LNSTPKRRKRASIHDLT 196
LNS K ++R+SIHD+T
Sbjct: 186 LNSGGKDKRRSSIHDIT 202
>UniRef100_Q8S9H6 MYB-like transcription factor DVL1 [Antirrhinum majus]
Length = 291
Score = 124 bits (310), Expect = 8e-27
Identities = 76/214 (35%), Positives = 116/214 (53%), Gaps = 25/214 (11%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S WT A ++A E ALA + P RWE++A V G+ +++ Y+ L V +IE G
Sbjct: 22 SPKWTAADNKAFENALAVFDEYTPHRWERVAEIVPGKTVWDVIRHYKELEDDVTSIEAGL 81
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P VP +S + S + +G ++V +
Sbjct: 82 VP----------VPGYNTSLPFTLEWGSGHGFDGFMQSYVV-----------GGRKSSCS 120
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
+ S + +K V WT++EH+ FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R
Sbjct: 121 RPSDQERKKGVPWTEEEHKLFLMGLKKYGKGDWRNISRNFVITRTPTQVASHAQKYFIRQ 180
Query: 181 NSTPKRRKRASIHDLT---IDDTDLVPQQNQVPQ 211
S K ++RASIHD+T ++D P++N++ Q
Sbjct: 181 LSGGKDKRRASIHDITTVNLNDGQTFPRENKIKQ 214
>UniRef100_Q9C773 MYB-family transcription factor, putative; alternative splicing
isoform 2 of 2;71559-70643 [Arabidopsis thaliana]
Length = 263
Score = 122 bits (307), Expect = 2e-26
Identities = 73/196 (37%), Positives = 102/196 (51%), Gaps = 34/196 (17%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S SWT +++ E ALA +D+PDRW ++A+ + G+ ++M++Y L V +IE G
Sbjct: 30 SGSWTKEENKMFERALAIYAEDSPDRWFKVASMIPGKTVFDVMKQYSKLEEDVFDIEAGR 89
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P+P P++S P D +
Sbjct: 90 V----------PIPGYPAASS-----------------------PLGFDTDMCRKRPSGA 116
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
+ S +K V WT++EHR FL GL KYGKG W+ ISR F+ SKTPTQ+ASHAQKYY R
Sbjct: 117 RGSDQDRKKGVPWTEEEHRRFLLGLLKYGKGDWRNISRNFVVSKTPTQVASHAQKYYQRQ 176
Query: 181 NSTPKRRKRASIHDLT 196
S K ++R SIHD+T
Sbjct: 177 LSGAKDKRRPSIHDIT 192
>UniRef100_Q8LG33 MYB-family transcription factor, putative [Arabidopsis thaliana]
Length = 263
Score = 122 bits (307), Expect = 2e-26
Identities = 73/196 (37%), Positives = 102/196 (51%), Gaps = 34/196 (17%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S SWT +++ E ALA +D+PDRW ++A+ + G+ ++M++Y L V +IE G
Sbjct: 30 SGSWTKEENKMFERALAIYAEDSPDRWFKVASMIPGKTVFDVMKQYSKLEEDVFDIEAGR 89
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P+P P++S P D +
Sbjct: 90 V----------PIPGYPAASS-----------------------PLGFDTDMCRKRPSGA 116
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
+ S +K V WT++EHR FL GL KYGKG W+ ISR F+ SKTPTQ+ASHAQKYY R
Sbjct: 117 RGSDQDRKKGVPWTEEEHRRFLLGLLKYGKGDWRNISRNFVVSKTPTQVASHAQKYYQRQ 176
Query: 181 NSTPKRRKRASIHDLT 196
S K ++R SIHD+T
Sbjct: 177 LSGAKDKRRPSIHDIT 192
>UniRef100_Q9FFJ9 Similarity to I-box binding factor [Arabidopsis thaliana]
Length = 277
Score = 122 bits (305), Expect = 3e-26
Identities = 77/196 (39%), Positives = 104/196 (52%), Gaps = 28/196 (14%)
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S SWT +++ E ALA DD PDRW ++AA + G+ +++M +Y L + +IE G
Sbjct: 28 SSSWTKEENKKFERALAVYADDTPDRWFKVAAMIPGKTISDVMRQYSKLEEDLFDIEAGL 87
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P +P P ++ +SP DF R +PN +
Sbjct: 88 VP------IPGYRSVTPCGFDQV---VSPR-------DFDAYR---KLPNGARGFDQ--- 125
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
K V WT++EHR FL GL KYGKG W+ ISR F+ SKTPTQ+ASHAQKYY R
Sbjct: 126 -----DRRKGVPWTEEEHRRFLLGLLKYGKGDWRNISRNFVGSKTPTQVASHAQKYYQRQ 180
Query: 181 NSTPKRRKRASIHDLT 196
S K ++R SIHD+T
Sbjct: 181 LSGAKDKRRPSIHDIT 196
>UniRef100_Q5ZBM1 Putative MYB-like transcription factor [Oryza sativa]
Length = 287
Score = 121 bits (303), Expect = 5e-26
Identities = 88/243 (36%), Positives = 125/243 (51%), Gaps = 32/243 (13%)
Query: 1 MSDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG 59
M+ W+P +++ E ALA + PD W+++A A+ GR E+M +R L L V IE G
Sbjct: 18 MARKWSPEENKQFERALAGLDLRCPD-WDRVARAIPGRSALEVMNHFRDLELDVQQIENG 76
Query: 60 PAPDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKIT 119
VP PV ++ T + G DF R +
Sbjct: 77 --------MVPFPVYGAAAAGGAFTLQWDGAHGVG---DF---RNAYRFGGGGGGKRHFG 122
Query: 120 KKISPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYL 178
+ +P Q K+ V WT++EH+ FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++
Sbjct: 123 R--TPEQERKKGVPWTEEEHKLFLLGLKKYGKGDWRNISRNFVQTRTPTQVASHAQKYFI 180
Query: 179 RLNSTPKRRKRASIHDLT----IDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQ 234
RLNS K ++R+SIHD+T DD P Q+ + +S Q N L A +
Sbjct: 181 RLNSGGKDKRRSSIHDITTVNLTDDRPPSPSQSSL---------ISNQSNTSTLTAAVAP 231
Query: 235 VSS 237
SS
Sbjct: 232 FSS 234
>UniRef100_Q9SCB4 I-box binding factor [Lycopersicon esculentum]
Length = 191
Score = 116 bits (291), Expect = 1e-24
Identities = 75/206 (36%), Positives = 103/206 (49%), Gaps = 37/206 (17%)
Query: 4 SWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAP 62
+WT D+ E L P+++ DRW+ IA V G+ +IM Y LV V I+ G
Sbjct: 6 TWTRDDDKLFEHGLVLYPENSADRWQLIADHVPGKTADDIMAHYDDLVHDVYEIDSG--- 62
Query: 63 DFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKI 122
R+ P D P + D + KK
Sbjct: 63 -----RIDLPSYTDD---------------------------PVELEGDCQITSGSNKKS 90
Query: 123 SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
+ + +K WT+DEHR FL GL KYGKG W++ISR + S+TPTQ+ASHAQKY++R +
Sbjct: 91 NEIERKKGTPWTEDEHRLFLIGLDKYGKGDWRSISRNVVVSRTPTQVASHAQKYFIRQQA 150
Query: 183 TPKRRKRASIHDLTID-DTDLVPQQN 207
K RKR+SIHD+T DT+ VP Q+
Sbjct: 151 MKKERKRSSIHDITTAVDTNPVPPQS 176
>UniRef100_Q9LT00 Similarity to I-box binding factor [Arabidopsis thaliana]
Length = 337
Score = 115 bits (289), Expect = 2e-24
Identities = 92/290 (31%), Positives = 128/290 (43%), Gaps = 35/290 (12%)
Query: 1 MSDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG 59
+ SW+ D A E ALA D RW++IA V G+ +++E Y +L V IE G
Sbjct: 9 VGSSWSKDDDIAFEKALAIYNDKTEIRWKKIATVVPGKTLEQVIEHYNILARDVMLIESG 68
Query: 60 PA--PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQK 117
PD+ D + EP N + I +E G ND K
Sbjct: 69 CVRLPDYD-DFLEEPNHNAFGKERSI-------LEGG---------------NDRKYESK 105
Query: 118 ITKKISPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKY 176
K Q +R V W EHR FL GLKKYGKG W++ISR + ++T TQ+ASHAQKY
Sbjct: 106 HKGKSKLKQKRRRGVPWKPFEHRQFLHGLKKYGKGDWRSISRHCVVTRTSTQVASHAQKY 165
Query: 177 YLRLNSTPKRRKRASIHDLTIDDTDLVPQQ------NQVPQLNAPMQELSPQQNGVQ--L 228
+ +NS K+RKR SIHD+TI + + + ++ A Q +Q L
Sbjct: 166 FAHINSEDKKRKRPSIHDITIAENKSISTKQRPITWQKINNNGATASNTQANQTTLQPSL 225
Query: 229 DAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPL 278
D P+ + A Q D P+ + QV Q + P+
Sbjct: 226 DIPIYGTHNNIWNAQATQVISQPSQNHPTYDAPIIWNTQVASQPSANIPM 275
>UniRef100_Q9M9A3 F27J15.20 [Arabidopsis thaliana]
Length = 314
Score = 114 bits (284), Expect = 8e-24
Identities = 82/255 (32%), Positives = 123/255 (48%), Gaps = 43/255 (16%)
Query: 4 SWTPAQDRALELALA---TVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG 59
+W+ +++A E A+A + D+W ++++ V + E+ + Y++L+ V IE G
Sbjct: 7 TWSREEEKAFENAIALHCVEEEITEDQWNKMSSMVPSKALEEVKKHYQILLEDVKAIENG 66
Query: 60 PAP--------DFIVDRVP----EPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEP 107
P IVD P D SS KK P P
Sbjct: 67 QVPLPRYHHRKGLIVDEAAAAATSPANRDSHSSGSSEKK------------------PNP 108
Query: 108 VPNDPSSSQKITKKISPNQNEKR--VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKT 165
+ SSS S + E+R + WT++EHR FL GL K+GKG W++ISR F+ S+T
Sbjct: 109 GTSGISSSNGGRSGGSRAEQERRKGIPWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRT 168
Query: 166 PTQIASHAQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNG 225
PTQ+ASHAQKY++RLNS + R+R+SIHD+T NQ P + Q+ ++
Sbjct: 169 PTQVASHAQKYFIRLNSMNRDRRRSSIHDIT-------TVNNQAPAVTGGGQQPQVVKHR 221
Query: 226 VQLDAPMQQVSSQQN 240
P Q QQ+
Sbjct: 222 PAQPQPQPQPQPQQH 236
>UniRef100_Q9LFB6 Hypothetical protein F7J8_180 [Arabidopsis thaliana]
Length = 267
Score = 112 bits (280), Expect = 2e-23
Identities = 70/203 (34%), Positives = 110/203 (53%), Gaps = 17/203 (8%)
Query: 4 SWTPAQDRALELALATVPD-DAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPA 61
+WT +++ E ALA + D D + W +IA + G+ A++++RY+ L V +IE G
Sbjct: 29 TWTAEENKRFEKALAYLDDKDNLESWSKIADLIPGKTVADVIKRYKELEDDVSDIEAG-- 86
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKK 121
+P P +SS + D++V + P+ +
Sbjct: 87 ------LIPIPGYGGDASSAANSDYFFGLENSSYGYDYVVGGKR----SSPAMTDCFRSP 136
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
+ + +K V WT+DEH FL GLKKYGKG W+ I++ F+ ++TPTQ+ASHAQKY+LR
Sbjct: 137 MPEKERKKGVPWTEDEHLRFLMGLKKYGKGDWRNIAKSFVTTRTPTQVASHAQKYFLRQL 196
Query: 182 STPKRRKRASIHDLT---IDDTD 201
+ K ++R+SIHD+T I D D
Sbjct: 197 TDGKDKRRSSIHDITTVNIPDAD 219
>UniRef100_Q60EH1 Hypothetical protein B1122D01.8 [Oryza sativa]
Length = 315
Score = 111 bits (277), Expect = 5e-23
Identities = 70/197 (35%), Positives = 110/197 (55%), Gaps = 23/197 (11%)
Query: 4 SWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAP 62
+W+ +++ E ALA + + P+RWE++A + G+ A++M Y L V IE G P
Sbjct: 39 AWSQEENKVFEQALAALDRNDPERWERVALLLPGKTVADVMTHYDDLENDVCFIEAGLVP 98
Query: 63 DFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKI 122
P+ ++ + + + G P + + S + K
Sbjct: 99 ----------FPHYGAAGGGGGSGFTLDWDGGDDPAGLGFK---------RSCYMVGGKR 139
Query: 123 S--PNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLR 179
+ P+Q K+ V WT++EH+ FL GLKKYG+G W+ ISR F+ S+TPTQ+ASHAQKY++R
Sbjct: 140 ARGPDQERKKGVPWTEEEHKLFLMGLKKYGRGDWRNISRNFVTSRTPTQVASHAQKYFIR 199
Query: 180 LNSTPKRRKRASIHDLT 196
LNS K ++R+SIHD+T
Sbjct: 200 LNSGGKDKRRSSIHDIT 216
>UniRef100_Q8LH59 Putative syringolide-induced protein 1-3-1B [Oryza sativa]
Length = 306
Score = 106 bits (264), Expect = 2e-21
Identities = 79/224 (35%), Positives = 111/224 (49%), Gaps = 35/224 (15%)
Query: 4 SWTPAQDRALELALAT----------VPDDAPDRWEQIAAAV--GRLPAEIMERYRLLVL 51
+WT D+A E ALA PDD D + +AA+V R E+ Y LV
Sbjct: 17 AWTREDDKAFENALAACAAPPPADGGAPDD--DWFAALAASVPGARSAEEVRRHYEALVE 74
Query: 52 HVDNIEEG--PAPDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVP 109
V I+ G P P + + P P+ ++ +K +E D
Sbjct: 75 DVAAIDAGRVPLPRYAGEESAAP-PDGAGAAAAASKDGGHRRDERKGGGGGYDG------ 127
Query: 110 NDPSSSQKITKKISPNQNEKR--VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPT 167
K S + E+R + WT++EHR FL GL K+GKG W++ISR F+ S+TPT
Sbjct: 128 ---------GKSCSKAEQERRKGIPWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRTPT 178
Query: 168 QIASHAQKYYLRLNSTPKRRKRASIHDLT-IDDTDLVPQQNQVP 210
Q+ASHAQKY++RLNS + R+R+SIHD+T + D V Q P
Sbjct: 179 QVASHAQKYFIRLNSMNRDRRRSSIHDITSVTAGDQVAAQQGAP 222
>UniRef100_O04544 F20P5.26 protein [Arabidopsis thaliana]
Length = 261
Score = 100 bits (250), Expect = 7e-20
Identities = 51/123 (41%), Positives = 75/123 (60%), Gaps = 5/123 (4%)
Query: 102 DRVPEPVPNDPS--SSQKITKKISPNQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISR 158
D+ P+P P D +S + N+ KR WT++EHR FL GL K GKG W+ ISR
Sbjct: 66 DQTPDPNPTDDGGYASDDVVHASGRNRERKRGTPWTEEEHRLFLTGLHKVGKGDWRGISR 125
Query: 159 EFLPSKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQE 218
F+ ++TPTQ+ASHAQKY+LR + +RR+R+S+ D+T D + + QL P++
Sbjct: 126 NFVKTRTPTQVASHAQKYFLRRTNQNRRRRRSSLFDITPD--SFIGSSKEENQLQTPLEL 183
Query: 219 LSP 221
+ P
Sbjct: 184 IRP 186
>UniRef100_Q9FKF9 Similarity to Myb-related transcription activator [Arabidopsis
thaliana]
Length = 317
Score = 100 bits (248), Expect = 1e-19
Identities = 52/142 (36%), Positives = 83/142 (57%), Gaps = 6/142 (4%)
Query: 125 NQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTP 184
++ +K WT++EHRNFL GL K GKG W+ I++ F+ ++TPTQ+ASHAQKY++RLN
Sbjct: 102 HEKKKGKPWTEEEHRNFLIGLNKLGKGDWRGIAKSFVSTRTPTQVASHAQKYFIRLNVND 161
Query: 185 KRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDA 244
KR++RAS+ D++++D + +Q P P+Q + P+ Q +Q +
Sbjct: 162 KRKRRASLFDISLEDQKEKERNSQDASTKTP-----PKQPITGIQQPVVQGHTQTEISNR 216
Query: 245 L-DFPMQQLSFQQNGDPPLNFP 265
+ M+ + Q P NFP
Sbjct: 217 FQNLSMEYMPIYQPIPPYYNFP 238
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.314 0.132 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 849,061,199
Number of Sequences: 2790947
Number of extensions: 39233576
Number of successful extensions: 104934
Number of sequences better than 10.0: 1677
Number of HSP's better than 10.0 without gapping: 231
Number of HSP's successfully gapped in prelim test: 1512
Number of HSP's that attempted gapping in prelim test: 94913
Number of HSP's gapped (non-prelim): 5502
length of query: 471
length of database: 848,049,833
effective HSP length: 131
effective length of query: 340
effective length of database: 482,435,776
effective search space: 164028163840
effective search space used: 164028163840
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 77 (34.3 bits)
Lotus: description of TM0213.5


