

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0133.5
(1481 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
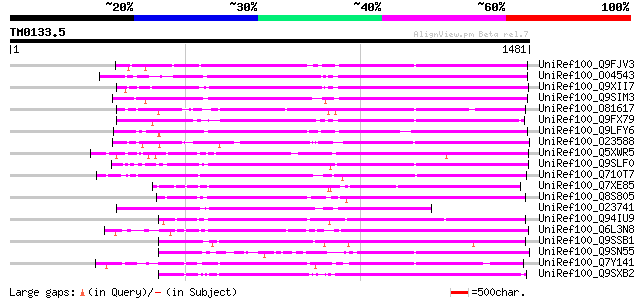
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q9FJV3 Retroelement pol polyprotein-like [Arabidopsis ... 958 0.0
UniRef100_O04543 F20P5.25 protein [Arabidopsis thaliana] 947 0.0
UniRef100_Q9XII7 Putative retroelement pol polyprotein [Arabidop... 943 0.0
UniRef100_Q9SIM3 Putative retroelement pol polyprotein [Arabidop... 929 0.0
UniRef100_O81617 F8M12.17 protein [Arabidopsis thaliana] 915 0.0
UniRef100_Q9FX79 Putative retroelement polyprotein [Arabidopsis ... 899 0.0
UniRef100_Q9LFY6 T7N9.5 [Arabidopsis thaliana] 881 0.0
UniRef100_O23588 Retrotransposon like protein [Arabidopsis thali... 872 0.0
UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Sol... 840 0.0
UniRef100_Q9SLF0 Putative retroelement pol polyprotein [Arabidop... 740 0.0
UniRef100_Q710T7 Gag-pol polyprotein [Populus deltoides] 708 0.0
UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa] 698 0.0
UniRef100_Q8S805 Putative copia-type polyprotein [Oryza sativa] 654 0.0
UniRef100_O23741 SLG-Sc and SLA-Sc genes and Melmoth retrotransp... 633 e-179
UniRef100_Q94IU9 Copia-like polyprotein [Arabidopsis thaliana] 632 e-179
UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum] 630 e-178
UniRef100_Q9SSB1 T18A20.5 protein [Arabidopsis thaliana] 615 e-174
UniRef100_Q9SN55 Putative retrotransposon polyprotein [Arabidops... 614 e-174
UniRef100_Q7Y141 Putative polyprotein [Oryza sativa] 614 e-174
UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana] 585 e-165
>UniRef100_Q9FJV3 Retroelement pol polyprotein-like [Arabidopsis thaliana]
Length = 1475
Score = 958 bits (2477), Expect = 0.0
Identities = 528/1204 (43%), Positives = 728/1204 (59%), Gaps = 68/1204 (5%)
Query: 303 KCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRS-------ANLVQTTKEDQESTSQG 355
+CS+ G GH + CYK HG+P GFK K+ K S ANL T + + S +QG
Sbjct: 309 QCSHCGYTGHNADTCYKIHGYPVGFKHKDKKTVTPSEKPKSVVANLALT--DGKVSVTQG 366
Query: 356 NTQEQEAARFGFTADQYHHLLLLLPPSEQKN-----------STSQHTASINSCVQMLPG 404
+ G + ++ ++ N ST+ S + + P
Sbjct: 367 IGPDGIVELVGSMSKSQIQDVIAYFSTQLHNPAKPITVASFASTNNDNGSTFTGISFSPS 426
Query: 405 KNGNSISQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNG-NIESA 463
S S+K L+T WI+D+GAT H+ + F S V+LP G N++ A
Sbjct: 427 TLRLLCSLTSSKKVLSLNT-WIIDSGATHHVSYDRNLFESLSDGLSNEVTLPTGSNVKIA 485
Query: 464 TIKGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIG 523
I G I+L+ + L NVL++P F NL+SV + K ++ ++ F +D C+IQD + IG
Sbjct: 486 GI-GVIKLNSNLTLKNVLYIPEFRLNLLSVSQQTKDMKCKIYFDEDCCVIQDPIKEQKIG 544
Query: 524 TARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQN-LWHYRLGHPSFVKGQS 582
+ LY+L+ S+ C SV S + V + LWH RLGHPS+ K
Sbjct: 545 RGNQIGGLYVLDTSSVE----CTSVDINSSVTEKQYCNAVVDSALWHSRLGHPSYEKNDV 600
Query: 583 IKDLFPYVQYTQDHV--CEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNG 640
+ D+ + ++ + C +C AKQK L FP N+ S+ F +IH+D WGP + + G
Sbjct: 601 LHDVLGLPKRNKEDLVHCSICQKAKQKHLSFPSKNNMSENKFDLIHIDTWGPFATPTTEG 660
Query: 641 FSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQ 700
+ YFLTIVDDYSR TWVYL+K+K++V + DF V Q+G VK VRSDN E
Sbjct: 661 YKYFLTIVDDYSRATWVYLMKAKNDVLQIFPDFLKMVETQYGTLVKAVRSDNAPELRFEA 720
Query: 701 FYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLIN 760
Y KGII + SC ETPQQNS+VERKHQHILNVARAL+F+A++P FW I+ ++FLIN
Sbjct: 721 LYQAKGIISYHSCPETPQQNSVVERKHQHILNVARALMFEANMPLEFWGDCILSAVFLIN 780
Query: 761 RLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSG 820
RLPTP+L N+ PF+LLH ++PD T LKVFG LC+ ST R KF RA+ CVFLG+ SG
Sbjct: 781 RLPTPLLSNKSPFELLHLKVPDYTSLKVFGCLCYESTSPQQRHKFAPRARACVFLGYPSG 840
Query: 821 TKGYLVYDLNTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPI 880
KGY + DL T I ISR+V+F+E +FP F +P + FD P+
Sbjct: 841 YKGYKLLDLETNTIHISRHVVFYETVFP--------------FTDKTIIPRDVFDLVDPV 886
Query: 881 PPDNPVQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKG 940
+ + + PS + P V S+R + P YLQDY C T K
Sbjct: 887 H-------ENIENPPSTSESAPKVS--SKRESRPPGYLQDYFCNAVPDVT--------KD 929
Query: 941 TSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTW 1000
+PL+ I+Y +L + A+I + EP Y++A K + W AM EI+ALE NTW
Sbjct: 930 VRYPLNAYINYTQLSEEFTAYICAVNKYPEPCTYAQAKKIKEWLDAMEIEIDALESTNTW 989
Query: 1001 LLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTT 1060
+ P K PIGCKWV+++K DG+++R+KARLV KGYTQ EG+D+ DTFSPVAKMTT
Sbjct: 990 SVCSLPQGKKPIGCKWVFKVKLNADGSLERFKARLVAKGYTQREGLDYYDTFSPVAKMTT 1049
Query: 1061 LRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK-----PNQVCLLQKS 1115
++ LLS+ + K W LHQLD+ NAFL+ L EEIYM+LP G + + N V LQKS
Sbjct: 1050 VKTLLSVAAIKEWSLHQLDISNAFLNGDLKEEIYMTLPPGYSMKQGGVLPQNPVLKLQKS 1109
Query: 1116 LYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDM 1175
LYGLKQASRQW+ L+ LGF +S ADHTL+ + S ++ ALL+YVDD+++AGN+
Sbjct: 1110 LYGLKQASRQWYLKFSSTLKKLGFKKSHADHTLFTRISGK-AYIALLVYVDDIVIAGNND 1168
Query: 1176 DEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSA 1235
+ I+ +K L + F+++DLG K+FLGL IAR+++GI + QRKY +ELL D+GLLG + +
Sbjct: 1169 ENIEELKKDLAKAFKLRDLGPMKYFLGLEIARTKEGISVCQRKYTMELLEDTGLLGCRPS 1228
Query: 1236 TTPMDCSQKLSASSGTPLSDISS-YRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMH 1294
T PM+ S KLS + + D YRRL+G+L+YLT TRPDI Y +N+L QF S+P + H
Sbjct: 1229 TIPMEPSLKLSQHNDEHVIDNPEVYRRLVGKLMYLTITRPDITYAINRLCQFSSSPKNSH 1288
Query: 1295 EAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVS 1354
AA +V+ Y+KG+ G GLFY + S L A++D+D CVD+R+S +G CMFLG SL+S
Sbjct: 1289 LKAAQKVVHYLKGTIGLGLFYSSKSDLCLKAYTDADWGSCVDSRRSTSGICMFLGDSLIS 1348
Query: 1355 WRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAH 1414
W+SKKQ S SS E+EYRAMA E+ WL+ LL + QV QT PV +FCD+ +A+HIA+
Sbjct: 1349 WKSKKQNMASSSSAESEYRAMAMGSREIAWLVKLLAEFQVKQTKPVPLFCDSTAAIHIAN 1408
Query: 1415 NPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRL 1474
N +HERTKHIE DCHI R++I QG++ + V ++ QLAD+ TKPL P F + K+ L
Sbjct: 1409 NAVFHERTKHIENDCHITRDRIEQGMLKTMHVDTTSQLADVLTKPLFPTLFNSLIGKMSL 1468
Query: 1475 YDIH 1478
I+
Sbjct: 1469 LSIY 1472
>UniRef100_O04543 F20P5.25 protein [Arabidopsis thaliana]
Length = 1315
Score = 947 bits (2447), Expect = 0.0
Identities = 525/1245 (42%), Positives = 736/1245 (58%), Gaps = 114/1245 (9%)
Query: 257 PLIAARENSNDGRSSQSNSGYQSNSGYQSGS---GNYHSNSGRN--RYSNKK---CSYYG 308
P ++ N D +Q ++ + G S N S S N Y K+ CSY
Sbjct: 157 PSLSEVFNMIDQDETQRSARISTTPGMTSSVFPVSNQSSQSALNGDTYQKKERPVCSYCS 216
Query: 309 KMGHTVEDCYKKHGFPPGFKFKNPKYA--NRSANLVQTTKEDQESTSQGNTQEQEAARFG 366
+ GH + CYKKHG+P FK K K+ + SAN ++E +TS +
Sbjct: 217 RPGHVEDTCYKKHGYPTSFKSKQ-KFVKPSISANAAIGSEEVVNNTS--------VSTGD 267
Query: 367 FTADQYHHLLLLLPPSEQKNST----SQHTASINSCVQMLPGKNGNSISQISTKNGNPLD 422
T Q L+ L Q ST H+ S++S
Sbjct: 268 LTTSQIQQLVSFLSSKLQPPSTPVQPEVHSISVSS------------------------- 302
Query: 423 TTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLF 482
D ++ +C I GS+ L IL +VLF
Sbjct: 303 -----DPSSSSTVC---------------------------PISGSVHLGRHLILNDVLF 330
Query: 483 LPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPF 542
+P F+FNL+SV L KS+ R+ F + C++QD+ M+G + V +LYI++ DSLS
Sbjct: 331 IPQFKFNLLSVSSLTKSMGCRIWFDETSCVLQDATRELMVGMGKQVANLYIVDLDSLSHP 390
Query: 543 PSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDL--FPYVQYTQDHVCEV 600
+ +S++ S + +LWH RLGHPS K Q + L FP + D C V
Sbjct: 391 GTDSSITVA---------SVTSHDLWHKRLGHPSVQKLQPMSSLLSFPKQKNNTDFHCRV 441
Query: 601 CPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLL 660
C I+KQK L F N+ S F +IH+D WGP SV + +G+ YFLTIVDDYSR TWVYLL
Sbjct: 442 CHISKQKHLPFVSHNNKSSRPFDLIHIDTWGPFSVQTHDGYRYFLTIVDDYSRATWVYLL 501
Query: 661 KSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTSCVETPQQN 720
++K +V T++ F V NQF ++K VRSDN E +QFY KGI+ + SC ETPQQN
Sbjct: 502 RNKSDVLTVIPTFVTMVENQFETTIKGVRSDNAPELNFTQFYHSKGIVPYHSCPETPQQN 561
Query: 721 SIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQL 780
S+VERKHQHILNVAR+L FQ+H+P +W I+ +++LINRLP PIL+++CPF++L K +
Sbjct: 562 SVVERKHQHILNVARSLFFQSHIPISYWGDCILTAVYLINRLPAPILEDKCPFEVLTKTV 621
Query: 781 PDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRNV 840
P +KVFG LC+AST R KF RAK C F+G+ SG KGY + DL T I +SR+V
Sbjct: 622 PTYDHIKVFGCLCYASTSPKDRHKFSPRAKACAFIGYPSGFKGYKLLDLETHSIIVSRHV 681
Query: 841 IFHENIFPYKLHRDDHECESSTFPQI---PCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQ 897
+FHE +FP+ L D + E + FP + P + + D+ P + V+I P S+ P+
Sbjct: 682 VFHEELFPF-LGSDLSQEEQNFFPDLNPTPPMQRQSSDHVNPSDSSSSVEILP-SANPTN 739
Query: 898 NSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDPS 957
N +P V + S R K+P YLQDY+C +S T H + + +SY +++
Sbjct: 740 NVPEPSV-QTSHRKAKKPAYLQDYYC-----------HSVVSSTPHEIRKFLSYDRINDP 787
Query: 958 YHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWV 1017
Y F+ + T EP+ Y+EA K + WR AM E + LE +TW + P DK IGC+W+
Sbjct: 788 YLTFLACLDKTKEPSNYTEAEKLQVWRDAMGAEFDFLEGTHTWEVCSLPADKRCIGCRWI 847
Query: 1018 YRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQ 1077
++IKY DG+++RYKARLV +GYTQ EGID+ +TFSPVAK+ ++++LL + + L Q
Sbjct: 848 FKIKYNSDGSVERYKARLVAQGYTQKEGIDYNETFSPVAKLNSVKLLLGVAARFKLSLTQ 907
Query: 1078 LDVDNAFLHAKLDEEIYMSLPQGMNSDK-----PNQVCLLQKSLYGLKQASRQWFSTLCQ 1132
LD+ NAFL+ LDEEIYM LPQG S + PN VC L+KSLYGLKQASRQW+
Sbjct: 908 LDISNAFLNGDLDEEIYMRLPQGYASRQGDSLPPNAVCRLKKSLYGLKQASRQWYLKFSS 967
Query: 1133 ALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIK 1192
L GLGF QS+ DHT ++K S G F +L+Y+DD+++A N+ + ++K + F+++
Sbjct: 968 TLLGLGFIQSYCDHTCFLKIS-DGIFLCVLVYIDDIIIASNNDAAVDILKSQMKSFFKLR 1026
Query: 1193 DLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTP 1252
DLGE K+FLGL I RS KGI ++QRKYAL+LL ++G LG K ++ PMD S + SG
Sbjct: 1027 DLGELKYFLGLEIVRSDKGIHISQRKYALDLLDETGQLGCKPSSIPMDPSMVFAHDSGGD 1086
Query: 1253 LSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCG 1312
++ YRRLIGRL+YL TRPDI + VN+L+QF AP H A +++L+YIKG+ G G
Sbjct: 1087 FVEVGPYRRLIGRLMYLNITRPDITFAVNKLAQFSMAPRKAHLQAVYKILQYIKGTIGQG 1146
Query: 1313 LFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEY 1372
LFY A S L ++++D C D+R+S +GYCMFLG SL+ W+S+KQ S+SS EAEY
Sbjct: 1147 LFYSATSELQLKVYANADYNSCRDSRRSTSGYCMFLGDSLICWKSRKQDVVSKSSAEAEY 1206
Query: 1373 RAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIV 1432
R+++ E+ WL L++LQV + P +FCDN++A+HIA+N +HERTKHIE DCH V
Sbjct: 1207 RSLSVATDELVWLTNFLKELQVPLSKPTLLFCDNEAAIHIANNHVFHERTKHIESDCHSV 1266
Query: 1433 REKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDI 1477
RE++ +GL L + + LQ+AD FTKPL P+ F + SK+ L +I
Sbjct: 1267 RERLLKGLFELYHINTELQIADPFTKPLYPSHFHRLISKMGLLNI 1311
>UniRef100_Q9XII7 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1454
Score = 943 bits (2438), Expect = 0.0
Identities = 513/1192 (43%), Positives = 711/1192 (59%), Gaps = 65/1192 (5%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPPGF--------KFKNPKYANRSANLVQTTKEDQESTSQ- 354
CS+Y ++GH E CYKKHGFPPGF K + PK +AN+ ++++ + S
Sbjct: 309 CSFYNRVGHIAERCYKKHGFPPGFTPKGKAGEKLQKPKPL--AANVAESSEVNTSLESMV 366
Query: 355 GNTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQIS 414
GN +++ +F + L PPS +++ + ++ C + I ++
Sbjct: 367 GNLSKEQLQQF---IAMFSSQLQNTPPSTYATASTSQSDNLGICFSP---STYSFIGILT 420
Query: 415 TKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPS 474
TW++D+GAT H+ + S FSS V+LP G + G+++L+
Sbjct: 421 VARHTLSSATWVIDSGATHHVSHDRSLFSSLDTSVLSAVNLPTGPTVKISGVGTLKLNDD 480
Query: 475 FILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYIL 534
+L NVLF+P F NLIS+ L + R+ F + C IQD +M+G R V +LY+L
Sbjct: 481 ILLKNVLFIPEFRLNLISISSLTDDIGSRVIFDKNSCEIQDLIKGRMLGQGRRVANLYLL 540
Query: 535 NNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQYTQ 594
+ S S N V + S +WH RLGH S + +I D ++
Sbjct: 541 DVGDQSI-----------SVNAVVDIS-----MWHRRLGHASLQRLDAISDSLGTTRHKN 584
Query: 595 --DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYS 652
C VC +AKQ++L FP SN IF ++H+D+WGP SV ++ G+ YFLTIVDD+S
Sbjct: 585 KGSDFCHVCHLAKQRKLSFPTSNKVCKEIFDLLHIDVWGPFSVETVEGYKYFLTIVDDHS 644
Query: 653 RYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTS 712
R TW+YLLK+K EV T+ F V NQ+ V VK VRSDN E + FYAEKGI+ S
Sbjct: 645 RATWMYLLKTKSEVLTVFPAFIQQVENQYKVKVKAVRSDNAPELKFTSFYAEKGIVSFHS 704
Query: 713 CVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCP 772
C ETP+QNS+VERKHQHILNVARAL+FQ+ +P W ++ ++FLINR P+ +L N+ P
Sbjct: 705 CPETPEQNSVVERKHQHILNVARALMFQSQVPLSLWGDCVLTAVFLINRTPSQLLMNKTP 764
Query: 773 FQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTR 832
+++L P L+ FG LC++ST R KF R++ C+FLG+ SG KGY + DL +
Sbjct: 765 YEILTGTAPVYEQLRTFGCLCYSSTSPKQRHKFQPRSRACLFLGYPSGYKGYKLMDLESN 824
Query: 833 DISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLS 892
+ ISRNV FHE +FP + P +P D + P P QI L
Sbjct: 825 TVFISRNVQFHEEVFPLAKNPGSESSLKLFTPMVPVSSGIISDTTHS-PSSLPSQISDL- 882
Query: 893 SGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYH 952
P Q S SQR RK P +L DYHC N+ +P+S ISY
Sbjct: 883 --PPQIS--------SQRVRKPPAHLNDYHC-----------NTMQSDHKYPISSTISYS 921
Query: 953 KLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPI 1012
K+ PS+ +I NIT PT Y+EA + W A++ EI A+E+ NTW + P K +
Sbjct: 922 KISPSHMCYINNITKIPIPTNYAEAQDTKEWCEAVDAEIGAMEKTNTWEITTLPKGKKAV 981
Query: 1013 GCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKN 1072
GCKWV+ +K+ DG ++RYKARLV KGYTQ EG+D+ DTFSPVAKMTT+++LL + ++K
Sbjct: 982 GCKWVFTLKFLADGNLERYKARLVAKGYTQKEGLDYTDTFSPVAKMTTIKLLLKVSASKK 1041
Query: 1073 WFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK-----PNQVCLLQKSLYGLKQASRQWF 1127
WFL QLDV NAFL+ +L+EEI+M +P+G K N V L++S+YGLKQASRQWF
Sbjct: 1042 WFLKQLDVSNAFLNGELEEEIFMKIPEGYAERKGIVLPSNVVLRLKRSIYGLKQASRQWF 1101
Query: 1128 STLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQ 1187
+L LGF ++ DHTL++ K G F +L+YVDD+++A + L Q
Sbjct: 1102 KKFSSSLLSLGFKKTHGDHTLFL-KMYDGEFVIVLVYVDDIVIASTSEAAAAQLTEELDQ 1160
Query: 1188 KFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSA 1247
+F+++DLG+ K+FLGL +AR+ GI + QRKYALELL +G+L K + PM + K+
Sbjct: 1161 RFKLRDLGDLKYFLGLEVARTTAGISICQRKYALELLQSTGMLACKPVSVPMIPNLKMRK 1220
Query: 1248 SSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKG 1307
G + DI YRR++G+L+YLT TRPDI + VN+L QF SAP H AA+RVL+YIKG
Sbjct: 1221 DDGDLIEDIEQYRRIVGKLMYLTITRPDITFAVNKLCQFSSAPRTTHLTAAYRVLQYIKG 1280
Query: 1308 SPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSS 1367
+ G GLFY A+S L F+DSD A C D+R+S T + MF+G SL+SWRSKKQ T SRSS
Sbjct: 1281 TVGQGLFYSASSDLTLKGFADSDWASCQDSRRSTTSFTMFVGDSLISWRSKKQHTVSRSS 1340
Query: 1368 CEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIEL 1427
EAEYRA+A CE+ WL LL LQ S P+ ++ D+ +A++IA NP +HERTKHI+L
Sbjct: 1341 AEAEYRALALATCEMVWLFTLLVSLQASPPVPI-LYSDSTAAIYIATNPVFHERTKHIKL 1399
Query: 1428 DCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHS 1479
DCH VRE++ G + LL V + Q+ADI TKPL P F H+ SK+ + +I S
Sbjct: 1400 DCHTVRERLDNGELKLLHVRTEDQVADILTKPLFPYQFEHLKSKMSILNIFS 1451
>UniRef100_Q9SIM3 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1461
Score = 929 bits (2400), Expect = 0.0
Identities = 510/1210 (42%), Positives = 715/1210 (58%), Gaps = 83/1210 (6%)
Query: 294 SGRNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFK---NPKYANRSANLVQTTKEDQE 350
SG N+ CS+ ++GH E CYKKHGFPPGF K + K A Q T +
Sbjct: 304 SGPNK-GRPTCSFCNRVGHIAERCYKKHGFPPGFTPKGKSSDKPPKPQAVAAQVTLSPDK 362
Query: 351 STSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQ------KNSTSQHTASINSCVQ---- 400
T Q E F+ DQ +L+ L Q + ++SQH AS + V
Sbjct: 363 MTGQ-----LETLAGNFSPDQIQNLIALFSSQLQPQIVSPQTASSQHEASSSQSVAPSGI 417
Query: 401 MLPGKNGNSISQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNI 460
+ I ++ + + TW++D+GAT H+ + F + V+LP G
Sbjct: 418 LFSPSTYCFIGILAVSHNSLSSDTWVIDSGATHHVSHDRKLFQTLDTSIVSFVNLPTGPN 477
Query: 461 ESATIKGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACK 520
+ G++ ++ IL NVLF+P F NLIS+ L L R+ F C IQD
Sbjct: 478 VRISGVGTVLINKDIILQNVLFIPEFRLNLISISSLTTDLGTRVIFDPSCCQIQDLTKGL 537
Query: 521 MIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKG 580
+G + + +LY+L D+ SP S N+V ++WH RLGHPSF +
Sbjct: 538 TLGEGKRIGNLYVL--DTQSPAISVNAVVDV--------------SVWHKRLGHPSFSRL 581
Query: 581 QSIKDLFPYVQYT--QDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSL 638
S+ ++ ++ + C VC +AKQK+L FP +N+ + F+++H+D+WGP SV ++
Sbjct: 582 DSLSEVLGTTRHKNKKSAYCHVCHLAKQKKLSFPSANNICNSTFELLHIDVWGPFSVETV 641
Query: 639 NGFSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVL 698
G+ YFLTIVDD+SR TW+YLLKSK +V T+ F V NQ+ VK VRSDN KE
Sbjct: 642 EGYKYFLTIVDDHSRATWIYLLKSKSDVLTVFPAFIDLVENQYDTRVKSVRSDNAKELAF 701
Query: 699 SQFYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFL 758
++FY KGI+ SC ETP+QNS+VERKHQHILNVARAL+FQ+++ +W ++ ++FL
Sbjct: 702 TEFYKAKGIVSFHSCPETPEQNSVVERKHQHILNVARALMFQSNMSLPYWGDCVLTAVFL 761
Query: 759 INRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFK 818
INR P+ +L N+ PF++L +LPD + LK FG LC++ST + R KF R++ CVFLG+
Sbjct: 762 INRTPSALLSNKTPFEVLTGKLPDYSQLKTFGCLCYSSTSSKQRHKFLPRSRACVFLGYP 821
Query: 819 SGTKGYLVYDLNTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEY 878
G KGY + DL + + ISRNV FHE +FP + S F P D
Sbjct: 822 FGFKGYKLLDLESNVVHISRNVEFHEELFPLASSQQSATTASDVF--------TPMD--- 870
Query: 879 PIPPDNPVQIQPLSSGPSQNSE------QPIVHRVSQRPRKQPTYLQDYHCTLAATSTVV 932
PLSSG S S P +R K P +LQDYHC
Sbjct: 871 -----------PLSSGNSITSHLPSPQISPSTQISKRRITKFPAHLQDYHCYFV------ 913
Query: 933 PANSSSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIE 992
+K SHP+S +SY ++ PS+ +I NI+ P Y EA + W A++ EI
Sbjct: 914 -----NKDDSHPISSSLSYSQISPSHMLYINNISKIPIPQSYHEAKDSKEWCGAIDQEIG 968
Query: 993 ALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTF 1052
A+ER +TW + PP K +GCKWV+ +K+ DG+++R+KAR+V KGYTQ EG+D+ +TF
Sbjct: 969 AMERTDTWEITSLPPGKKAVGCKWVFTVKFHADGSLERFKARIVAKGYTQKEGLDYTETF 1028
Query: 1053 SPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK-----PN 1107
SPVAKM T+++LL + ++K W+L+QLD+ NAFL+ L+E IYM LP G K PN
Sbjct: 1029 SPVAKMATVKLLLKVSASKKWYLNQLDISNAFLNGDLEETIYMKLPDGYADIKGTSLPPN 1088
Query: 1108 QVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDD 1167
VC L+KS+YGLKQASRQWF +L LGF + DHTL+V + F LL+YVDD
Sbjct: 1089 VVCRLKKSIYGLKQASRQWFLKFSNSLLALGFEKQHGDHTLFV-RCIGSEFIVLLVYVDD 1147
Query: 1168 VLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDS 1227
+++A + + +L F++++LG K+FLGL +AR+ +GI L+QRKYALELL+ +
Sbjct: 1148 IVIASTTEQAAQSLTEALKASFKLRELGPLKYFLGLEVARTSEGISLSQRKYALELLTSA 1207
Query: 1228 GLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFL 1287
+L K ++ PM + +LS + G L D YRRL+G+L+YLT TRPDI + VN+L QF
Sbjct: 1208 DMLDCKPSSIPMTPNIRLSKNDGLLLEDKEMYRRLVGKLMYLTITRPDITFAVNKLCQFS 1267
Query: 1288 SAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMF 1347
SAP H AA ++VL+YIKG+ G GLFY A L ++D+D C D+R+S TG+ MF
Sbjct: 1268 SAPRTAHLAAVYKVLQYIKGTVGQGLFYSAEDDLTLKGYTDADWGTCPDSRRSTTGFTMF 1327
Query: 1348 LGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQ 1407
+GSSL+SWRSKKQ T SRSS EAEYRA+A CE+ WL LL L+V P+ ++ D+
Sbjct: 1328 VGSSLISWRSKKQPTVSRSSAEAEYRALALASCEMAWLSTLLLALRVHSGVPI-LYSDST 1386
Query: 1408 SAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRH 1467
+A++IA NP +HERTKHIE+DCH VREK+ G + LL V + Q+ADI TKPL P F H
Sbjct: 1387 AAVYIATNPVFHERTKHIEIDCHTVREKLDNGQLKLLHVKTKDQVADILTKPLFPYQFAH 1446
Query: 1468 IFSKLRLYDI 1477
+ SK+ + +I
Sbjct: 1447 LLSKMSIQNI 1456
>UniRef100_O81617 F8M12.17 protein [Arabidopsis thaliana]
Length = 1633
Score = 915 bits (2366), Expect = 0.0
Identities = 517/1232 (41%), Positives = 713/1232 (56%), Gaps = 131/1232 (10%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQGNTQEQEAA 363
C+ GK+GHTV+ CYK G+PPG+K + R + + SQ Q+
Sbjct: 265 CTNCGKVGHTVQKCYKIIGYPPGYKAAT---SYRQPQIQTQPRMQMPQQSQPRMQQP--- 318
Query: 364 RFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQIST----KNGN 419
Q++ + + P+ TS TA+I M +I ST +N N
Sbjct: 319 -IQHLISQFNAQVRVQEPAATSIYTSSPTATITEHGLMAQTSTSGTIPFPSTSLKYENNN 377
Query: 420 PL--------------DTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATI 465
WI+D+GA+ H+C+ L+ F HV+ + V+LPNG + T
Sbjct: 378 LTFQNHTLSSLQNVLSSDAWIIDSGASSHVCSDLTMFRELIHVSGVTVTLPNGTRVAITH 437
Query: 466 KGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTA 525
G+I ++ + IL NVL +P+F+FNLISV C ++ + MIG
Sbjct: 438 TGTICITSTLILHNVLLVPDFKFNLISV-----------------CCLELTRGL-MIGRG 479
Query: 526 RAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKD 585
+ +LYIL S PS + ++ HPS Q +
Sbjct: 480 KTYNNLYILETQRTSFSPSLPAATS----------------------RHPSLPALQKLVS 517
Query: 586 LFPY---VQYTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFS 642
P V T H C + P+AKQKRL + N+ + F +IH+DIWGP S+ S++GF
Sbjct: 518 SIPSLKSVSSTASH-CRISPLAKQKRLAYVSHNNLASSPFDLIHLDIWGPFSIESVDGFR 576
Query: 643 YFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFY 702
YFLT+VDD +R TWVY++K+K EV + F + Q+ +K +RSDN KE ++F
Sbjct: 577 YFLTLVDDCTRTTWVYMMKNKSEVSNIFPVFVKLIFTQYNAKIKAIRSDNVKELAFTKFV 636
Query: 703 AEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRL 762
E+G+IH SC TPQQNS+VERKHQH+LN+AR+LLFQ+++P +W+ ++ + +LINRL
Sbjct: 637 KEQGMIHQFSCAYTPQQNSVVERKHQHLLNIARSLLFQSNVPLQYWSDCVLTAAYLINRL 696
Query: 763 PTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTK 822
P+P+LDN+ PF+LL K++PD T LK LC+AST R KF RA+ CVFLG+ SG K
Sbjct: 697 PSPLLDNKTPFELLLKKIPDYTLLK--SCLCYASTNVHDRNKFSPRARPCVFLGYPSGYK 754
Query: 823 GYLVYDLNTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPP 882
GY V DL + ISI+RNV+FHE FP+K + E I LP+ P + +P
Sbjct: 755 GYKVLDLESHSISITRNVVFHETKFPFKTSKFLKESVDMFPNSILPLPA-PLHFVESMPL 813
Query: 883 DNPVQ-----IQPLSSGPSQNSEQPIVHRVS-------------------QRPRKQPTYL 918
D+ ++ +S S +S P+ V+ +R K P YL
Sbjct: 814 DDDLRADDNNASTSNSASSASSIPPLPSTVNTQNTDALDIDTNSVPIARPKRNAKAPAYL 873
Query: 919 QDYHCTLA--------ATSTVVPANSSS-----KGTSHPLSQVISYHKLDPSYHAFIMNI 965
+YHC TST + SSS T +P+S ISY KL P +H++I
Sbjct: 874 SEYHCNSVPFLSSLSPTTSTSIETPSSSIPPKKITTPYPMSTAISYDKLTPLFHSYICAY 933
Query: 966 TTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQD 1025
EP +++A+K E W A N E+ ALE+N TW++ K +GCKWV+ IKY D
Sbjct: 934 NVETEPKAFTQAMKSEKWTRAANEELHALEQNKTWIVESLTEGKNVVGCKWVFTIKYNPD 993
Query: 1026 GTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFL 1085
G+I+RYKARLV +G+TQ EGID+M+TFSPVAK ++++LL L + W L Q+DV NAFL
Sbjct: 994 GSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLGLAAATGWSLTQMDVSNAFL 1053
Query: 1086 HAKLDEEIYMSLPQGMN-----SDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFT 1140
H +LDEEIYMSLPQG S VC L KSLYGLKQASRQW+ L G F
Sbjct: 1054 HGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQASRQWYKRLSSVFLGANFI 1113
Query: 1141 QSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFF 1200
QS AD+T++VK S T S +L+YVDD+++A ND ++ +K L +F+IKDLG A+FF
Sbjct: 1114 QSPADNTMFVKVSCT-SIIVVLVYVDDLMIASNDSSAVENLKELLRSEFKIKDLGPARFF 1172
Query: 1201 LGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYR 1260
LGL IARS +GI + QRKYA LL D GL G K ++ PMD + L+ GT L + +SYR
Sbjct: 1173 LGLEIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDPNLHLTKEMGTLLPNATSYR 1232
Query: 1261 RLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASS 1320
L+GRLLYL TRPDI + V+ LSQFLSAPTD+H AAH+VLRY+KG+PG
Sbjct: 1233 ELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKVLRYLKGNPG---------- 1282
Query: 1321 TVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVC 1380
D+D C D+R+S+TG+C++LG+SL++W+SKKQ+ SRSS E+EYR++A C
Sbjct: 1283 ------QDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQSVVSRSSTESEYRSLAQATC 1336
Query: 1381 EVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGL 1440
E+ WL LL+DL V+ T P +FCDN+SA+H+A NP +HERTKHIE+DCH VR++I G
Sbjct: 1337 EIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHERTKHIEIDCHTVRDQIKAGK 1396
Query: 1441 VHLLPVTSSLQLADIFTKPLSPAPFRHIFSKL 1472
+ L V + QLADI TKPL P PF + ++
Sbjct: 1397 LKTLHVPTGNQLADILTKPLHPGPFHSLLKRI 1428
>UniRef100_Q9FX79 Putative retroelement polyprotein [Arabidopsis thaliana]
Length = 1413
Score = 899 bits (2324), Expect = 0.0
Identities = 497/1189 (41%), Positives = 696/1189 (57%), Gaps = 111/1189 (9%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYAN--RSANLVQTTKEDQESTSQ--GNTQE 359
CS+YG GHT E CYK HG+P G+K Y +++ Q K + T+Q GN+Q
Sbjct: 312 CSHYGYTGHTSERCYKLHGYPVGWKKGKSFYEKIAQASQSSQAPKPNSAVTAQVTGNSQN 371
Query: 360 Q----EAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPG------KNGNS 409
E+ + DQ +L+ L Q S +TA +++ P +
Sbjct: 372 TPAGLESLIGNMSKDQIQNLIALFSSQLQPASPVLNTAPMSTSHNNDPSGITFSSSTFSF 431
Query: 410 ISQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSI 469
I ++ TWI+D+GAT H+C+ F + V+LP G G+I
Sbjct: 432 IGILTVSETEMTHGTWIVDSGATHHVCHVKDMFLNLDTSVQHHVNLPTGTTIRVGGVGNI 491
Query: 470 QLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVR 529
++ IL NVL++P F NL+SV L + R+ F C++ D IG+
Sbjct: 492 AVNADLILKNVLYIPEFRLNLLSVSALTTDIGARVVFDPTCCVVHDLTKGSTIGSD---- 547
Query: 530 SLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPY 589
L D L F + N + L H
Sbjct: 548 ----LLTDVLGIFKTKN------------------KGLLH-------------------- 565
Query: 590 VQYTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVD 649
C++C AKQK+L +P ++ F ++H+D+WGP S + G+ YFLTIVD
Sbjct: 566 --------CDICQRAKQKKLTYPSRHNICLAPFDLLHIDVWGPFSEPTQEGYHYFLTIVD 617
Query: 650 DYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIH 709
D++R TWVYL+K K +V T+ DF V Q+ VK VRSDN E + Y KGI+
Sbjct: 618 DHTRVTWVYLMKYKSDVLTIFPDFITMVETQYDTKVKAVRSDNAPELKFEELYRRKGIVA 677
Query: 710 HTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDN 769
+ SC ETP+QNS+VERKHQHILNVARALLFQ+ +P +W I+ ++F+INR P+P++ N
Sbjct: 678 YHSCPETPEQNSVVERKHQHILNVARALLFQSQIPLSYWGDCILTAVFIINRTPSPVISN 737
Query: 770 QCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDL 829
+ F++L K++PD T LK FG LC+AST R KF+ RA+ C FLG+ SG KGY + DL
Sbjct: 738 KTLFEMLTKKVPDYTHLKSFGCLCYASTSPKQRHKFEDRARTCAFLGYPSGYKGYKLLDL 797
Query: 830 NTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQI-----PCLPSEPFDYEYPIPPDN 884
+ I ISRNV+F+E++FP+K ++E S FP I PS+P
Sbjct: 798 ESHTIFISRNVVFYEDLFPFKTKPAENEESSVFFPHIYVDRNDSHPSQPL---------- 847
Query: 885 PVQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHP 944
PVQ S+ P++ + R + P YL+DYHC NS + T HP
Sbjct: 848 PVQETSASNVPAEKQ--------NSRVSRPPAYLKDYHC-----------NSVTSSTDHP 888
Query: 945 LSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVD 1004
+S+V+SY L Y FI + EP Y++A + + W AM EI ALE N TW++
Sbjct: 889 ISEVLSYSSLSDPYMIFINAVNKIPEPHTYAQARQIKEWCDAMGMEITALEDNGTWVVCS 948
Query: 1005 KPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVL 1064
P K +GCKWVY+IK DG+++RYKARLV KGYTQ EG+D++DTFSPVAK+TT+++L
Sbjct: 949 LPVGKKAVGCKWVYKIKLNADGSLERYKARLVAKGYTQTEGLDYVDTFSPVAKLTTVKLL 1008
Query: 1065 LSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMN-----SDKPNQVCLLQKSLYGL 1119
+++ + K W L QLD+ NAFL+ LDEEIYM+LP G + S PN VC L+KSLYGL
Sbjct: 1009 IAVAAAKGWSLSQLDISNAFLNGSLDEEIYMTLPPGYSPRQGDSFPPNAVCRLKKSLYGL 1068
Query: 1120 KQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIK 1179
KQASRQW+ ++L+ LGFTQS DHTL+ +KS S+ A+L+YVDD+++A + E +
Sbjct: 1069 KQASRQWYLKFSESLKALGFTQSSGDHTLFTRKSKN-SYMAVLVYVDDIIIASSCDRETE 1127
Query: 1180 LVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPM 1239
L++ +L + +++DLG ++FLGL IAR+ GI + QRKY LELL+++GLLG KS++ PM
Sbjct: 1128 LLRDALQRSSKLRDLGTLRYFLGLEIARNTDGISICQRKYTLELLAETGLLGCKSSSVPM 1187
Query: 1240 DCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAH 1299
+ +QKLS G + D YR+L+G+L+YLT TRPDI Y V++L QF SAP H A +
Sbjct: 1188 EPNQKLSQEDGELIDDAEHYRKLVGKLMYLTFTRPDITYAVHRLCQFTSAPRVPHLKAVY 1247
Query: 1300 RVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKK 1359
+++ Y+KG+ G GLFY A L+ F+DSD + C D+RK TGYCMFLG+SLV+W+SKK
Sbjct: 1248 KIIYYLKGTVGQGLFYSANVDLKLSGFADSDFSSCSDSRKLTTGYCMFLGTSLVAWKSKK 1307
Query: 1360 QTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYH 1419
Q S SS EAEY+AM+ V E+ WL +LL+DL + + ++CDN +A+HIA+NP +H
Sbjct: 1308 QEVISMSSAEAEYKAMSMAVREMMWLRFLLEDLWIDVSEASVLYCDNTAAIHIANNPVFH 1367
Query: 1420 ERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHI 1468
ERTKHIE D H +REKI GL+ L V + QLADI P + FR +
Sbjct: 1368 ERTKHIERDYHHIREKIILGLIRTLHVRTENQLADI---PYKSSIFRSV 1413
>UniRef100_Q9LFY6 T7N9.5 [Arabidopsis thaliana]
Length = 1436
Score = 881 bits (2277), Expect = 0.0
Identities = 505/1235 (40%), Positives = 707/1235 (56%), Gaps = 117/1235 (9%)
Query: 296 RNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQG 355
+ + KC++ ++GHTV+ CYK HG+PPG S NL T + + ++
Sbjct: 263 QGNFKKPKCTHCNRIGHTVDKCYKVHGYPPGHPRAKENTYVGSTNLASTDQIETQAPPTM 322
Query: 356 NTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGN---SISQ 412
+ E + D L+ L ST + SI SC + N SISQ
Sbjct: 323 SATGHET----MSNDHIQQLISYL-------STKLQSPSITSCFDKAIASSSNPVPSISQ 371
Query: 413 ISTK----NGNPL--------------DTT-------------------WILDTGATDHI 435
I+ K + NP+ D+T W++D+GA+ H+
Sbjct: 372 ITDKAIASSSNPVPSISQITGTFFSLYDSTYYEMLTSSIPIETELSLRAWVIDSGASHHV 431
Query: 436 CNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLPNFEFNLISVHK 495
+ + + +YK + V LPNG+ G IQL+ + L NVLF+P F+FNL+SV
Sbjct: 432 THERNLYHTYKALDRTFVRLPNGHTVKIEGTGFIQLTDALSLHNVLFIPEFKFNLLSVSV 491
Query: 496 LVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDS----LSPFPSCNSVSTC 551
L K+L+ +++F+ D+C+IQ M+G V +LYILN D +S FP + S+
Sbjct: 492 LTKTLQSKVSFTSDECMIQALTKELMLGKGSQVGNLYILNLDKSLVDVSSFPGKSVCSSV 551
Query: 552 KSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKD--LFPYVQYTQDHV-CEVCPIAKQKR 608
K+++ +WH RLGHPSF K ++ D + P + +D C VC ++KQK
Sbjct: 552 KNES----------EMWHKRLGHPSFAKIDTLSDVLMLPKQKINKDSSHCHVCHLSKQKH 601
Query: 609 LKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQT 668
L F N + F+++H+D WGP SV +++ + YFLTIVDD+SR TW+YLLK K +V T
Sbjct: 602 LPFKSVNHIREKAFELVHIDTWGPFSVPTVDSYRYFLTIVDDFSRATWIYLLKQKSDVLT 661
Query: 669 LVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTSCVETPQQNSIVERKHQ 728
+ F V Q+ V VRSDN E ++ +A++GI C ETP+QN +VERKHQ
Sbjct: 662 VFPSFLKMVETQYHTKVCSVRSDNAHELKFNELFAKEGIKADHPCPETPEQNFVVERKHQ 721
Query: 729 HILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPDITFLKV 788
H+LNVARAL+FQ+ +P +W ++ ++FLINRL +P+++N+ P++ L K PD + LK
Sbjct: 722 HLLNVARALMFQSGIPLEYWGDCVLTAVFLINRLLSPVINNETPYERLTKGKPDYSSLKA 781
Query: 789 FGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIFHENIFP 848
FG LC+ ST RTKFD RAK C+FLG+ G KGY + D+ T +SISR+VIF+E+IFP
Sbjct: 782 FGCLCYCSTSPKSRTKFDPRAKACIFLGYPMGYKGYKLLDIETYSVSISRHVIFYEDIFP 841
Query: 849 YKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNSEQPIVHRV- 907
+ + + FP I LP+ D P+ +Q S P + E + V
Sbjct: 842 FA-SSNITDAAKDFFPHI-YLPAPNNDEHLPL-------VQSSSDAPHNHDESSSMIFVP 892
Query: 908 ----SQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDPSYHAFIM 963
S R RK P++LQD+HC +T +K + +PL+ ISY L + AFI
Sbjct: 893 SEPKSTRQRKLPSHLQDFHCYNNTPTT-------TKTSPYPLTNYISYSYLSEPFGAFIN 945
Query: 964 NITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYK 1023
IT T P +YSEA + W AM EI A R TW + D P K +GCKW+ IK+
Sbjct: 946 IITATKLPQKYSEARLDKVWNDAMGKEISAFVRTGTWSICDLPAGKVAVGCKWIITIKFL 1005
Query: 1024 QDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNA 1083
DG+I+R+KARLV KGYTQ EGIDF +TFSPVAKM T++VLLSL W+LHQLD+ NA
Sbjct: 1006 ADGSIERHKARLVAKGYTQQEGIDFFNTFSPVAKMVTVKVLLSLAPKMKWYLHQLDISNA 1065
Query: 1084 FLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSF 1143
L+ L+EEIYM LP G + + +V K
Sbjct: 1066 LLNGDLEEEIYMKLPPGYSEIQGQEVSPNAKC---------------------------H 1098
Query: 1144 ADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGL 1203
DHTL+VK + G F +L+YVDD+L+A + L F+++DLGE KFFLG+
Sbjct: 1099 GDHTLFVK-AQDGFFLVVLVYVDDILIASTTEAASAELTSQLSSFFQLRDLGEPKFFLGI 1157
Query: 1204 AIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLI 1263
IAR+ GI L QRKY L+LL+ S K ++ PM+ +QKLS +GT L D YRR++
Sbjct: 1158 EIARNADGISLCQRKYVLDLLASSDFSDCKPSSIPMEPNQKLSKDTGTLLEDGKQYRRIL 1217
Query: 1264 GRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVL 1323
G+L YL TRPDI + V++L+Q+ SAPTD+H A H++LRY+KG+ G GLFY A ++ L
Sbjct: 1218 GKLQYLCLTRPDINFAVSKLAQYSSAPTDIHLQALHKILRYLKGTIGQGLFYGADTNFDL 1277
Query: 1324 TAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQ 1383
FSDSD C DTR+ +TG+ +F+G+SLVSWRSKKQ S SS EAEYRAM+ E+
Sbjct: 1278 RGFSDSDWQTCPDTRRCVTGFAIFVGNSLVSWRSKKQDVVSMSSAEAEYRAMSVATKELI 1337
Query: 1384 WLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHL 1443
WL Y+L ++ T P ++CDN++A+HIA+N +HERTKHIE DCH VRE I G++
Sbjct: 1338 WLGYILTAFKIPFTHPAYLYCDNEAALHIANNSVFHERTKHIENDCHKVRECIEAGILKT 1397
Query: 1444 LPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIH 1478
+ V + QLAD TKPL P PFR SKL L +I+
Sbjct: 1398 IFVRTDNQLADTLTKPLYPKPFRENNSKLGLLNIY 1432
>UniRef100_O23588 Retrotransposon like protein [Arabidopsis thaliana]
Length = 1433
Score = 872 bits (2252), Expect = 0.0
Identities = 489/1220 (40%), Positives = 688/1220 (56%), Gaps = 107/1220 (8%)
Query: 293 NSGRNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQEST 352
N + Y KCSY K+GH V+ CYKKHG+PPG K+ + S NL T + T
Sbjct: 288 NMAQGSYKKPKCSYCNKLGHLVDKCYKKHGYPPGSKWTKGQTIG-STNLASTQLQPVNET 346
Query: 353 SQGNTQEQEAARFGFTADQYHHLL------LLLPPSEQKNSTSQHTASINSCVQMLPGKN 406
T E F+ DQ ++ L + + +TS + S + V M+ +
Sbjct: 347 PNEKTDSYEE----FSTDQIQTMISYLSTKLHIASASPMPTTSSASISASPSVPMISQIS 402
Query: 407 GNSISQISTKNGNPLDTT-----------WILDTGATDHICNTLSYFSSYKHVAPIPVSL 455
G +S S + L ++ W++D+GAT H+ + + +++ + V L
Sbjct: 403 GTFLSLFSNAYYDMLISSVSQEPAVSPRGWVIDSGATHHVTHNRDLYLNFRSLENTFVRL 462
Query: 456 PNGNIESATIKGSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQD 515
PN G IQLS + L NVL++P F+FNLIS +L K L
Sbjct: 463 PNDCTVKIAGIGFIQLSDAISLHNVLYIPEFKFNLIS--ELTKEL--------------- 505
Query: 516 SNACKMIGTARAVRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHP 575
MIG V +LY+L+ + + S ++ + VC WH RLGHP
Sbjct: 506 -----MIGRGSQVGNLYVLDFNENNHTVSLKGTTSMCPEFSVCSSVVVDSVTWHKRLGHP 560
Query: 576 SFVKGQSIKDLFPYVQYTQD-------HVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVD 628
++ K + D+ + HVC VC ++KQK L F + F ++H+D
Sbjct: 561 AYSKIDLLSDVLNLKVKKINKEHSPVCHVCHVCHLSKQKHLSFQSRQNMCSAAFDLVHID 620
Query: 629 IWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIV 688
WGP SV + + TW+YLLK+K +V + F V Q+ +K V
Sbjct: 621 TWGPFSVPTNDA--------------TWIYLLKNKSDVLHVFPAFINMVHTQYQTKLKSV 666
Query: 689 RSDNGKEFVLSQFYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFW 748
RSDN E + +A GI+ + SC ETP+QNS+VERKHQHILNVARALLFQ+++P FW
Sbjct: 667 RSDNAHELKFTDLFAAHGIVAYHSCPETPEQNSVVERKHQHILNVARALLFQSNIPLEFW 726
Query: 749 AHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHR 808
++ ++FLINRLPTP+L+N+ P++ L P LK FG LC++ST R KF+ R
Sbjct: 727 GDCVLTAVFLINRLPTPVLNNKSPYEKLKNIPPAYESLKTFGCLCYSSTSPKQRHKFEPR 786
Query: 809 AKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIFHENIFPY--KLHRDDHECESSTFPQI 866
A+ CVFLG+ G KGY + D+ T +SISR+VIFHE+IFP+ +DD + FP +
Sbjct: 787 ARACVFLGYPLGYKGYKLLDIETHAVSISRHVIFHEDIFPFISSTIKDDIK---DFFPLL 843
Query: 867 PCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLA 926
P+ D P+ + + P S + P +S+R +K P +LQD+HC
Sbjct: 844 Q-FPARTDDL--PLEQTSIIDTHPHQDVSSSKALVPF-DPLSKRQKKPPKHLQDFHC--- 896
Query: 927 ATSTVVPANSSSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVA 986
Y+ +HAFI NIT V P RYSEA + W A
Sbjct: 897 ------------------------YNNTTEPFHAFINNITNAVIPQRYSEAKDFKAWCDA 932
Query: 987 MN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGI 1046
M EI A+ R NTW +V PP+K IGCKWV+ IK+ DG+I+RYKARLV KGYTQ EG+
Sbjct: 933 MKEEIGAMVRTNTWSVVSLPPNKKAIGCKWVFTIKHNADGSIERYKARLVAKGYTQEEGL 992
Query: 1047 DFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGM----- 1101
D+ +TFSPVAK+T++R++L L + W +HQLD+ NAFL+ LDEEIYM +P G
Sbjct: 993 DYEETFSPVAKLTSVRMMLLLAAKMKWSVHQLDISNAFLNGDLDEEIYMKIPPGYADLVG 1052
Query: 1102 NSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTAL 1161
+ P+ +C L KS+YGLKQASRQW+ L L+G+GF +S ADHTL++K A G +
Sbjct: 1053 EALPPHAICRLHKSIYGLKQASRQWYLKLSNTLKGMGFQKSNADHTLFIKY-ANGVLMGV 1111
Query: 1162 LLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYAL 1221
L+YVDD+++ N D + L F+++DLG AK+FLG+ IARS+KGI + QRKY L
Sbjct: 1112 LVYVDDIMIVSNSDDAVAQFTAELKSYFKLRDLGAAKYFLGIEIARSEKGISICQRKYIL 1171
Query: 1222 ELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVN 1281
ELLS +G LG K ++ P+D S KL+ G PL+D +SYR+L+G+L+YL TRPDIAY VN
Sbjct: 1172 ELLSTTGFLGSKPSSIPLDPSVKLNKEDGVPLTDSTSYRKLVGKLMYLQITRPDIAYAVN 1231
Query: 1282 QLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSI 1341
L QF APT +H +A H+VLRY+KG+ G GLFY A L ++DSD C D+R+ +
Sbjct: 1232 TLCQFSHAPTSVHLSAVHKVLRYLKGTVGQGLFYSADDKFDLRGYTDSDFGSCTDSRRCV 1291
Query: 1342 TGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVS 1401
YCMF+G LVSW+SKKQ T S S+ EAE+RAM+ E+ WL L D +V P
Sbjct: 1292 AAYCMFIGDYLVSWKSKKQDTVSMSTAEAEFRAMSQGTKEMIWLSRLFDDFKVPFIPPAY 1351
Query: 1402 MFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLS 1461
++CDN +A+HI +N +HERTK +ELDC+ RE + G + + V + Q+AD TK +
Sbjct: 1352 LYCDNTAALHIVNNSVFHERTKFVELDCYKTREAVESGFLKTMFVETGEQVADPLTKAIH 1411
Query: 1462 PAPFRHIFSKLRLYDIHSPV 1481
PA F + K+ + +I +P+
Sbjct: 1412 PAQFHKLIGKMGVCNIFAPL 1431
>UniRef100_Q5XWR5 Putative retroelement pol polyprotein-like [Solanum tuberosum]
Length = 1476
Score = 840 bits (2171), Expect = 0.0
Identities = 506/1314 (38%), Positives = 725/1314 (54%), Gaps = 121/1314 (9%)
Query: 232 CPMLIVSLLSLHSKKGSSLLKTFLGPLIAARENSNDGRSSQSNSGYQSNSGYQSGSGNYH 291
C +IV S S GS +T + G SQ + G SG S +GN H
Sbjct: 200 CYAMIVQDESQRSLSGSG--QTIDPTALFTHRPGGSGFGSQGSQG----SGNGSSNGNSH 253
Query: 292 S-NSGRNRYSN---------------KKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYA 335
+ G N Y + K C++ GHT + CY+ G+P +K K
Sbjct: 254 RFHKGGNIYCDFCNMKGHIRANCNKLKHCTHCNMQGHTKDTCYQLIGYPADYKGK----- 308
Query: 336 NRSANLVQTTKEDQESTSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASI 395
+ AN+V Q + N + +T D H + P + +T+ H++
Sbjct: 309 -KKANIVTAPSLPQMQHNNFNNNLNYPMQ--YTGDGIGH---FVSPMQFTGNTNGHSSGS 362
Query: 396 -------NSCVQMLPGKNGNSISQI-------------------STKNGNPLDTTWILDT 429
S Q P + N + + S N N + WI+D+
Sbjct: 363 IAGNFGPGSVPQFTPSQYNNILQMLNKPMLSESSANVAGIFAGSSHCNSNTHSSAWIVDS 422
Query: 430 GATDH-ICNT--LSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLPNF 486
GATDH + NT L++ S H P V LP G+ T GS QL+ ++ NVL +P F
Sbjct: 423 GATDHMVSNTTLLNHGLSVSH--PGKVQLPTGDSAVVTHSGSSQLTGGDVVKNVLCVPTF 480
Query: 487 EFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFPSCN 546
+FNL+SV KL K L + F D +IQD K+ + LYI P +
Sbjct: 481 QFNLLSVSKLTKELNCCVIFFPDFFIIQDLFTGKVKEIGEEINGLYITR-----PHQHHD 535
Query: 547 SVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQYTQDHVCEVCPIAKQ 606
+ + CE + +WH RLGH + IK +F Q C+VCP+A+Q
Sbjct: 536 TSKKTLAAIKGCEEA----EMWHKRLGHIPMSVLRKIK-MFDSPQKLVLPSCDVCPLARQ 590
Query: 607 KRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEV 666
RL FP+S S S+ F +IH+D+WGP + N YFLT+VDD+SR+TW++L+ K +V
Sbjct: 591 VRLPFPISQSRSENCFDLIHLDVWGPYKAATHNKMRYFLTVVDDHSRWTWIFLMHLKSDV 650
Query: 667 QTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQ---FYAEKGIIHHTSCVETPQQNSIV 723
T++++F + QFG +KI RSDNG EF +Q + GI+H +SC TPQQN +V
Sbjct: 651 STVLQNFILMIDTQFGQKIKIFRSDNGTEFFNAQCDGLFKSHGIVHQSSCPHTPQQNGVV 710
Query: 724 ERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPDI 783
ER+H+HIL ARAL FQ HLP FW ++ ++ +INR+P+ +L N+ PF+L++K+ PD+
Sbjct: 711 ERRHKHILETARALRFQGHLPIRFWGECVLSAVHIINRIPSSVLHNKSPFELMYKRSPDL 770
Query: 784 TFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIFH 843
++++V G LC A+ L + T+ KGY +YDL + +SR+++F+
Sbjct: 771 SYMRVIGCLCHATNLVNTSTQ-----------------KGYKLYDLEHQHFFVSRDMVFN 813
Query: 844 ENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPP-DNPVQIQPLSSGPSQNSEQ- 901
E +FP++ ++ F P S D + P +I P++S PS S+
Sbjct: 814 EAVFPFQSPALADPHDTPVFLASPPCSSHTEDADAVQPAIITSEEIIPVASPPSAVSDDH 873
Query: 902 ---PIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDPSY 958
P R S R K P + +D+ + + S S +P+S I Y L +Y
Sbjct: 874 LHPPPERRRSYRTGKPPIWQKDF---------ITTSTSRSNHCLYPISDNIDYSCLSSTY 924
Query: 959 HAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVY 1018
+I + + EP Y +A W AM EI+ALE N TW +V P K IGCKWVY
Sbjct: 925 QCYIASSSVETEPQFYYQAANDCRWVHAMKEEIQALEDNKTWEVVSLPKGKKAIGCKWVY 984
Query: 1019 RIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQL 1078
+IKYK G I+R+KARLV KGY Q EG+D+ +TFSPV KM TLR +L+L +K W + Q+
Sbjct: 985 KIKYKASGEIERFKARLVAKGYNQKEGLDYQETFSPVVKMVTLRTVLTLAVSKGWDIQQM 1044
Query: 1079 DVDNAFLHAKLDEEIYMSLPQGMNSDKPN--QVCLLQKSLYGLKQASRQWFSTLCQALQG 1136
DV NAFL L EE+YM LPQG DK +VC L KSLYGLKQASRQW L AL
Sbjct: 1045 DVYNAFLQGDLIEEVYMQLPQGFQYDKTGDPKVCRLLKSLYGLKQASRQWNVKLTTALLA 1104
Query: 1137 LGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGE 1196
GF QS D++L +K++A G +L+YVDD+L+ G+ + I K L F+IKDLG
Sbjct: 1105 AGFQQSHLDYSLMLKRTADG-IVIVLIYVDDLLITGSSLQLIDDAKQVLKANFKIKDLGT 1163
Query: 1197 AKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKL----------S 1246
++FLG+ AR+ G++++QRKYALEL+SD GL G K + TP++ KL S
Sbjct: 1164 LRYFLGMEFARNASGMLMHQRKYALELISDLGLGGSKPSVTPVELHLKLTTREFDLHVGS 1223
Query: 1247 ASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIK 1306
+ + + L+D + Y+RL+GRLLYLT TRPDI++ V LSQF+ AP H AA RV++Y+K
Sbjct: 1224 SGADSLLADPTEYQRLVGRLLYLTITRPDISFAVQHLSQFMHAPKVSHMEAAIRVVKYVK 1283
Query: 1307 GSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRS 1366
+PG GL+ ++ L A+ D+D C++TRKSITGY + GS+L+SW+SKKQ T SRS
Sbjct: 1284 QAPGLGLYMAVQTADTLQAYCDADWGSCINTRKSITGYMIQFGSALLSWKSKKQPTISRS 1343
Query: 1367 SCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIE 1426
S EAEYR++A+TV E+ WL L ++L + + PVS++CD+++A+ IA NP +HERTKHI+
Sbjct: 1344 SAEAEYRSLASTVAELVWLTGLFKELDMPLSLPVSLYCDSKAAIQIAANPVFHERTKHID 1403
Query: 1427 LDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHSP 1480
+DCH +REK+ GLV + + + Q ADI TK LS A ++ SKL L +I P
Sbjct: 1404 IDCHFIREKVQAGLVMIHYLPTQEQPADILTKGLSSAQHSYLVSKLGLKNIFIP 1457
>UniRef100_Q9SLF0 Putative retroelement pol polyprotein [Arabidopsis thaliana]
Length = 1333
Score = 740 bits (1911), Expect = 0.0
Identities = 454/1227 (37%), Positives = 682/1227 (55%), Gaps = 95/1227 (7%)
Query: 291 HSNSGRNRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQE 350
+ G + + C + + GH E CY G+P ++ + +RS +QT
Sbjct: 164 YQGRGDEKDKSMVCKHCNRSGHASESCYAVIGYP---EWWGDRPRSRS---LQTRGRGGT 217
Query: 351 STSQGNTQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSI 410
++S G G A Y + + + N + A+ + G NG +
Sbjct: 218 NSSGGR---------GRGAAAYANRVTV------PNHDTYEQANYALTDEDRDGVNGLTD 262
Query: 411 SQISTKNGNPLDTTWILDTG---ATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKG 467
SQ T IL++G AT+ + + + PI + L +G + +G
Sbjct: 263 SQWRTIKS-------ILNSGKDAATEKLTD----------MDPILIVLADGRERISVKEG 305
Query: 468 SIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARA 527
+++L + ++I+V ++ F+ +LIS+ +L+ R L SD ++QD + ++G R
Sbjct: 306 TVRLGSNLVMISVFYVEEFQSDLISIGQLMDENRCVLQMSDRFLVVQDRTSRMVMGAGRR 365
Query: 528 VRSLYILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLF 587
V + + ++ SV+ + +N LWH R+GHP+ I +
Sbjct: 366 VGGTFHFRSTEIAA-----SVTVKEEKNY---------ELWHSRMGHPAARVVSLIPESS 411
Query: 588 PYVQYTQ-DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLT 646
V T + C+VC AKQ R FPLS + + IF++I+ D+WGP S G YFLT
Sbjct: 412 VSVSSTHLNKACDVCHRAKQTRNSFPLSINKTLRIFELIYCDLWGPYRTPSHTGARYFLT 471
Query: 647 IVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV-LSQFYAEK 705
I+DDYSR W+YLL K E +K+F A QF V +K VRSDNG EF+ L++F+ E+
Sbjct: 472 IIDDYSRGVWLYLLNDKSEAPCHLKNFFAMTDRQFNVKIKTVRSDNGTEFLCLTKFFQEQ 531
Query: 706 GIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTP 765
G+IH SCV TP++N VERKH+H+LNVARAL FQA+LP FW ++ + +LINR P+
Sbjct: 532 GVIHERSCVATPERNDRVERKHRHLLNVARALRFQANLPIQFWGECVLTAAYLINRTPSS 591
Query: 766 ILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYL 825
+L++ P++ LHK+ P L+VFGSLC+A KF R++RCVF+G+ G KG+
Sbjct: 592 VLNDSTPYERLHKKQPRFDHLRVFGSLCYAHNRNRGGDKFAERSRRCVFVGYPHGQKGWR 651
Query: 826 VYDLNTRDISISRNVIFHENIFPYKLHRD----DHECESSTFPQIPCLPSEPFDYEY--- 878
++DL + +SR+V+F E FP+++ + + E E+ P + L E
Sbjct: 652 LFDLEQNEFFVSRDVVFSELEFPFRISHEQNVIEEEEEALWAPIVDGLIEEEVHLGQNAG 711
Query: 879 PIPP---DNPVQIQPLSSGPSQNSEQPIVHRVSQRPR---------KQPTYLQDYHCTLA 926
P PP +P+ SS ++ P+ V P PT LQ + A
Sbjct: 712 PTPPICVSSPISPSATSSRSEHSTSSPLDTEVVPTPATSTTSASSPSSPTNLQFLPLSRA 771
Query: 927 ATSTVVPANSSSKGTSHPLSQVISYHKLDP-SYHAFIMNITTTVE-PTR-----YSEAVK 979
+T A + + P + + +K P + F++N T E P++ Y +
Sbjct: 772 KPTT---AQAVAPPAVPPPRRQSTRNKAPPVTLKDFVVNTTVCQESPSKLNSILYQLQKR 828
Query: 980 HECWRVAMN*E----IEALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARL 1035
+ R + + I+A E N+TW + D PP K IG +WVY++K+ DG+++RYKARL
Sbjct: 829 DDTRRFSASHTTYVAIDAQEENHTWTIEDLPPGKRAIGSQWVYKVKHNSDGSVERYKARL 888
Query: 1036 VVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYM 1095
V G Q EG D+ +TF+PVAKM T+R+ L + +NW +HQ+DV NAFLH L EE+YM
Sbjct: 889 VALGNKQKEGEDYGETFAPVAKMATVRLFLDVAVKRNWEIHQMDVHNAFLHGDLREEVYM 948
Query: 1096 SLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLY--VKKS 1153
LP G + PN+VC L+K+LYGLKQA R WF L AL+ GF QS AD++L+ VK S
Sbjct: 949 KLPPGFEASHPNKVCRLRKALYGLKQAPRCWFEKLTTALKRYGFQQSLADYSLFTLVKGS 1008
Query: 1154 ATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGII 1213
+L+YVDD+++ GN + K L F +KDLG K+FLG+ +ARS GI
Sbjct: 1009 VR---IKILIYVDDLIITGNSQRATQQFKEYLASCFHMKDLGPLKYFLGIEVARSTTGIY 1065
Query: 1214 LNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTR 1273
+ QRKYAL+++S++GLLG K A P++ + KL S+ L+D YRRL+GRL+YL TR
Sbjct: 1066 ICQRKYALDIISETGLLGVKPANFPLEQNHKLGLSTSPLLTDPQRYRRLVGRLIYLAVTR 1125
Query: 1274 PDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAG 1333
D+A+ V+ L++F+ P + H AAA RV+RY+K PG G+F + +T + DSD AG
Sbjct: 1126 LDLAFSVHILARFMQEPREDHWAAALRVVRYLKADPGQGVFLRRSGDFQITGWCDSDWAG 1185
Query: 1334 CVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQ 1393
+R+S+TGY + G S +SW++KKQ T S+SS EAEYRAM+ E+ WL LL L
Sbjct: 1186 DPMSRRSVTGYFVQFGDSPISWKTKKQDTVSKSSAEAEYRAMSFLASELLWLKQLLFSLG 1245
Query: 1394 VSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLA 1453
VS P+ M CD++SA++IA NP +HERTKHIE+D H VR++ +G++ V ++ QLA
Sbjct: 1246 VSHVQPMIMCCDSKSAIYIATNPVFHERTKHIEIDYHFVRDEFVKGVITPRHVGTTSQLA 1305
Query: 1454 DIFTKPLSPAPFRHIFSKLRLYDIHSP 1480
DIFTKPL F KL + ++++P
Sbjct: 1306 DIFTKPLGRDCFSAFRIKLGIRNLYAP 1332
>UniRef100_Q710T7 Gag-pol polyprotein [Populus deltoides]
Length = 1382
Score = 708 bits (1828), Expect = 0.0
Identities = 434/1243 (34%), Positives = 646/1243 (51%), Gaps = 76/1243 (6%)
Query: 249 SLLKTFLGPLIAARENSNDGRSSQSNSGYQSNSGYQSGSGNYHSNSGRNRYSNKKCSYYG 308
S++ L I + S G S SN S S + H N R +CS+
Sbjct: 193 SVVSELLAEEIRLQSYSEKGILSASNP---SVLAVPSKPFSNHQNKPYTRVGFDECSFCK 249
Query: 309 KMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQGNTQEQEAARFGFT 368
+ GH C PK ++ Q K +S S + Q G+
Sbjct: 250 QKGHWKAQC--------------PKLRQQN----QAWKSGSQSQSNAHRSPQ-----GYK 286
Query: 369 ADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLP-GKNGNSISQISTKNGNPLDTTWIL 427
++ + P S +T + + P + +SI Q+ + + W+L
Sbjct: 287 PPHHNTAAVASPGSITDPNTLAE--QFQKFLSLQPQAMSASSIGQLPHSSSGISHSEWVL 344
Query: 428 DTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLPNFE 487
D+GA+ H+ S F+S ++ IPV +G GS+ ++ L NV +P +
Sbjct: 345 DSGASHHMSPDSSSFTSVSPLSSIPVMTADGTPMPLAGVGSV-VTLHLSLPNVYLIPKLK 403
Query: 488 FNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFPSCNS 547
NL S+ ++ S + + FS C +QD + K+IGT R LYIL+ + P +
Sbjct: 404 LNLASIGQICDSGDYLVMFSGSFCCVQDLQSQKLIGTGRRENGLYILDELKV---PVVVA 460
Query: 548 VSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQYTQDHV--CEVCPIAK 605
+T S S LWH RLGH S + + + + C C +AK
Sbjct: 461 ATTVDLSFFRLSLSSSSFYLWHSRLGHVSSSRLRFLASTGALGNLKTCDISDCSGCKLAK 520
Query: 606 QKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDE 665
L F S S S F +IH D+WGP V + G Y+++ +DD++RY WVYL+K + E
Sbjct: 521 FSALPFNRSTSVSSSPFDLIHSDVWGPSPVSTKGGSRYYVSFIDDHTRYCWVYLMKHRSE 580
Query: 666 VQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQF---YAEKGIIHHTSCVETPQQNSI 722
+ F A + Q +K R D G E+ ++F A G IH TSC +TP+QN +
Sbjct: 581 FFEIYAAFRALIKTQHSAVIKCFRCDLGGEYTSNKFCQMLALDGTIHQTSCTDTPEQNGV 640
Query: 723 VERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPD 782
ERKH+HI+ AR+LL A + FW A++ ++ LIN +P+ PF+ L+ +PD
Sbjct: 641 AERKHRHIVETARSLLLSAFVLSEFWGEAVLTAVSLINTIPSSHSSGLSPFEKLYGHVPD 700
Query: 783 ITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIF 842
+ +VFG F R K R+ CVFLG+ G KGY +D T+ + +S +V+F
Sbjct: 701 YSSFRVFGCTYFVLHPHVERNKLSSRSAICVFLGYGEGKKGYRCFDPITQKLYVSHHVVF 760
Query: 843 HENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNSEQP 902
E+I + + H S I +PF S S N P
Sbjct: 761 LEHIPFFSIPSTTHSLTKSDLIHI-----DPF------------------SEDSGNDTSP 797
Query: 903 IVHRV-SQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQV----------ISY 951
V + + T L A+ S+ P SS P + +Y
Sbjct: 798 YVRSICTHNSAGTGTLLSG--TPEASFSSTAPQASSEIVDPPPRQSIRIRKSTKLPDFAY 855
Query: 952 HKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTP 1011
S+ +F+ I EP+ Y EA+ + AM+ E+ AL + +TW LV PP K+
Sbjct: 856 SCYSSSFTSFLAYIHCLFEPSSYKEAILDPLGQQAMDEELSALHKTDTWDLVPLPPGKSV 915
Query: 1012 IGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTK 1071
+GC+WVY+IK DG+I+RYKARLV KGY+Q G+D+ +TF+P+AKMTT+R L+++ S +
Sbjct: 916 VGCRWVYKIKTNSDGSIERYKARLVAKGYSQQYGMDYEETFAPIAKMTTIRTLIAVASIR 975
Query: 1072 NWFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFSTLC 1131
W + QLDV NAFL+ L EE+YM+ P G++ D VC L+K+LYGLKQA R WF
Sbjct: 976 QWHISQLDVKNAFLNGDLQEEVYMAPPPGISHDS-GYVCKLKKALYGLKQAPRAWFEKFS 1034
Query: 1132 QALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRI 1191
+ LGF S D L++K + G L LYVDD+++ G+D+D I ++K L ++F +
Sbjct: 1035 IVISSLGFVSSSHDSALFIKCTDAGRI-ILSLYVDDMIITGDDIDGISVLKTELARRFEM 1093
Query: 1192 KDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGT 1251
KDLG ++FLG+ +A S +G +L+Q KY +L + L K+ TP++ + + S+S G
Sbjct: 1094 KDLGYLRYFLGIEVAYSPRGYLLSQSKYVANILERARLTDNKTVDTPIEVNARYSSSDGL 1153
Query: 1252 PLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGC 1311
PL D + YR ++G L+YLT T PDIAY V+ +SQF+++PT +H AA R+LRY++G+
Sbjct: 1154 PLIDPTLYRTIVGSLVYLTITHPDIAYAVHVVSQFVASPTTIHWAAVLRILRYLRGTVFQ 1213
Query: 1312 GLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAE 1371
L + SS L A+SD+D RKS+TG+C+FLG SL+SW+SKKQ+ S+SS EAE
Sbjct: 1214 SLLLSSTSSLELRAYSDADHGSDPTDRKSVTGFCIFLGDSLISWKSKKQSIVSQSSTEAE 1273
Query: 1372 YRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHI 1431
Y AMA+T E+ W +LL D+ +S + M+CDNQS++ IAHN +HERTKHIE+DCH+
Sbjct: 1274 YCAMASTTKEIVWSRWLLADMGISFSHLTPMYCDNQSSIQIAHNSVFHERTKHIEIDCHL 1333
Query: 1432 VREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRL 1474
R + G + L V SSLQ+AD FTK S + F + KL +
Sbjct: 1334 TRHHLKHGTIALPFVPSSLQIADFFTKAHSISRFCFLVGKLSM 1376
>UniRef100_Q7XE85 Putative pol polyprotein [Oryza sativa]
Length = 1688
Score = 698 bits (1802), Expect = 0.0
Identities = 426/1101 (38%), Positives = 606/1101 (54%), Gaps = 77/1101 (6%)
Query: 408 NSISQISTKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAP-IPVSLPNGNIESATIK 466
+SIS ++ P WILD+GA+ H+ S+ +S + V V NG + T +
Sbjct: 171 SSISTVTPIASQP----WILDSGASFHMSFDDSWLTSCRLVKNGATVHTANGTLCKVTHQ 226
Query: 467 GSIQLSPSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTA- 525
GSI SP F + NV +P NLISV +L + F + F D C +QD + +IGT
Sbjct: 227 GSIS-SPQFTVPNVSLVPKLSMNLISVGQLTDTNCF-VGFDDTSCFVQDRHTGAVIGTGH 284
Query: 526 RAVRS--LYILNNDSLSPFPSCNSVSTCKSQ-NLVCEFSPSVQNLWHYRLGHPSFVKGQS 582
R RS LYIL++ SL P S N+ S + C+ P WH+RLGH + +
Sbjct: 285 RQKRSCGLYILDSLSL-PSSSTNTPSVYSPMCSTACKSFPQ----WHHRLGHLCGSRLAT 339
Query: 583 I--KDLFPYVQYTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNG 640
+ + + V VC+ C + KQ +L +P S S S F ++H D+WG S G
Sbjct: 340 LINQGVLGSVPVDTTFVCKGCKLGKQVQLPYPSSTSRSSRPFDLVHSDVWGKSPFPSKGG 399
Query: 641 FSYFLTIVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLS- 699
+Y++ VDDYSRYTW+Y +K + ++ ++ + F + QF +++I RSD+G E++ +
Sbjct: 400 HNYYVIFVDDYSRYTWIYFMKHRSQLISIYQSFAQMIHTQFSSAIRIFRSDSGGEYMSNA 459
Query: 700 --QFYAEKGIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIF 757
+F +G + SC QN + ERKH+HI+ AR LL + +P FWA AI +++
Sbjct: 460 FREFLVSQGTLPQLSCPGAHAQNGVAERKHRHIIETARTLLIASFVPAHFWAEAISTAVY 519
Query: 758 LINRLPTPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGF 817
LIN P+ L + P ++L P L+VFG C+ RTK ++ CVFLG+
Sbjct: 520 LINMQPSSSLQGRSPGEVLFGSPPRYDHLRVFGCTCYVLLAPRERTKLTAQSVECVFLGY 579
Query: 818 KSGTKGYLVYDLNTRDISISRNVIFHENI-FPYKLHRDDHECESST----FPQIPC---L 869
KGY YD + R I ISR+V F EN F Y E+S P IP L
Sbjct: 580 SLEHKGYRCYDPSARRIRISRDVTFDENKPFFYSSTNQPSSPENSISFLYLPPIPSPESL 639
Query: 870 PSEPFDYE-YPIPPD--NPVQIQPLSSGPSQNSEQPIVHRV------------------- 907
PS P PIPP +P + P PS + P +
Sbjct: 640 PSSPITPSPSPIPPSVPSPTYVPPPPPSPSPSPVSPPPSHIPASSSPPHVPSTITLDTFP 699
Query: 908 ---SQRPR-------KQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDPS 957
S+RP+ QPT L+D C++ +S N ++ P+
Sbjct: 700 FHYSRRPKIPNESQPSQPT-LEDPTCSVDDSSPAPRYNLRARDALRA-----------PN 747
Query: 958 YHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWV 1017
F++ + EP+ Y EA+ W++AM+ E+ ALER NTW +V P PI CKWV
Sbjct: 748 RDDFVVGVV--FEPSTYQEAIVLPHWKLAMSEELAALERTNTWDVVPLPSHAVPITCKWV 805
Query: 1018 YRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQ 1077
Y++K K DG ++RYKARLV +G+ Q G D+ +TF+PVA MTT+R L+++ +T++W + Q
Sbjct: 806 YKVKTKSDGQVERYKARLVARGFQQAHGRDYDETFAPVAHMTTVRTLIAVAATRSWTISQ 865
Query: 1078 LDVDNAFLHAKLDEEIYMSLPQGMNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGL 1137
+DV NAFLH L EE+YM P G+ + P V L+++LYGLKQA R WF+ +
Sbjct: 866 MDVKNAFLHGDLHEEVYMHPPPGVEAP-PGHVFRLRRALYGLKQAPRAWFARFSSVVLAA 924
Query: 1138 GFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEA 1197
GF+ S D L++ S+ G T LLLYVDD+L+ G+D++ I VK L ++F + DLG
Sbjct: 925 GFSPSDHDPALFIHTSSRGR-TLLLLYVDDMLITGDDLEYIAFVKGKLSEQFMMSDLGPL 983
Query: 1198 KFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDIS 1257
+FLG+ + + G L+Q +Y +LL+ SGL ++ TTPM+ +L ++ GTPL D S
Sbjct: 984 SYFLGIEVTSTVDGYYLSQHRYIEDLLAQSGLTDSRTTTTPMELHVRLRSTDGTPLDDPS 1043
Query: 1258 SYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPA 1317
YR L+G L+YLT TRPDIAY V+ LSQF+SAP +H RVLRY++G+ LFY A
Sbjct: 1044 RYRHLVGSLVYLTVTRPDIAYAVHILSQFVSAPISVHYGHLLRVLRYLRGTTTQCLFYAA 1103
Query: 1318 ASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAA 1377
+S L AFSDS A R+S+TGYC+FLG+SL++W+SKKQT SRSS EAE RA+A
Sbjct: 1104 SSPLQLRAFSDSTWASDPIDRRSVTGYCIFLGTSLLTWKSKKQTAVSRSSTEAELRALAT 1163
Query: 1378 TVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKIT 1437
T E+ WL +LL D VS P + CDN A+ IA++P HE TKHI +D R
Sbjct: 1164 TTSEIVWLRWLLADFGVSCDVPTPLLCDNTGAIQIANDPIKHELTKHIGVDASFTRSHCQ 1223
Query: 1438 QGLVHLLPVTSSLQLADIFTK 1458
Q + L V S LQ+AD FTK
Sbjct: 1224 QSTIALHYVPSELQVADFFTK 1244
>UniRef100_Q8S805 Putative copia-type polyprotein [Oryza sativa]
Length = 1803
Score = 654 bits (1686), Expect = 0.0
Identities = 399/1076 (37%), Positives = 571/1076 (52%), Gaps = 65/1076 (6%)
Query: 420 PLDTTWILDTGATDHICNTLSYFSSYKHVAPIP----VSLPNGNIESATIKGSIQL---S 472
P W LDTGA+ H+ +T + H P+P +++ NG T S + S
Sbjct: 363 PSAADWFLDTGASAHMSSTPGILA---HPRPLPFSSCITVGNGAKLPVTHTASTHIPTSS 419
Query: 473 PSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLY 532
L NVL P NLISV KL + + F I+D + + LY
Sbjct: 420 TDLHLHNVLVSPPLIKNLISVKKLTRDNNVSIEFDPTGFSIKDLQTQVVKLRCDSPGDLY 479
Query: 533 ILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPY-VQ 591
L PS +++S S PSV++ WH RLGHP + F +
Sbjct: 480 PLR------LPSPHALSATSS--------PSVEH-WHLRLGHPGSASLSKVLGSFDFQCN 524
Query: 592 YTQDHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDY 651
+ H C C + RL F S+S + FQ++H D+W + S +G+ Y++ +DD+
Sbjct: 525 KSAPHHCSACHVGTNVRLPFHSSSSQTLFPFQLVHTDVWTS-PIYSNSGYKYYVVFLDDF 583
Query: 652 SRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEF---VLSQFYAEKGII 708
+ Y W + +++K EV V+ F A+ QFG+ V +++DNGKE+ L + G +
Sbjct: 584 THYIWTFPVRNKSEVFHTVRSFFAYAHTQFGLPVLALQTDNGKEYDSYALRSLLSLHGAV 643
Query: 709 HHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILD 768
SC + QQN ER + I + R +L + P FWA A+ ++ LINR P
Sbjct: 644 LRLSCPYSSQQNGKAERILRTINDCVRTMLVHSAAPLSFWAEALQTAMHLINRRPCRATG 703
Query: 769 NQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYD 828
+ P+QLL P L+VFG LC+ +T+A+ K R+ CVF+G+ + +GY YD
Sbjct: 704 SLKPYQLLLGAPPTYDHLRVFGCLCYPNTIATAPHKLSPRSLACVFIGYPADHRGYRCYD 763
Query: 829 LNTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQI 888
+ +R + SR+V F E++FP+ RD S P P P D +P +
Sbjct: 764 MVSRRVFTSRHVTFVEDVFPF---RD----APSPRPSAPPPPDHGDDTIVLLPAPAQHVV 816
Query: 889 QPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQV 948
P+ + P+ ++ P P+ + A V P S TS P S
Sbjct: 817 TPVGTAPAHDAASP------------PSPASSTPSSAAPAHDVAPPPSPE--TSSPASAS 862
Query: 949 ISYHKLDPSYHA--------FIMNITTTVEPTRYSE--AVKHECWRVAMN*EIEALERNN 998
H + A + M T+T+ PT S A++ WR AM E +AL N
Sbjct: 863 PPRHAMTTRARAGISKPNPRYAMTATSTLSPTPSSVRVALRDPNWRAAMQAEFDALLANR 922
Query: 999 TWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKM 1058
TW LV +PP I KWV++ K DG++D+YKAR VV+G+ Q G+DF +TFSPV K
Sbjct: 923 TWTLVPRPPGARIITGKWVFKTKLHADGSLDKYKARWVVRGFNQRPGVDFGETFSPVVKP 982
Query: 1059 TTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGM-NSDKPNQVCLLQKSLY 1117
T+R +L+L+S+K W HQLDV NAFLH L E + P G ++ +P VCLL +SLY
Sbjct: 983 ATIRTVLTLISSKQWPAHQLDVSNAFLHGHLQERVLCQQPTGFEDAARPADVCLLSRSLY 1042
Query: 1118 GLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTA-LLLYVDDVLLAGNDMD 1176
GL+QA R WF LGF QS AD +L+V + GS TA LLLYVDD++L+ +
Sbjct: 1043 GLRQAPRAWFKRFADHATSLGFVQSRADPSLFVLRR--GSDTAYLLLYVDDMILSASSSS 1100
Query: 1177 EIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSAT 1236
++ + L +F++KD+G K+FLG+ + R+ G +L+Q KYA ++L +G+ K+
Sbjct: 1101 LLQRIIDRLQAEFKVKDMGPLKYFLGIEVQRTADGFVLSQSKYATDVLERAGMANCKAVA 1160
Query: 1237 TPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEA 1296
TP D KLS+ G D S YR + G L YLT TRPDIAY V Q+ + AP + H
Sbjct: 1161 TPADAKPKLSSDEGPLFQDSSWYRSIAGALQYLTLTRPDIAYAVQQVCLHMHAPREAHVT 1220
Query: 1297 AAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWR 1356
R+LRYIKG+ GL A++S LTAFSD+D AGC DTR+S +G+C+FLG SL+SW
Sbjct: 1221 LLKRILRYIKGTAAFGLHLRASTSPTLTAFSDADWAGCPDTRRSTSGFCIFLGDSLISWS 1280
Query: 1357 SKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNP 1416
SK+QTT SRSS EAEYR +A V E WL LL +L +CDN S+++++ NP
Sbjct: 1281 SKRQTTVSRSSAEAEYRGVANAVAECTWLRQLLGELHCRVPQATIAYCDNISSVYMSKNP 1340
Query: 1417 SYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKL 1472
+H+RTKHIELD H VREK+ G + +LP+ S+ Q AD+FTK L + F + L
Sbjct: 1341 VHHKRTKHIELDIHFVREKVALGELRVLPIPSAHQFADVFTKGLPSSMFNEFRASL 1396
>UniRef100_O23741 SLG-Sc and SLA-Sc genes and Melmoth retrotransposon sequence
[Brassica oleracea]
Length = 1131
Score = 633 bits (1633), Expect = e-179
Identities = 349/914 (38%), Positives = 509/914 (55%), Gaps = 62/914 (6%)
Query: 304 CSYYGKMGHTVEDCYKKHGFPPGF--KFKNPKYANRSANLVQTTKEDQESTSQGNTQEQE 361
CS+ ++GH E CYKKHGFPPGF K+K+ +R Q + S
Sbjct: 261 CSFCNRVGHIAERCYKKHGFPPGFVSKYKSQSSGDRLQKPKQVAAQVSFSPPNSGQSPMT 320
Query: 362 AARF--GFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNS-----ISQIS 414
+ +Q + L + + AS + G + N I ++
Sbjct: 321 MDHLVGNHSKEQLQQFIALFSSQLPNVTMGSNEASSSKQPMDNSGISFNPTTLVFIGLLT 380
Query: 415 TKNGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPS 474
+ TWI+D+GAT H+C+ S ++S V+LPNG I + G +QL+
Sbjct: 381 VSRHTLANETWIIDSGATHHVCHDRSMYTSIDITTTSNVNLPNGMIVKISGVGIVQLNEH 440
Query: 475 FILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYIL 534
L NVL++P F NL+S+ L + ++ F C IQD IG R V +LY+L
Sbjct: 441 ITLHNVLYIPEFRLNLLSISSLTSDIGSQVIFDVSSCAIQDPTKGWTIGQGRRVANLYVL 500
Query: 535 NNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSIKDLFPYVQYTQ 594
+ S SP N V + S LWH RLGHPS+ + I + ++
Sbjct: 501 DVKS-SPMKI----------NAVVDIS-----LWHKRLGHPSYTRLDKISEALGTTKHKN 544
Query: 595 --DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYS 652
D C VC +AKQK+L + N FQ++HVD+WGP SV +L G+ YFLTIVDD+S
Sbjct: 545 KGDAHCHVCHLAKQKKLSYSSQNHICTASFQLLHVDVWGPFSVETLEGYKYFLTIVDDHS 604
Query: 653 RYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTS 712
R TW+YLL+SK +V + F + Q+ +K VR DN E ++ + EKGI+ + S
Sbjct: 605 RATWIYLLQSKSDVLHIFPTFVNQIETQYNTKIKSVRRDNAPELSFTELFKEKGIVSYHS 664
Query: 713 CVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCP 772
C ET +QNS++ERKHQH+LNVARAL+FQ+ +P +W ++ + FLINR P+P+L N+ P
Sbjct: 665 CPETLEQNSVLERKHQHLLNVARALMFQSQVPLQYWGDCVLTAAFLINRTPSPLLANKSP 724
Query: 773 FQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTR 832
+++L + P L+ FG LC+ ST R KF R++ CVFLG+ SG KGY + DL +
Sbjct: 725 YEVLMGKAPQYDQLRTFGCLCYGSTSPKQRHKFMPRSRACVFLGYPSGYKGYKLLDLESN 784
Query: 833 DISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLS 892
I ISRNV FHE+IFP H+ E FP +PS P S
Sbjct: 785 KIYISRNVTFHEDIFPMAKHQKMDESSLHFFPPKVTVPSAP------------------S 826
Query: 893 SGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYH 952
S + + ++S+R R P +L+D+HC S +++P+S +SY
Sbjct: 827 PNISSSPFSTLSPQISKRQRTVPAHLKDFHC------------YSVHDSAYPISSTLSYS 874
Query: 953 KLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPI 1012
++ + A+I +IT P Y+E + + W + + E++A+E N+TW +V P K I
Sbjct: 875 QISSHHLAYINSITNIPIPQSYAEVRQSKEWTESADKELDAMEENDTWDVVPLPKGKKAI 934
Query: 1013 GCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKN 1072
GC+WV+ +K+ DGT++R K+RLV KGYTQ EG+D+++TFSPVAKM T+++LL + ++K
Sbjct: 935 GCRWVHTLKFNADGTLERRKSRLVGKGYTQKEGLDYIETFSPVAKMATVKLLLKVGASKK 994
Query: 1073 WFLHQLDVDNAFLHAKLDEEIYMSLPQGMNSDK----PNQVCLLQKSLYGLKQASRQWFS 1128
WFLHQLD+ NAFL+ +LDEEIYM LP+G K PN VC L+KS+YGLKQASRQWF
Sbjct: 995 WFLHQLDISNAFLNGELDEEIYMKLPEGYAERKGDLPPNAVCKLKKSIYGLKQASRQWFK 1054
Query: 1129 TLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQK 1188
+L LGF ++ DHTL+V+++ F A+L+YVDD+++A D +K L
Sbjct: 1055 KFSTSLFQLGFQKAHGDHTLFVRQT-ENDFVAVLVYVDDIVIASTDDAVAVKLKSDLKSF 1113
Query: 1189 FRIKDLGEAKFFLG 1202
F+++DLG K+F G
Sbjct: 1114 FKLRDLGSLKYFFG 1127
>UniRef100_Q94IU9 Copia-like polyprotein [Arabidopsis thaliana]
Length = 1466
Score = 632 bits (1630), Expect = e-179
Identities = 390/1081 (36%), Positives = 575/1081 (53%), Gaps = 60/1081 (5%)
Query: 425 WILDTGATDHIC-------NTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFIL 477
W D+ AT HI N +Y + + LP ++ S TI S P L
Sbjct: 324 WYPDSAATAHITASTSGLQNATTYEGNDAVLVGDGTYLPITHVGSTTISSSKGTIP---L 380
Query: 478 INVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNND 537
VL P + +L+SV KL + F + I D K++ LY+L N
Sbjct: 381 NEVLVCPAIQKSLLSVSKLCDDYPCGVYFDANKVCIIDLTTQKVVSKGPRNNGLYMLEN- 439
Query: 538 SLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSFVKGQSI---KDLFPYVQYTQ 594
S F + S C + WH+RLGH + Q + K++ T
Sbjct: 440 --SEFVALYSNRQCAAS----------METWHHRLGHSNSKILQQLLTRKEIQVNKSRTS 487
Query: 595 DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRY 654
VCE C + K RL+F S+ + +H D+WGP V+S GF Y+ VDD+SR+
Sbjct: 488 P-VCEPCQMGKSTRLQFFSSDFRALKPLDRVHCDLWGPSPVVSNQGFKYYAVFVDDFSRF 546
Query: 655 TWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFV---LSQFYAEKGIIHHT 711
+W + L+ K + ++ + V NQ G +K +SD G EF L + + E GI H
Sbjct: 547 SWFFPLRMKSKFISVFIAYQKLVENQLGTKIKEFQSDGGGEFTSNKLKEHFREHGIHHRI 606
Query: 712 SCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQC 771
SC TPQQN + ERKH+H++ + ++L+ +H P FW A + +L N LP+ +L
Sbjct: 607 SCPYTPQQNGVAERKHRHLVELGLSMLYHSHTPLKFWVEAFFTANYLSNLLPSSVLKEIS 666
Query: 772 PFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNT 831
P++ L +Q D T L+VFG+ C+ + KFD R+ +CVFLG+ + KGY T
Sbjct: 667 PYETLFQQKVDYTPLRVFGTACYPCLRPLAKNKFDPRSLQCVFLGYHNQYKGYRCLYPPT 726
Query: 832 RDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPV-QIQP 890
+ ISR+VIF E FP+K E S P+ + + + PP P Q+QP
Sbjct: 727 GKVYISRHVIFDEAQFPFK------EKYHSLVPKYQTTLLQAWQHTDLTPPSVPSSQLQP 780
Query: 891 LSSG--PSQNSE-QPIVHRVSQRP--------RKQPTYLQDY--HCTLAATSTVVPANSS 937
L+ P SE QP+++ ++ + T D H + N+
Sbjct: 781 LARQMTPMATSENQPMMNYETEEAVNVNMETSSDEETESNDEFDHEVAPVLNDQNEDNAL 840
Query: 938 SKGTSHPLSQVIS-----YHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIE 992
+G+ L +I+ K +P Y A I++ ++ EP + A+KH W A+ EI+
Sbjct: 841 GQGSLENLHPMITRSKDGIQKPNPRY-ALIVSKSSFDEPKTITTAMKHPSWNAAVMDEID 899
Query: 993 ALERNNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTF 1052
+ NTW LV D + KWV++ K K DGTID+ KARLV KG+ Q EG+D+++TF
Sbjct: 900 RIHMLNTWSLVPATEDMNILTSKWVFKTKLKPDGTIDKLKARLVAKGFDQEEGVDYLETF 959
Query: 1053 SPVAKMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQG-MNSDKPNQVCL 1111
SPV + T+R++L + W L QLDV NAFLH +L E ++M P G ++ +KPN VC
Sbjct: 960 SPVVRTATIRLVLDTATANEWPLKQLDVSNAFLHGELQEPVFMFQPSGFVDPNKPNHVCR 1019
Query: 1112 LQKSLYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLA 1171
L K+LYGLKQA R WF T L GF S +D +L+V G LLLYVDD+LL
Sbjct: 1020 LTKALYGLKQAPRAWFDTFSNFLLDFGFECSTSDPSLFVCHQ-NGQSLILLLYVDDILLT 1078
Query: 1172 GNDMDEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLG 1231
G+D + + +L+ +F +KDLG ++FLG+ I G+ L+Q YA ++L +G+
Sbjct: 1079 GSDQLLMDKLLQALNNRFSMKDLGPPRYFLGIEIESYNNGLFLHQHAYASDILHQAGMTE 1138
Query: 1232 GKSATTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPT 1291
TP+ Q L + P + + +R L G+L YLT TRPDI Y VN + Q + APT
Sbjct: 1139 CNPMPTPLP--QHLEDLNSEPFEEPTYFRSLAGKLQYLTITRPDIQYAVNFICQRMHAPT 1196
Query: 1292 DMHEAAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSS 1351
+ R+LRY+KG+ GL + VL+ F DSD AGC DTR+S TG+C+ LGS+
Sbjct: 1197 NSDFGLLKRILRYVKGTINMGLPIRKHHNPVLSGFCDSDYAGCKDTRRSTTGFCILLGST 1256
Query: 1352 LVSWRSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMH 1411
L+SW +K+Q T S SS EAEYRA++ T E+ W+ LL+DL +SQ P +FCDN SA++
Sbjct: 1257 LISWSAKRQPTISHSSTEAEYRALSDTAREITWISSLLRDLGISQHQPTRVFCDNLSAVY 1316
Query: 1412 IAHNPSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSK 1471
++ NP+ H+R+KH + D H +RE++ GL+ + +++QLAD+FTK L PF + +K
Sbjct: 1317 LSANPALHKRSKHFDKDFHYIRERVALGLIETQHIPATIQLADVFTKSLPRRPFITLRAK 1376
Query: 1472 L 1472
L
Sbjct: 1377 L 1377
>UniRef100_Q6L3N8 Putative gag-pol polyprotein [Solanum demissum]
Length = 1333
Score = 630 bits (1624), Expect = e-178
Identities = 409/1239 (33%), Positives = 633/1239 (51%), Gaps = 160/1239 (12%)
Query: 271 SQSNSGYQSNSGYQSGSGNYHSNSGRNRY-------SNKKCSYYGKMGHTVEDCYKKHGF 323
+++++G G G G S GRN+ SN +C Y K GH DC+ K
Sbjct: 220 AENSAGRGHGRGNFRGRGRGGSGRGRNQVGEFRQYKSNIQCRYCKKFGHKEVDCWTKQ-- 277
Query: 324 PPGFKFKNPKYANRSANLVQTTKEDQESTSQGNTQEQEAARFGFTADQYHHLLLLLPPSE 383
K + AN Q +E+ S+
Sbjct: 278 ---------KDEQKDANFTQNVEEE---------------------------------SK 295
Query: 384 QKNSTSQHTASINSCVQMLPGKNGNSISQISTKNGNPLDTTWILDTGATDHICNTLSYFS 443
++SQ T S N+ W +D+G ++H+ ++ S F
Sbjct: 296 LFMASSQITESANA--------------------------VWFIDSGCSNHMSSSKSLFR 329
Query: 444 SYKHVAPIPVSLPN---------GNIESATIKGSIQLSPSFILINVLFLPNFEFNLISVH 494
V L + G +E T++G+++ L +V ++P NL+SV
Sbjct: 330 DLDESQKSEVRLGDDKQVHIEGKGTVEIKTVQGNVKF-----LYDVQYVPTLAHNLLSVG 384
Query: 495 KLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRS-LYILNNDSLSPFPSCNSVSTCKS 553
+L+ S + + F D+ C I+D + + I ++ ++ L+ ++ NS K
Sbjct: 385 QLMTS-GYSVVFYDNACDIKDKESGRTIARVPMTQNKMFPLDISNVG-----NSALVVKE 438
Query: 554 QNLVCEFSPSVQNLWHYRLGH--PSFVKGQSIKDLFPYVQYTQD-HVCEVCPIAKQKRLK 610
+N NLWH R GH +++K KD+ + ++ +CE C KQ R
Sbjct: 439 KNET--------NLWHLRYGHLNVNWLKLLVQKDMVIGLPNIKELDLCEGCIYGKQTRKS 490
Query: 611 FPLSNS--TSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLKSKDEVQT 668
FP+ S + C+ +++H D+ GP+ + SL G YFL DDYSR++WVY LK K E
Sbjct: 491 FPVGKSWRATTCL-ELVHADLCGPMKMESLGGSRYFLMFTDDYSRFSWVYFLKFKSETFE 549
Query: 669 LVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYA---EKGIIHHTSCVETPQQNSIVER 725
K F AFV NQ G +K +R+D G EF+ + F E GI + TP+QN + ER
Sbjct: 550 TFKKFKAFVENQSGNKIKSLRTDRGGEFLSNDFNLFCEENGIRRELTAPYTPEQNGVAER 609
Query: 726 KHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQCPFQLLHKQLPDITF 785
K++ ++ +AR+ L LP FW A+ ++ +N PT + N P + + + P ++
Sbjct: 610 KNRTVVEMARSSLKAKGLPDYFWGEAVATVVYFLNISPTKDVWNTTPLEAWNGKKPRVSH 669
Query: 786 LKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNTRDISISRNVIFHEN 845
L++FG C A L + +K D ++ +C+F+G+ +K Y +Y+ + + ISRNV+F+E+
Sbjct: 670 LRIFG--CIAYALVNFHSKLDEKSTKCIFVGYSLQSKAYRLYNPISGKVIISRNVVFNED 727
Query: 846 IFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQPLSSGPSQNSEQPIVH 905
+ + + I LP+ D E + N P+SS S
Sbjct: 728 V-------SWNFNSGNMMSNIQLLPT---DEESAVDFGNSPNSSPVSSSVSS-------- 769
Query: 906 RVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVISYHKLDPSYHAFIMN- 964
+A ++TV P SS + PL + K +P Y +
Sbjct: 770 ------------------PIAPSTTVAPDESSVEPI--PLRRSTREKKPNPKYSNTVNTS 809
Query: 965 ---ITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIK 1021
+P Y EAV+ W+ AM EI+A+ERN+TW LVD P K IG KWV+R K
Sbjct: 810 CQFALLVSDPICYEEAVEQSEWKNAMIEEIQAIERNSTWELVDAPEGKNVIGLKWVFRTK 869
Query: 1022 YKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVD 1081
Y DG+I ++KARLV KGY+Q +G+DF +TFSPVA+ T+RV+L+L + + ++Q DV
Sbjct: 870 YNADGSIQKHKARLVAKGYSQQQGVDFDETFSPVARFETVRVVLALAAQLHLPVYQFDVK 929
Query: 1082 NAFLHAKLDEEIYMSLPQG-MNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFT 1140
+AFL+ L+EE+Y+S PQG M + N+V L+K+LYGLKQA R W+S + QG GF
Sbjct: 930 SAFLNGDLEEEVYVSQPQGFMITGNENKVYKLRKALYGLKQAPRAWYSKIDSFFQGSGFR 989
Query: 1141 QSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFF 1200
+S + TLY+KK T F + LYVDD++ G+ + K ++ + F + DLG K+F
Sbjct: 990 RSDNEPTLYLKKQGTDEFLLVCLYVDDMIYIGSSKSLVNDFKSNMMRNFEMSDLGLLKYF 1049
Query: 1201 LGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYR 1260
LGL + + + GI ++Q+KYA +LL ++ + ATTPM+ ++KL + GT ++ +R
Sbjct: 1050 LGLEVIQDKDGIFISQKKYAEDLLKKFQMMNCEVATTPMNINEKLQRADGTEKANPKLFR 1109
Query: 1261 RLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASS 1320
L+G L YLT TRPDIA+ V+ +S+FL +PT H AA RVLRY+ G+ G++Y A +
Sbjct: 1110 SLVGGLNYLTHTRPDIAFSVSVVSRFLQSPTKQHFGAAKRVLRYVAGTTDFGIWYSKAPN 1169
Query: 1321 TVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAEYRAMAATVC 1380
L F+DSD AGC+D RKS +G C GS +V+W SKKQ T + S+ EAEY A +
Sbjct: 1170 FRLVGFTDSDYAGCLDDRKSTSGSCFSFGSGVVTWSSKKQETVALSTSEAEYTAASLAAR 1229
Query: 1381 EVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHIVREKITQGL 1440
+ WL LL+D Q +F D++SA+ +A NPS+H RTKHI++ H +R + G
Sbjct: 1230 QALWLRKLLEDFSYEQKESTEIFSDSKSAIAMAKNPSFHGRTKHIDVQYHFIRTLVADGR 1289
Query: 1441 VHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHS 1479
+ L +++ Q ADIFTK L A + +L + D S
Sbjct: 1290 IVLKFCSTNEQAADIFTKSLPQAKHEYFRLQLGVCDFES 1328
>UniRef100_Q9SSB1 T18A20.5 protein [Arabidopsis thaliana]
Length = 1522
Score = 615 bits (1587), Expect = e-174
Identities = 387/1121 (34%), Positives = 571/1121 (50%), Gaps = 93/1121 (8%)
Query: 425 WILDTGATDHICNTLSYFS-SYKHVAPIPVSLPNGNIESATIKGSIQLSPS---FILINV 480
WI D+ A+ H+ N S + + + +GN T GS ++ S L V
Sbjct: 326 WIPDSAASAHVTNNRHVLQQSQPYHGSDSIMVADGNFLPITHTGSGSIASSSGKIPLKEV 385
Query: 481 LFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLS 540
L P+ +L+SV KL + F D I D K++ R LY L L
Sbjct: 386 LVCPDIVKSLLSVSKLTSDYPCSVEFDADSVRINDKATKKLLVMGRNRDGLYSLEEPKLQ 445
Query: 541 PFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPS------FVKGQSIKDLFPYVQYTQ 594
S S +WH RLGH + +SI + V+
Sbjct: 446 VLYSTRQNSASSE-------------VWHRRLGHANAEVLHQLASSKSIIIINKVVKT-- 490
Query: 595 DHVCEVCPIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRY 654
VCE C + K RL F LS + + IH D+WGP S+ GF Y++ +D YSR+
Sbjct: 491 --VCEACHLGKSTRLPFMLSTFNASRPLERIHCDLWGPSPTSSVQGFRYYVVFIDHYSRF 548
Query: 655 TWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYA---EKGIIHHT 711
TW Y LK K + + F V NQ G +KI + D G EF+ SQF + GI +
Sbjct: 549 TWFYPLKLKSDFFSTFVMFQKLVENQLGHKIKIFQCDGGGEFISSQFLKHLQDHGIQQNM 608
Query: 712 SCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDN-Q 770
SC TPQQN + ERKH+HI+ + +++FQ+ LP +W + + F+IN LPT LDN +
Sbjct: 609 SCPYTPQQNGMAERKHRHIVELGLSMIFQSKLPLKYWLESFFTANFVINLLPTSSLDNNE 668
Query: 771 CPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLN 830
P+Q L+ + P+ + L+VFG C+ + TKFD R+ +CVFLG+ KGY
Sbjct: 669 SPYQKLYGKAPEYSALRVFGCACYPTLRDYASTKFDPRSLKCVFLGYNEKYKGYRCLYPP 728
Query: 831 TRDISISRNVIFHENIFPYK-----LHRDDH----ECESSTFPQIPCLPSEPFDYEYPIP 881
T I ISR+V+F EN P++ LH D E +F + P++P YP+
Sbjct: 729 TGRIYISRHVVFDENTHPFESIYSHLHPQDKTPLLEAWFKSFHHVT--PTQPDQSRYPVS 786
Query: 882 PDNPVQIQPLSSGPSQ--------------NSEQPIVHRVSQRPRK-----QPTYLQDYH 922
+ LS+ P+ + + + VS P + + YH
Sbjct: 787 SIPQPETTDLSAAPASVAAETAGPNASDDTSQDNETISVVSGSPERTTGLDSASIGDSYH 846
Query: 923 CTLAATSTVVPANSS--SKGTSHPLSQVISYHKLDP--SYHAFIMN-------------- 964
A +S PA SS S P+ + P + HA +
Sbjct: 847 SPTADSSHPSPARSSPASSPQGSPIQMAPAQQVQAPVTNEHAMVTRGKEGISKPNKRYVL 906
Query: 965 ITTTV---EPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKTPIGCKWVYRIK 1021
+T V EP +EA+KH W AM E+ + TW LV P+ +G WV+R K
Sbjct: 907 LTHKVSIPEPKTVTEALKHPGWNNAMQEEMGNCKETETWTLVPYSPNMNVLGSMWVFRTK 966
Query: 1022 YKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVSTKNWFLHQLDVD 1081
DG++D+ KARLV KG+ Q EGID+++T+SPV + T+R++L + + W L Q+DV
Sbjct: 967 LHADGSLDKLKARLVAKGFKQEEGIDYLETYSPVVRTPTVRLILHVATVLKWELKQMDVK 1026
Query: 1082 NAFLHAKLDEEIYMSLPQG-MNSDKPNQVCLLQKSLYGLKQASRQWFSTLCQALQGLGFT 1140
NAFLH L E +YM P G ++ KP+ VCLL KSLYGLKQ+ R WF L GF
Sbjct: 1027 NAFLHGDLTETVYMRQPAGFVDKSKPDHVCLLHKSLYGLKQSPRAWFDRFSNFLLEFGFI 1086
Query: 1141 QSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKFRIKDLGEAKFF 1200
S D +L+V S+ LLLYVDD+++ GN+ + + +L+++FR+KD+G+ +F
Sbjct: 1087 CSLFDPSLFVY-SSNNDVILLLLYVDDMVITGNNSQSLTHLLAALNKEFRMKDMGQVHYF 1145
Query: 1201 LGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASSGTPLSDISSYR 1260
LG+ I G+ ++Q+KYA +LL + + TP+ ++ SD + +R
Sbjct: 1146 LGIQIQTYDGGLFMSQQKYAEDLLITASMANCSPMPTPLPLQLDRVSNQDEVFSDPTYFR 1205
Query: 1261 RLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSPGCGLFYPAASS 1320
L G+L YLT TRPDI + VN + Q + P+ R+LRYIKG+ G+ Y + SS
Sbjct: 1206 SLAGKLQYLTLTRPDIQFAVNFVCQKMHQPSVSDFNLLKRILRYIKGTVSMGIQYNSNSS 1265
Query: 1321 TV---------LTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCEAE 1371
+V L+A+SDSD A C +TR+S+ GYC F+G +++SW SKKQ T SRSS EAE
Sbjct: 1266 SVVSAYESDYDLSAYSDSDYANCKETRRSVGGYCTFMGQNIISWSSKKQPTVSRSSTEAE 1325
Query: 1372 YRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDCHI 1431
YR+++ T E++W+ +L+++ VS +FCDN SA+++ NP++H RTKH ++D H
Sbjct: 1326 YRSLSETASEIKWMSSILREIGVSLPDTPELFCDNLSAVYLTANPAFHARTKHFDVDHHY 1385
Query: 1432 VREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKL 1472
+RE++ + + + LQLADIFTK L F + KL
Sbjct: 1386 IRERVALKTLVVKHIPGHLQLADIFTKSLPFEAFTRLRFKL 1426
>UniRef100_Q9SN55 Putative retrotransposon polyprotein [Arabidopsis thaliana]
Length = 1203
Score = 614 bits (1583), Expect = e-174
Identities = 401/1080 (37%), Positives = 561/1080 (51%), Gaps = 163/1080 (15%)
Query: 425 WILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLSPSFILINVLFLP 484
WI+D+GA+ H+C+ L+ F HV+ + V+LPNG + T G+I ++ + IL NVL +P
Sbjct: 99 WIIDSGASSHVCSDLTMFRELIHVSGVTVTLPNGTRVAITHTGTICITSTLILHNVLLVP 158
Query: 485 NFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLSPFPS 544
+F+FNLISV LVK+L + F D C IQ+ MIG + +LYIL S PS
Sbjct: 159 DFKFNLISVCCLVKTLSYSAHFFADCCYIQELTRGLMIGRGKTYNNLYILETQRTSFSPS 218
Query: 545 CNSVSTCKSQNLVCEFSPSVQN---LWHYRLGHPSFVKGQSIKDLFPYVQYTQDHVCEVC 601
+ S+ F+ +VQ+ LWH RLG + Y+ Y + +
Sbjct: 219 LPAASS---------FTGTVQDDCLLWHQRLG------------IRHYLHYRNLYFLTLV 257
Query: 602 PIAKQKRLKFPLSNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRYTWVYLLK 661
+ + + N + VS N F F+ ++ +++Y
Sbjct: 258 DDCTRTTWVYMMKNKSE--------------VS----NIFPVFVKLI--FTQYNAKIKAI 297
Query: 662 SKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQFYAEKGIIHHTSCVETPQQNS 721
D V+ L F FV Q G+IH SC TPQQNS
Sbjct: 298 RSDNVKELA--FTKFVKEQ-------------------------GMIHQFSCAYTPQQNS 330
Query: 722 IVER------------KHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDN 769
+VER H +++ R ++F + + S ++ P IL
Sbjct: 331 VVERYPSGYKGYKVLDLESHSISITRNVVFHETKFPFKTSKFLKES---VDMFPNSILPL 387
Query: 770 QCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDL 829
P + + +P L+ + S AS + L T+ D+
Sbjct: 388 PAPLHFV-ESMPLDDDLRADDNNASTSNSASSASSIPP-------LPSTVNTQNTDALDI 439
Query: 830 NTRDISISRNVIFHENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYP---IPPDNPV 886
+T + I+R + + ++ C S F P+ E P IPP
Sbjct: 440 DTNSVPIAR----PKRNAKAPAYLSEYHCNSVPFLS-SLSPTTSTSIETPSSSIPPKKIT 494
Query: 887 QIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLS 946
P+S+ S + P+ H Y C
Sbjct: 495 TPYPMSTAISYDKLTPLFH--------------SYICA---------------------- 518
Query: 947 QVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKP 1006
+ ++ AF T ++ +++ A E + ALE+N TW++
Sbjct: 519 -----YNVETEPKAF----TQAMKSEKWTRAANEE---------LHALEQNKTWIVESLT 560
Query: 1007 PDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLS 1066
K +GCKWV+ IKY DG+I+RYKARLV +G+TQ EGID+M+TFSPVAK ++++LL
Sbjct: 561 EGKNVVGCKWVFTIKYNPDGSIERYKARLVAQGFTQQEGIDYMETFSPVAKFGSVKLLLG 620
Query: 1067 LVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMN-----SDKPNQVCLLQKSLYGLKQ 1121
L + W L Q+DV NAFLH +LDEEIYMSLPQG S VC L KSLYGLKQ
Sbjct: 621 LAAATGWSLTQMDVSNAFLHGELDEEIYMSLPQGYTPPTGISLPSKPVCRLLKSLYGLKQ 680
Query: 1122 ASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLV 1181
ASRQW+ L G F QS AD+T++VK S T S +L+YVDD+++A ND ++ +
Sbjct: 681 ASRQWYKRLSSVFLGANFIQSPADNTMFVKVSCT-SIIVVLVYVDDLMIASNDSSAVENL 739
Query: 1182 KHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDC 1241
K L +F+IKDLG A+FFLGL IARS +GI + QRKYA LL D GL G K ++ PMD
Sbjct: 740 KELLRSEFKIKDLGPARFFLGLEIARSSEGISVCQRKYAQNLLEDVGLSGCKPSSIPMDP 799
Query: 1242 SQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRV 1301
+ L+ GT L + +SYR L+GRLLYL TRPDI + V+ LSQFLSAPTD+H AAH+V
Sbjct: 800 NLHLTKEMGTLLPNATSYRELVGRLLYLCITRPDITFAVHTLSQFLSAPTDIHMQAAHKV 859
Query: 1302 LRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQT 1361
LRY+KG+PG GL Y A+S L FSD+D C D+R+S+TG+C++LG+SL++W+SKKQ+
Sbjct: 860 LRYLKGNPGQGLMYSASSELCLNGFSDADWGTCKDSRRSVTGFCIYLGTSLITWKSKKQS 919
Query: 1362 TTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHER 1421
SRSS E+EYR++A CE+ WL LL+DL V+ T P +FCDN+SA+H+A NP +HER
Sbjct: 920 VVSRSSTESEYRSLAQATCEIIWLQQLLKDLHVTMTCPAKLFCDNKSALHLATNPVFHER 979
Query: 1422 TKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLRLYDIHSPV 1481
TKHIE+DCH VR++I G + L V + QLADI TKPL P IFS L + SPV
Sbjct: 980 TKHIEIDCHTVRDQIKAGKLKTLHVPTGNQLADILTKPLHPVQ-SPIFSLLFRFTGTSPV 1038
>UniRef100_Q7Y141 Putative polyprotein [Oryza sativa]
Length = 1335
Score = 614 bits (1583), Expect = e-174
Identities = 395/1250 (31%), Positives = 623/1250 (49%), Gaps = 148/1250 (11%)
Query: 245 KKGSSLLKTFLGPLIAARENSNDGRSSQSNS------GYQSNSGY--QSGSGNYHSNSGR 296
++GSS+ F L +NS + Q N GY +G+ Q G G
Sbjct: 193 REGSSIENAFQSKLSFRPQNSRFRGNFQKNGFPMRDRGYFQKNGFSRQKEDGQERREKGT 252
Query: 297 NRYSNKKCSYYGKMGHTVEDCYKKHGFPPGFKFKNPKYANRSANLVQTTKEDQESTSQGN 356
+ SN C K HT + C+KK K A +T + ++ + SQ
Sbjct: 253 SS-SNLWCDICQKSSHTTDMCWKK------MTCNKCKRKGHIAKYCRTREINRANFSQEK 305
Query: 357 TQEQEAARFGFTADQYHHLLLLLPPSEQKNSTSQHTASINSCVQMLPGKNGNSISQISTK 416
+ +E S HTA
Sbjct: 306 EKSEEMV------------------------FSCHTAQEEK------------------- 322
Query: 417 NGNPLDTTWILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQLS---- 472
D W++D+G T+H+ + F + + NG+I + KG++ +
Sbjct: 323 -----DDVWVIDSGCTNHMAADPNLFREMDSSYHAKIHMGNGSIAQSEGKGTVAVQTADG 377
Query: 473 PSFILINVLFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLY 532
P FI +VL +P+ + NL+S+ +L++ + + F D C I D +++ ++
Sbjct: 378 PKFIK-DVLLVPDLKQNLLSIGQLLEH-GYAVYFEDFSCKILDRKNNRLVAKINMEKNRN 435
Query: 533 ILNNDSLSPFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSF-----VKGQSIKDLF 587
L + + + + +LWH R+GH ++ ++ + +
Sbjct: 436 FL-------------LRMNHTTQMALRSEVDISDLWHKRMGHLNYRALKLLRTKGMVQGL 482
Query: 588 PYVQYTQDHVCEVCPIAKQKRLKFPLSNS-TSDCIFQMIHVDIWGPVSVLSLNGFSYFLT 646
P++ D CE C KQ R FP S + + +++H DI G V +S G YF+T
Sbjct: 483 PFITLKSDP-CEGCVFGKQIRASFPHSGAWRASAPLELVHADIVGKVPTISEGGNWYFIT 541
Query: 647 IVDDYSRYTWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQF--YAE 704
+DDY+R WVY LK K + K F A V NQ +K++RSD G+E++ +F Y E
Sbjct: 542 FIDDYTRMIWVYFLKEKSAALEIFKKFKAMVENQSNRKIKVLRSDQGREYISKEFEKYCE 601
Query: 705 K-GIIHHTSCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLP 763
GI + + QQN + ERK++ I ++A ++L +PK FWA A+ +++++NR P
Sbjct: 602 NAGIRRQLTAGYSAQQNGVAERKNRTINDMANSMLQDKGMPKSFWAEAVNTAVYILNRSP 661
Query: 764 TPILDNQCPFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKG 823
T + N+ PF+ + + P I ++VFG +C+A A R KFD+++ RC+F+G+ G KG
Sbjct: 662 TKAVTNRTPFEAWYGKKPVIGHMRVFGCICYAQVPAQKRVKFDNKSDRCIFVGYADGIKG 721
Query: 824 YLVYDLNTRDISISRNVIFHENI-FPYKLHRDDHECESSTFPQIPCL------PSEPFDY 876
Y +Y+L + I ISR+ IF E+ + +K E+S+ P +P P +
Sbjct: 722 YRLYNLEKKKIIISRDAIFDESATWNWK------SPEASSTPLLPTTTITLGQPHMHGTH 775
Query: 877 EYPIPPDNPVQIQPLSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANS 936
E +P P+SS + + P P P ++ S V S
Sbjct: 776 EVEDHTPSPQPSSPMSSSSASSDSSPSSEEQISTPESAPRRVR---------SMVELLES 826
Query: 937 SSKGTSHPLSQVISYHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALER 996
+S+ + +Y + VEP + EA KH+ W AM EI +E+
Sbjct: 827 TSQQRGSEQHEFCNY---------------SVVEPQSFQEAEKHDNWIKAMEDEIHMIEK 871
Query: 997 NNTWLLVDKPPDKTPIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVA 1056
NNTW LVD+P D+ IG KWVY+ K DG++ +YKARLV KG+ Q GID+ +T++PVA
Sbjct: 872 NNTWELVDRPRDREVIGVKWVYKTKLNPDGSVQKYKARLVAKGFKQKPGIDYYETYAPVA 931
Query: 1057 KMTTLRVLLSLVSTKNWFLHQLDVDNAFLHAKLDEEIYMSLPQGMN-SDKPNQVCLLQKS 1115
++ T+R +++L + K W ++QLDV +AFL+ LDEEIY+ P+G + N+V L+K+
Sbjct: 932 RLETIRTIIALAAQKRWKIYQLDVKSAFLNGYLDEEIYVEQPEGFSVQGGENKVFRLKKA 991
Query: 1116 LYGLKQASRQWFSTLCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDM 1175
LYGLKQA R W+S + + GF +S ++ TLYV K+ T + LYVDD++ GN
Sbjct: 992 LYGLKQAPRAWYSQIDKYFIQKGFAKSISEPTLYVNKTGT-DILIVSLYVDDLIYTGNSE 1050
Query: 1176 DEIKLVKHSLHQKFRIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSA 1235
++ K + + + DLG +FLG+ + +S +GI ++QRKYA +L + KS
Sbjct: 1051 KMMQDFKKDMMHTYEMSDLGLLHYFLGMEVHQSDEGIFISQRKYAENILKKFKMDNCKSV 1110
Query: 1236 TTPMDCSQKLSASSGTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHE 1295
TTP+ ++K A G +D + YR L+G LLYLT TRPDI + + LS+++S+P+ ++
Sbjct: 1111 TTPLLPNEKQKARDGADKADPTIYRSLVGSLLYLTATRPDIMFAASLLSRYMSSPSQLNF 1170
Query: 1296 AAAHRVLRYIKGSPGCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSW 1355
AA RVLRYIKG+ G++Y + L ++DSD AGC+D KS +GY LGS+
Sbjct: 1171 TAAKRVLRYIKGTADYGIWYKPVKESKLIGYTDSDWAGCLDDMKSTSGYAFSLGSA---- 1226
Query: 1356 RSKKQTTTSRSSCEAEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHN 1415
EAEY A + V +V WL +++DL Q P +++CD++SA+ I+ N
Sbjct: 1227 -------------EAEYVAASKAVSQVVWLRRIMEDLGEKQYQPTTIYCDSKSAIAISEN 1273
Query: 1416 PSYHERTKHIELDCHIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPF 1465
P H+RTKHI + H +RE + + V L + QLADIFTK LS F
Sbjct: 1274 PVSHDRTKHIAIKYHYIREAVDRQEVKLEFCRTDEQLADIFTKALSKEKF 1323
>UniRef100_Q9SXB2 T28P6.8 protein [Arabidopsis thaliana]
Length = 1352
Score = 585 bits (1509), Expect = e-165
Identities = 355/1064 (33%), Positives = 575/1064 (53%), Gaps = 77/1064 (7%)
Query: 425 WILDTGATDHICNTLSYFSSYKHVAPIPVSLPNGNIESATIKGSIQL----SPSFILINV 480
W LD+GA++H+C S F+ V+L + + KG+I + + NV
Sbjct: 335 WYLDSGASNHMCGRKSMFAELDESVRGNVALGDESKMEVKGKGNILIRLKNGDHQFISNV 394
Query: 481 LFLPNFEFNLISVHKLVKSLRFRLTFSDDDCLIQDSNACKMIGTARAVRSLYILNNDSLS 540
++P+ + N++S+ +L++ + + D++ I+D + + + +++LN
Sbjct: 395 YYIPSMKTNILSLGQLLEK-GYDIRLKDNNLSIRDQESNLITKVPMSKNRMFVLNIR--- 450
Query: 541 PFPSCNSVSTCKSQNLVCEFSPSVQNLWHYRLGHPSF--VKGQSIKDL---FPYVQYTQD 595
N ++ C +C S LWH R GH +F ++ S K++ P + + +
Sbjct: 451 -----NDIAQCLK---MCYKEESW--LWHLRFGHLNFGGLELLSRKEMVRGLPCINHP-N 499
Query: 596 HVCEVCPIAKQKRLKFPL-SNSTSDCIFQMIHVDIWGPVSVLSLNGFSYFLTIVDDYSRY 654
VCE C + KQ ++ FP S+S + ++IH D+ GP+ SL +YFL +DD+SR
Sbjct: 500 QVCEGCLLGKQFKMSFPKESSSRAQKPLELIHTDVCGPIKPKSLGKSNYFLLFIDDFSRK 559
Query: 655 TWVYLLKSKDEVQTLVKDFCAFVTNQFGVSVKIVRSDNGKEFVLSQF--YAE-KGIIHHT 711
TWVY LK K EV + K F A V + G+ +K +RSD G EF +F Y E GI
Sbjct: 560 TWVYFLKEKSEVFEIFKKFKAHVEKESGLVIKTMRSDRGGEFTSKEFLKYCEDNGIRRQL 619
Query: 712 SCVETPQQNSIVERKHQHILNVARALLFQAHLPKIFWAHAIIHSIFLINRLPTPILDNQC 771
+ +PQQN +VERK++ IL +AR++L LPK WA A+ +++L+NR PT + +
Sbjct: 620 TVPRSPQQNGVVERKNRTILEMARSMLKSKRLPKELWAEAVACAVYLLNRSPTKSVSGKT 679
Query: 772 PFQLLHKQLPDITFLKVFGSLCFASTLASHRTKFDHRAKRCVFLGFKSGTKGYLVYDLNT 831
P + + P ++ L+VFGS+ A R+K D ++++ +F+G+ + +KGY +Y+ +T
Sbjct: 680 PQEAWSGRKPGVSHLRVFGSIAHAHVPDEKRSKLDDKSEKYIFIGYDNNSKGYKLYNPDT 739
Query: 832 RDISISRNVIF-HENIFPYKLHRDDHECESSTFPQIPCLPSEPFDYEYPIPPDNPVQIQP 890
+ ISRN++F E + + + +D+ + FP F+ + P P
Sbjct: 740 KKTIISRNIVFDEEGEWDWNSNEEDY----NFFPH--------FEEDEPEP--------- 778
Query: 891 LSSGPSQNSEQPIVHRVSQRPRKQPTYLQDYHCTLAATSTVVPANSSSKGTSHPLSQVIS 950
E+P S+ P PT + TS+ + +SS + Q +
Sbjct: 779 -------TREEP----PSEEPTTPPT---------SPTSSQIEESSSERTPRFRSIQEL- 817
Query: 951 YHKLDPSYHAFIMNITTTVEPTRYSEAVKHECWRVAMN*EIEALERNNTWLLVDKPPDKT 1010
Y + + + + EP + +A++ + WR AM+ EI+++++N+TW L P
Sbjct: 818 YEVTENQENLTLFCLFAECEPMDFQKAIEKKTWRNAMDEEIKSIQKNDTWELTSLPNGHK 877
Query: 1011 PIGCKWVYRIKYKQDGTIDRYKARLVVKGYTQIEGIDFMDTFSPVAKMTTLRVLLSLVST 1070
IG KWVY+ K G ++RYKARLV KGY+Q GID+ + F+PVA++ T+R+++SL +
Sbjct: 878 AIGVKWVYKAKKNSKGEVERYKARLVAKGYSQRVGIDYDEVFAPVARLETVRLIISLAAQ 937
Query: 1071 KNWFLHQLDVDNAFLHAKLDEEIYMSLPQG-MNSDKPNQVCLLQKSLYGLKQASRQWFST 1129
W +HQ+DV +AFL+ L+EE+Y+ PQG + + ++V L+K LYGLKQA R W +
Sbjct: 938 NKWKIHQMDVKSAFLNGDLEEEVYIEQPQGYIVKGEEDKVLRLKKVLYGLKQAPRAWNTR 997
Query: 1130 LCQALQGLGFTQSFADHTLYVKKSATGSFTALLLYVDDVLLAGNDMDEIKLVKHSLHQKF 1189
+ + + F + +H LY+K A L YVDD++ GN+ + K + ++F
Sbjct: 998 IDKYFKEKDFIKCPYEHALYIKIQKEDILIACL-YVDDLIFTGNNPSIFEEFKKEMTKEF 1056
Query: 1190 RIKDLGEAKFFLGLAIARSQKGIILNQRKYALELLSDSGLLGGKSATTPMDCSQKLSASS 1249
+ D+G ++LG+ + + GI + Q YA E+L + TPM+C KLS
Sbjct: 1057 EMTDIGLMSYYLGIEVKQEDNGIFITQEGYAKEVLKKFKMDDSNPVCTPMECGIKLSKKE 1116
Query: 1250 GTPLSDISSYRRLIGRLLYLTTTRPDIAYVVNQLSQFLSAPTDMHEAAAHRVLRYIKGSP 1309
D ++++ L+G L YLT TRPDI Y V +S+++ PT H AA R+LRYIKG+
Sbjct: 1117 EGEGVDPTTFKSLVGSLRYLTCTRPDILYAVGVVSRYMEHPTTTHFKAAKRILRYIKGTV 1176
Query: 1310 GCGLFYPAASSTVLTAFSDSDGAGCVDTRKSITGYCMFLGSSLVSWRSKKQTTTSRSSCE 1369
GL Y S L +SDSD G VD RKS +G+ ++G + +W SKKQ + S+CE
Sbjct: 1177 NFGLHYSTTSDYKLVGYSDSDWGGDVDDRKSTSGFVFYIGDTAFTWMSKKQPIVTLSTCE 1236
Query: 1370 AEYRAMAATVCEVQWLIYLLQDLQVSQTSPVSMFCDNQSAMHIAHNPSYHERTKHIELDC 1429
AEY A + VC WL LL++L + Q P +F DN+SA+ +A NP +H+R+KHI+
Sbjct: 1237 AEYVAATSCVCHAIWLRNLLKELSLPQEEPTKIFVDNKSAIALAKNPVFHDRSKHIDTRY 1296
Query: 1430 HIVREKITQGLVHLLPVTSSLQLADIFTKPLSPAPFRHIFSKLR 1473
H +RE +++ V L V + Q+AD FTKPL R F K+R
Sbjct: 1297 HYIRECVSKKDVQLEYVKTHDQVADFFTKPLK----RENFIKMR 1336
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.326 0.138 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,403,906,033
Number of Sequences: 2790947
Number of extensions: 100127972
Number of successful extensions: 331315
Number of sequences better than 10.0: 1965
Number of HSP's better than 10.0 without gapping: 1456
Number of HSP's successfully gapped in prelim test: 517
Number of HSP's that attempted gapping in prelim test: 321231
Number of HSP's gapped (non-prelim): 4340
length of query: 1481
length of database: 848,049,833
effective HSP length: 140
effective length of query: 1341
effective length of database: 457,317,253
effective search space: 613262436273
effective search space used: 613262436273
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0133.5


