

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0069.5
(595 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
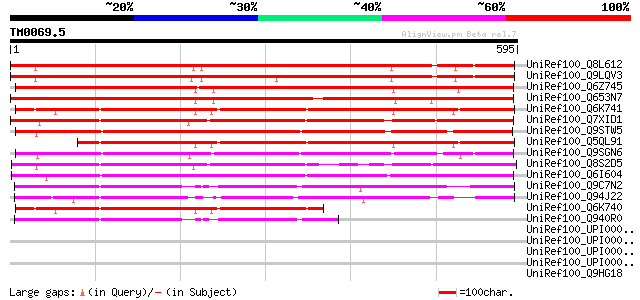
UniRef100_Q8L612 Hypothetical protein At1g14780 [Arabidopsis tha... 684 0.0
UniRef100_Q9LQV3 F10B6.18 [Arabidopsis thaliana] 669 0.0
UniRef100_Q6Z745 Hypothetical protein P0487D09.21 [Oryza sativa] 612 e-173
UniRef100_Q653N7 Hypothetical protein P0431E05.15 [Oryza sativa] 584 e-165
UniRef100_Q6K741 Hypothetical protein P0419C03.10-1 [Oryza sativa] 526 e-148
UniRef100_Q7XID1 Hypothetical protein P0039H02.131 [Oryza sativa] 504 e-141
UniRef100_Q9STW5 Hypothetical protein T22A6.120 [Arabidopsis tha... 496 e-139
UniRef100_Q5QL91 Hypothetical protein P0419C03.10-3 [Oryza sativa] 464 e-129
UniRef100_Q9SGN6 F3M18.18 [Arabidopsis thaliana] 462 e-128
UniRef100_Q8S2D5 P0401G10.6 protein [Oryza sativa] 430 e-119
UniRef100_Q6I604 Hypothetical protein OJ1214_E03.15 [Oryza sativa] 429 e-119
UniRef100_Q9C7N2 Hypothetical protein F15D2.24 [Arabidopsis thal... 401 e-110
UniRef100_Q94J22 P0481E12.26 protein [Oryza sativa] 375 e-102
UniRef100_Q6K740 Hypothetical protein P0419C03.10-2 [Oryza sativa] 354 5e-96
UniRef100_Q940R0 At1g29690/F15D2_24 [Arabidopsis thaliana] 279 1e-73
UniRef100_UPI0000439A00 UPI0000439A00 UniRef100 entry 48 0.001
UniRef100_UPI00003AFDA5 UPI00003AFDA5 UniRef100 entry 40 0.19
UniRef100_UPI0000363E83 UPI0000363E83 UniRef100 entry 38 0.74
UniRef100_UPI00003624A1 UPI00003624A1 UniRef100 entry 37 2.1
UniRef100_Q9HG18 PH-regulated protein 2 [Candida dubliniensis] 36 3.7
>UniRef100_Q8L612 Hypothetical protein At1g14780 [Arabidopsis thaliana]
Length = 627
Score = 684 bits (1765), Expect = 0.0
Identities = 356/622 (57%), Positives = 446/622 (71%), Gaps = 38/622 (6%)
Query: 2 GEGIVEKALNSLGKGFDLTSDFRLKFCK------GEERLVILNETERRELTVPGFGPVTD 55
G ++E A+ SLGKGFDLT+DFRLK+CK G++RLV+L++T+ REL +PGFG +
Sbjct: 5 GGDVIETAVKSLGKGFDLTADFRLKYCKDGDGSAGDDRLVVLDQTQNRELHIPGFGVFQN 64
Query: 56 VSVDIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAA 115
VS DI CDKG+ TR++SDIL F +MSE FNQ+SS+ GKIPSG FN FGF++GSWA+DAA
Sbjct: 65 VSADINCDKGERTRFRSDILDFNKMSEYFNQRSSVTGKIPSGNFNATFGFQSGSWATDAA 124
Query: 116 NTKFLGLDGYVIKLFNVHIDR-YPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLS 174
N K LGLD V+ LFN+HI L L+ +V AVPSSWDP LARFIE++GTH++ G+S
Sbjct: 125 NVKSLGLDASVVTLFNLHIHNPNRLRLTDRVRNAVPSSWDPQLLARFIERYGTHVITGVS 184
Query: 175 IGGKDLVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNF----LPKTKDQKH---KVPP 227
+GG+D+V+V+QD SS+L+ L+ +L +LGDQLFTG+C L K H K P
Sbjct: 185 VGGQDVVVVRQDKSSDLDNDLLRHHLYDLGDQLFTGSCLLSTRRLNKAYHHSHSQPKFPE 244
Query: 228 AFDVFGP-QIVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFI 286
AF+VF Q VAFNN + + +++GITVICAKRGGD + +HSEWL+TVP+KPDA++F+FI
Sbjct: 245 AFNVFDDKQTVAFNNFS-INSQNGITVICAKRGGDGRAKSHSEWLITVPDKPDAINFNFI 303
Query: 287 PFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRT 346
P TSLLK PG G LSHA++LYLRYKPPL DL YFLD+ R WAP+HNDLP G N
Sbjct: 304 PITSLLKDVPGSGLLSHAMSLYLRYKPPLMDLQYFLDFSGPRAWAPVHNDLPFGAAPNMA 363
Query: 347 THSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNN 406
+ P+L +N MGPKLYVNT VT K PVTGMR FLEG KCNRLAIHLQ+L NT T +
Sbjct: 364 SAYPALHINFMGPKLYVNTTPVTSEKNPVTGMRFFLEGKKCNRLAIHLQHLDNTRTTVGE 423
Query: 407 KIEDTTTW--SEEIID-DRFLEAISGKKFSHVCTAPVKYNPSW-------SSDKDVAFIV 456
KI D W S++I D DR+ E ++GKKFSHVCT PVKY+P+W S DVAFIV
Sbjct: 424 KITDEHIWRGSDQITDNDRYFEPLNGKKFSHVCTVPVKYDPNWIKTTSNHKSQNDVAFIV 483
Query: 457 TGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTS 516
TG QL VKKH S+SVLHLRL ++KVS+ VV+++W G G+ S +FS++S
Sbjct: 484 TGAQLEVKKHGSKSVLHLRLRYTKVSDHYVVQNSWVHGPI-----GTSQKSGIFSSMSMP 538
Query: 517 IQGKD------QKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLV 570
+ QK VV+DS VFP GPPVP K++KFVD SQLC+GPQ SPGHWLV
Sbjct: 539 LTSGSVHHNMIQKDKNEVVLDSGVFPGGPPVPAN-NKIVKFVDLSQLCRGPQHSPGHWLV 597
Query: 571 TGARLMLDKGKICLWAKFSLLN 592
TG RL LDKGK+CL KF+LL+
Sbjct: 598 TGVRLYLDKGKLCLHVKFALLH 619
>UniRef100_Q9LQV3 F10B6.18 [Arabidopsis thaliana]
Length = 645
Score = 669 bits (1727), Expect = 0.0
Identities = 355/640 (55%), Positives = 446/640 (69%), Gaps = 56/640 (8%)
Query: 2 GEGIVEKALNSLGKGFDLTSDFRLKFCK------GEERLVILNETERRELTVPGFGPVTD 55
G ++E A+ SLGKGFDLT+DFRLK+CK G++RLV+L++T+ REL +PGFG +
Sbjct: 5 GGDVIETAVKSLGKGFDLTADFRLKYCKDGDGSAGDDRLVVLDQTQNRELHIPGFGVFQN 64
Query: 56 VSVDIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAA 115
VS DI CDKG+ TR++SDIL F +MSE FNQ+SS+ GKIPSG FN FGF++GSWA+DAA
Sbjct: 65 VSADINCDKGERTRFRSDILDFNKMSEYFNQRSSVTGKIPSGNFNATFGFQSGSWATDAA 124
Query: 116 NTKFLGLDGYVIKLFNVHI-DRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLS 174
N K LGLD V+ LFN+HI + L L+ +V AVPSSWDP LARFIE++GTH++ G+S
Sbjct: 125 NVKSLGLDASVVTLFNLHIHNPNRLRLTDRVRNAVPSSWDPQLLARFIERYGTHVITGVS 184
Query: 175 IGGKDLVLVKQDVSSNLEPSELKKNLDELGDQLFTGTC----NFLPKTKDQKH---KVPP 227
+GG+D+V+V+QD SS+L+ L+ +L +LGDQLFTG+C L K H K P
Sbjct: 185 VGGQDVVVVRQDKSSDLDNDLLRHHLYDLGDQLFTGSCLLSTRRLNKAYHHSHSQPKFPE 244
Query: 228 AFDVF-GPQIVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFI 286
AF+VF Q VAFNN + + +++GITVICAKRGGD + +HSEWL+TVP+KPDA++F+FI
Sbjct: 245 AFNVFDDKQTVAFNNFS-INSQNGITVICAKRGGDGRAKSHSEWLITVPDKPDAINFNFI 303
Query: 287 PFTSLLKAAPGRGFLSHAINLYLRY------------------KPPLSDLPYFLDYQAHR 328
P TSLLK PG G LSHA++LYLR KPPL DL YFLD+ R
Sbjct: 304 PITSLLKDVPGSGLLSHAMSLYLRCNYSSCLILTTEMSSNTFDKPPLMDLQYFLDFSGPR 363
Query: 329 LWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCN 388
WAP+HNDLP G N + P+L +N MGPKLYVNT VT K PVTGMR FLEG KCN
Sbjct: 364 AWAPVHNDLPFGAAPNMASAYPALHINFMGPKLYVNTTPVTSEKNPVTGMRFFLEGKKCN 423
Query: 389 RLAIHLQYLLNTPTMLNNKIEDTTTW--SEEIID-DRFLEAISGKKFSHVCTAPVKYNPS 445
RLAIHLQ+L NT T + KI D W S++I D DR+ E ++GKKFSHVCT PVKY+P+
Sbjct: 424 RLAIHLQHLDNTRTTVGEKITDEHIWRGSDQITDNDRYFEPLNGKKFSHVCTVPVKYDPN 483
Query: 446 W-------SSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQGSTQR 498
W S DVAFIVTG QL VKKH S+SVLHLRL ++KVS+ VV+++W G
Sbjct: 484 WIKTTSNHKSQNDVAFIVTGAQLEVKKHGSKSVLHLRLRYTKVSDHYVVQNSWVHGPI-- 541
Query: 499 SGSGSGSGSSVFSAISTSIQGKD------QKKPALVVVDSSVFPTGPPVPVQAQKLLKFV 552
G+ S +FS++S + QK VV+DS VFP GPPVP K++KFV
Sbjct: 542 ---GTSQKSGIFSSMSMPLTSGSVHHNMIQKDKNEVVLDSGVFPGGPPVPAN-NKIVKFV 597
Query: 553 DTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
D SQLC+GPQ SPGHWLVTG RL LDKGK+CL KF+LL+
Sbjct: 598 DLSQLCRGPQHSPGHWLVTGVRLYLDKGKLCLHVKFALLH 637
>UniRef100_Q6Z745 Hypothetical protein P0487D09.21 [Oryza sativa]
Length = 620
Score = 612 bits (1577), Expect = e-173
Identities = 320/610 (52%), Positives = 421/610 (68%), Gaps = 26/610 (4%)
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGEER-LVILNETERRELTVPGFGPVTDVSVDIKCDKG 65
EKA+ LG+GFD+ D RLK+CKG ++ E LTVPG G + DV D++CDKG
Sbjct: 8 EKAVRCLGRGFDMAGDLRLKYCKGGGAGCLVERRGETTPLTVPGVGVIADVPADVRCDKG 67
Query: 66 DLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGY 125
D R++SD+L F +MSE+FNQ+SS+ GKIPSG FN F ++GSWA DA +T+ L +DGY
Sbjct: 68 DRVRFKSDVLEFNKMSELFNQRSSVEGKIPSGQFNASFDLDSGSWAHDAPHTRCLAMDGY 127
Query: 126 VIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVKQ 185
I LF++ +D L L V+ VP +WDP+A+ARFIEK+GTH++VGLS+GG+D+V VKQ
Sbjct: 128 FISLFDLRLDHRHLALDAGVLADVPPAWDPSAIARFIEKYGTHVIVGLSMGGQDVVYVKQ 187
Query: 186 DVSSNLEPSELKKNLDELGDQLFTGTCNFLP---KTKDQKHKVPPAFDVFGPQIV---AF 239
D SS+L PSE+K++LD LGDQLFTGTC P ++KD K K+P AF+VF Q+
Sbjct: 188 DKSSSLSPSEIKEHLDRLGDQLFTGTCAMPPLHCRSKD-KFKIPEAFNVFDAQVAQQRLH 246
Query: 240 NNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRG 299
+T V +K+G+TVI +KRGG+T VS+HSEWLLTVP PD ++ +P TSL++ PG G
Sbjct: 247 GITTLVSSKEGVTVIYSKRGGNTTVSSHSEWLLTVPAMPDVINVKLVPITSLIRGVPGTG 306
Query: 300 FLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGP 359
FLSHAINLYLRYKPP++DL YFLD+Q H +WAP+ +LPLGP S+R SP+L +L+G
Sbjct: 307 FLSHAINLYLRYKPPVADLRYFLDFQHHCVWAPVLGELPLGPCSHRQGSSPALHFSLLGS 366
Query: 360 KLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTML-NNKIEDTTTW--SE 416
KLYV++ +V V K PVTGMRL LEG K NRL IHLQ+L TPT + + + W +E
Sbjct: 367 KLYVSSTEVVVPKLPVTGMRLHLEGKKNNRLGIHLQHLSTTPTFVAAARADKPPVWRGTE 426
Query: 417 EIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSD----KDVAFIVTGVQLHVKKHE-SRSV 471
+ DDR+ E + + + VCTAPVKY+P W + + A +V G QLHV H+ + +V
Sbjct: 427 AVTDDRYYEPVQWRMLARVCTAPVKYDPRWCAGDRRRRPAACVVAGAQLHVVAHDAANNV 486
Query: 472 LHLRLLFSKVSNAIVVKSNWTQGSTQ-RSGSGSGSGSSVFSAISTSIQGKDQK--KP--- 525
LHLRLL+S++ VV+S W +G+ + SG S S FS ++ G +K +P
Sbjct: 487 LHLRLLYSQLPGYAVVQSKWARGAARPPSGRSSSFLSIPFSGSPSTSSGAAEKGGRPEQG 546
Query: 526 ----ALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGK 581
+ V+S VF GPPVPV AQKLLKFVDTSQ+ GPQDSPG+WLVTGARL +DKGK
Sbjct: 547 ASPVGVANVNSGVFAGGPPVPVGAQKLLKFVDTSQVTMGPQDSPGYWLVTGARLDVDKGK 606
Query: 582 ICLWAKFSLL 591
I L KFSLL
Sbjct: 607 IMLHVKFSLL 616
>UniRef100_Q653N7 Hypothetical protein P0431E05.15 [Oryza sativa]
Length = 614
Score = 584 bits (1506), Expect = e-165
Identities = 311/614 (50%), Positives = 408/614 (65%), Gaps = 34/614 (5%)
Query: 1 MGEGIVEKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRE-LTVPGFGPVTDVSVD 59
M V++A+ LG+G D+ D RLK CK E ++ E+ + VPG G V V D
Sbjct: 8 MAAAAVQRAVRCLGRGVDMAGDLRLKHCKDEGGCLVARSGEKAAAVAVPGVGVVAGVPAD 67
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKF 119
+K KGD R++SD+L F +MS++FN +SS+PGKIPSG FN+ F F + SWASDA +T+
Sbjct: 68 VKFGKGDRIRFKSDVLEFNKMSDLFNHRSSLPGKIPSGLFNSCFDFGSDSWASDAGDTRC 127
Query: 120 LGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKD 179
L DGY I L ++ +D PL L+ V+ VP++WDP+A+A FIEK+GTHI+VGLS+GG+D
Sbjct: 128 LAFDGYFISLLDLRLDCRPLALAGHVVADVPAAWDPSAIASFIEKYGTHIIVGLSMGGQD 187
Query: 180 LVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLP---KTKDQKHKVPPAFDVFGPQI 236
+V VKQD SS L PS +K++LD+LGDQLFTGTC P K++D K KVP AF+VF Q+
Sbjct: 188 VVYVKQDKSSPLSPSVIKEHLDKLGDQLFTGTCTLPPSHCKSRDHKFKVPEAFNVFDAQM 247
Query: 237 V---AFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLK 293
+ + K+G+TVI +KRGGDT SNHSEWL TVP PDA++F +P TSLLK
Sbjct: 248 TRQRIEGMTAPMSCKEGVTVIYSKRGGDTAASNHSEWLPTVPLMPDAINFKLVPITSLLK 307
Query: 294 AAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLS 353
G GFLSHAINLYLRYKPP+++L YFLD+Q HRLWAP+ +DLPLG SNR +P+L
Sbjct: 308 GVAGVGFLSHAINLYLRYKPPVAELRYFLDFQHHRLWAPVLSDLPLGLCSNRQGTNPALH 367
Query: 354 LNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIED-TT 412
+L V V K P+TGMRL LEG K NRL IHLQ+L TPT +
Sbjct: 368 FSL-----------VIVPKLPITGMRLHLEGKKNNRLGIHLQHLSTTPTFIAGGWSGRPP 416
Query: 413 TW--SEEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKD------VAFIVTGVQLHVK 464
W SE I D+R+ E + + F+HVCT PVK++P W + D A++V+G QLHVK
Sbjct: 417 AWRGSEAIADERYYEPVQRRMFAHVCTVPVKHDPRWLAAGDGGGGRPAAYVVSGAQLHVK 476
Query: 465 KHESRSVLHLRLLFSKVSNAIVVKSNWTQ-----GSTQRSGSGSGSGSSVFSAISTSIQG 519
HES SVLHLRLL++++ VV+S W G+ + SG S F++++ +
Sbjct: 477 AHESTSVLHLRLLYTELPGHSVVQSRWAHGGGGGGAARMSGVKGSFLSMSFASMAAAAAE 536
Query: 520 KDQKKPAL--VVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLML 577
K+Q+K A + VDS VF GPP PV AQ+LLKFV+TSQ+ GPQD PG+WLVTGA+L +
Sbjct: 537 KEQQKQAAARLNVDSGVFAGGPPAPVGAQRLLKFVETSQVTMGPQDCPGYWLVTGAKLDV 596
Query: 578 DKGKICLWAKFSLL 591
DKG+I L KFSLL
Sbjct: 597 DKGRISLHVKFSLL 610
>UniRef100_Q6K741 Hypothetical protein P0419C03.10-1 [Oryza sativa]
Length = 634
Score = 526 bits (1356), Expect = e-148
Identities = 287/615 (46%), Positives = 397/615 (63%), Gaps = 36/615 (5%)
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGP-------VTDVSVD 59
E+A +LG GFDLTSDFRLKF K E RLV L+E R++ VPG G + V D
Sbjct: 5 ERAAMALGAGFDLTSDFRLKFAK-EGRLVELDEAGARDVPVPGGGVGGGAAAVLRGVPRD 63
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKF 119
+ DKGD R++SD+L F QMSE+ NQKSS+ GK+PSGYFNT+F +G+W +DA TK
Sbjct: 64 VGVDKGDRIRFRSDVLEFNQMSELLNQKSSVQGKVPSGYFNTLFDL-SGAWMTDAKETKH 122
Query: 120 LGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKD 179
L DGY I L+ +H+ PLVL +V AVP WDP AL+RFI+ +GTHI+V +++GG+D
Sbjct: 123 LAFDGYFISLYKLHLKTSPLVLRDEVRSAVPPKWDPAALSRFIKTYGTHIIVEMAVGGQD 182
Query: 180 LVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLP---KTKDQKHKVPPAFDVFGPQ- 235
++ VKQ SS + ++LK +L++LGD LF+ N P KT+D K KVP F Q
Sbjct: 183 VICVKQSPSSTISSADLKLHLEDLGDFLFSDGRNHSPIHRKTRDGKSKVPDVFVRMEQQP 242
Query: 236 ----IVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSL 291
+ +++ S+ KDG+T+ C+KRGGD +++HS+WL TVP PDA+ F F+P TSL
Sbjct: 243 NNLHLSSYSESS---TKDGLTITCSKRGGDASIASHSKWLQTVPRVPDAIMFKFVPITSL 299
Query: 292 LKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPS 351
L PG G+LSHAINLYLRYKP DL +FL++Q WAP+ N+L LGP + ++ PS
Sbjct: 300 LTGIPGSGYLSHAINLYLRYKPDPEDLQHFLEFQVPLQWAPLFNELILGPQKRKGSY-PS 358
Query: 352 LSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDT 411
L +GPKL V+T++V+ +PV G+RL+LEG KCNRLAIH+Q+L + P+ML + + +
Sbjct: 359 LQFRFLGPKLQVSTSQVSSSHKPVVGLRLYLEGRKCNRLAIHVQHLSSAPSMLGDSLSSS 418
Query: 412 -TTWSE-EIIDDRFLEAISGKKFSHVCTAPVKYNPSWSS---DKDVAFIVTGVQLHVKKH 466
+ W E E + ++E I K +S VCT+ V YNP W F+VTG QL K
Sbjct: 419 MSEWRESEEVGAGYIEPIQWKSYSCVCTSKVDYNPEWLKRVPGGRGVFVVTGAQLVTKGT 478
Query: 467 ESRSVLHLRLLFSKVSNAIVVKSNWTQG-STQRSGSGSGSGSSVFSAISTSIQG------ 519
SR VLHLRL ++ V + ++ W + + GS + S+ S+ T +Q
Sbjct: 479 WSRKVLHLRLHYTHVPGCAIQRTEWAAAPAASQRGSFLTTISTTLSSPFTQLQAAAAPAA 538
Query: 520 --KDQKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLML 577
+++ PA +++S V+P GPPVP+Q++KLLKFVD S++ KGP D PGHWLVT A+L+
Sbjct: 539 PPRNEPAPA-ALLNSGVYPDGPPVPLQSRKLLKFVDMSEVVKGPHDVPGHWLVTAAKLVK 597
Query: 578 DKGKICLWAKFSLLN 592
D GKI L KF+LLN
Sbjct: 598 DGGKIGLNVKFALLN 612
>UniRef100_Q7XID1 Hypothetical protein P0039H02.131 [Oryza sativa]
Length = 608
Score = 504 bits (1299), Expect = e-141
Identities = 280/600 (46%), Positives = 386/600 (63%), Gaps = 22/600 (3%)
Query: 1 MGEGIVEKALNSLGKGFDLTSDFRLKFCKGEER----LVILNETERRELTVPGFGPVTDV 56
M + E A+ S+G G+D+ D RLK+CK L+ L+ E +++ +PG V V
Sbjct: 6 MLQSAAESAIQSIGLGYDIAHDIRLKYCKQRSSPDPLLIELDHDEVQDIVLPGGLTVAGV 65
Query: 57 SVDIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAAN 116
S IKCDKG+ TR++SD+LSF QMSE FNQ+ S+ GKIPSG FNT+F F G W DAAN
Sbjct: 66 SKSIKCDKGERTRFRSDVLSFQQMSEQFNQELSLSGKIPSGLFNTMFEF-TGCWQKDAAN 124
Query: 117 TKFLGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIG 176
TK L DG+ I L+ V + + +VL V +AVPS+W+P ALARFI KFGTH++VG+ +G
Sbjct: 125 TKSLAFDGWCITLYTVALSKAQIVLRDHVKQAVPSTWEPAALARFIRKFGTHVVVGIKMG 184
Query: 177 GKDLVLVKQDVSSNLEPSELKKNLDELGDQLF---TGTCNFLPKTKDQKHKVPPAFDVFG 233
GKD++ +KQ SS L+ +++K L E+ D+ F G +F K K K+
Sbjct: 185 GKDIIYLKQQHSSTLQAVDVQKRLKEMSDRRFLDANGQSDFSFKDSYGKDKIDTREHRL- 243
Query: 234 PQIVAFNNSTCVCAKDGITVICAKRGG-DTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLL 292
+ V + +K+ + ++ +RGG D + +HSEWL TV +PD + SFIP TSLL
Sbjct: 244 -RFVDSSPLNSYSSKEDLVMMPKRRGGRDKDILSHSEWLNTVQAEPDVISMSFIPITSLL 302
Query: 293 KAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSL 352
PG GFL+HAINLYLRYKP + +L FL++Q R WAP+++DLPLGP R + + SL
Sbjct: 303 NGVPGCGFLNHAINLYLRYKPQIEELHQFLEFQLPRQWAPVYSDLPLGPQRKRQS-TVSL 361
Query: 353 SLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTT 412
+NL+GPKLYV T V VGKRPVTG+RLFLEG + N+LAIHLQ+L + P +L + +
Sbjct: 362 PVNLIGPKLYVCTNMVDVGKRPVTGIRLFLEGKRSNKLAIHLQHLCSLPQILQLEDDPYN 421
Query: 413 TWSEEIIDDRFLEAI-SGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSV 471
+ E D ++ E I S K+FSHVCTAPV + D + IVTG QL V H + +
Sbjct: 422 DQTPEAYDRKYYEPIGSWKRFSHVCTAPV--------ESDDSSIVTGAQLEVVSHGFKKI 473
Query: 472 LHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVD 531
L LRL FSKV NA V++ +GS + SG S++ S ++ K +PA V ++
Sbjct: 474 LFLRLHFSKVCNATSVRNPEWEGSPNLA-QKSGLISTLISTHFSTAAQKPAPRPADVNIN 532
Query: 532 SSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLL 591
S+V+P GPPVPVQ KLLKFVD +++ +GPQD PG+W+V+GA+L L++GKI L K+SLL
Sbjct: 533 SAVYPGGPPVPVQTPKLLKFVDPTEMMRGPQDLPGYWVVSGAKLQLERGKISLRVKYSLL 592
>UniRef100_Q9STW5 Hypothetical protein T22A6.120 [Arabidopsis thaliana]
Length = 606
Score = 496 bits (1277), Expect = e-139
Identities = 274/594 (46%), Positives = 384/594 (64%), Gaps = 25/594 (4%)
Query: 7 EKALNSLGKGFDLTSDFRLKFCKG---EERLVILNETERR-ELTVPGFGPVTDVSVDIKC 62
E A+ S+G G+DL D RLK+CKG + RL+ + E + E+ +PG + +VS IKC
Sbjct: 12 EVAIGSIGCGYDLAIDLRLKYCKGGSKDSRLLDIKEGDDNCEIVLPGGISIPNVSKSIKC 71
Query: 63 DKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGL 122
DKG+ R++SDIL F QM+E FNQ+ S+ GKIPSG FN +F F + W DAA TK L
Sbjct: 72 DKGERMRFRSDILPFQQMAEQFNQELSLAGKIPSGLFNAMFEFSS-CWQKDAAYTKNLAF 130
Query: 123 DGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVL 182
DG I L++V +D+ ++L + V +AVPS+WDP ALARFI+ +GTHI+V + +GGKD++
Sbjct: 131 DGVFISLYSVALDKSQVLLREHVKQAVPSTWDPAALARFIDIYGTHIIVSVKMGGKDVIY 190
Query: 183 VKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVFGPQIVAFNNS 242
KQ SS L+P +L+K L E+ D+ F + + T ++ + + ++ + S
Sbjct: 191 AKQQHSSKLQPEDLQKRLKEVADKRFV-EASVVHNTGSERVQASSKVETKEQRLRFADTS 249
Query: 243 TC--VCAKDGITVICAKRGG-DTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRG 299
+ K+ +C +RGG D + H+EWL TV +PD + SFIP TSLL PG G
Sbjct: 250 SLGSYANKEDYVFMCKRRGGNDNRNLMHNEWLQTVQMEPDVISMSFIPITSLLNGVPGSG 309
Query: 300 FLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGP 359
FLSHAINLYLRYKPP+ +L FL++Q R WAP+ ++LPLGP + SL + GP
Sbjct: 310 FLSHAINLYLRYKPPIEELHQFLEFQLPRQWAPVFSELPLGP-QRKQQSCASLQFSFFGP 368
Query: 360 KLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWSEEII 419
KLYVNT V VGKRP+TGMRL+LEG + NRLAIHLQ+L + P + + + + +E
Sbjct: 369 KLYVNTTPVDVGKRPITGMRLYLEGRRSNRLAIHLQHLSSLPKIYQLEDDLNRSIRQESH 428
Query: 420 DDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFS 479
D R+ E ++ K +SHVCT PV+ SD D++ +VTG QLHV+ H ++VL LRL FS
Sbjct: 429 DRRYYEKVNWKNYSHVCTEPVE------SDDDLS-VVTGAQLHVESHGFKNVLFLRLCFS 481
Query: 480 KVSNAIVVK-SNWTQ--GSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFP 536
+V A +VK S W + G +SG S S F+A K +PA V ++S+++P
Sbjct: 482 RVVGATLVKNSEWDEAVGFAPKSGLISTLISHHFTAAQ-----KPPPRPADVNINSAIYP 536
Query: 537 TGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSL 590
GPPVP QA KLLKFVDTS++ +GPQ+SPG+W+V+GARL+++KGKI L K+SL
Sbjct: 537 GGPPVPTQAPKLLKFVDTSEMTRGPQESPGYWVVSGARLLVEKGKISLKVKYSL 590
>UniRef100_Q5QL91 Hypothetical protein P0419C03.10-3 [Oryza sativa]
Length = 551
Score = 464 bits (1193), Expect = e-129
Identities = 247/535 (46%), Positives = 347/535 (64%), Gaps = 28/535 (5%)
Query: 80 MSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYVIKLFNVHIDRYPL 139
MSE+ NQKSS+ GK+PSGYFNT+F +G+W +DA TK L DGY I L+ +H+ PL
Sbjct: 1 MSELLNQKSSVQGKVPSGYFNTLFDL-SGAWMTDAKETKHLAFDGYFISLYKLHLKTSPL 59
Query: 140 VLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVKQDVSSNLEPSELKKN 199
VL +V AVP WDP AL+RFI+ +GTHI+V +++GG+D++ VKQ SS + ++LK +
Sbjct: 60 VLRDEVRSAVPPKWDPAALSRFIKTYGTHIIVEMAVGGQDVICVKQSPSSTISSADLKLH 119
Query: 200 LDELGDQLFTGTCNFLP---KTKDQKHKVPPAFDVFGPQ-----IVAFNNSTCVCAKDGI 251
L++LGD LF+ N P KT+D K KVP F Q + +++ S+ KDG+
Sbjct: 120 LEDLGDFLFSDGRNHSPIHRKTRDGKSKVPDVFVRMEQQPNNLHLSSYSESS---TKDGL 176
Query: 252 TVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRY 311
T+ C+KRGGD +++HS+WL TVP PDA+ F F+P TSLL PG G+LSHAINLYLRY
Sbjct: 177 TITCSKRGGDASIASHSKWLQTVPRVPDAIMFKFVPITSLLTGIPGSGYLSHAINLYLRY 236
Query: 312 KPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVG 371
KP DL +FL++Q WAP+ N+L LGP + ++ PSL +GPKL V+T++V+
Sbjct: 237 KPDPEDLQHFLEFQVPLQWAPLFNELILGPQKRKGSY-PSLQFRFLGPKLQVSTSQVSSS 295
Query: 372 KRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDT-TTWSE-EIIDDRFLEAISG 429
+PV G+RL+LEG KCNRLAIH+Q+L + P+ML + + + + W E E + ++E I
Sbjct: 296 HKPVVGLRLYLEGRKCNRLAIHVQHLSSAPSMLGDSLSSSMSEWRESEEVGAGYIEPIQW 355
Query: 430 KKFSHVCTAPVKYNPSWSS---DKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIV 486
K +S VCT+ V YNP W F+VTG QL K SR VLHLRL ++ V +
Sbjct: 356 KSYSCVCTSKVDYNPEWLKRVPGGRGVFVVTGAQLVTKGTWSRKVLHLRLHYTHVPGCAI 415
Query: 487 VKSNWTQG-STQRSGSGSGSGSSVFSAISTSIQG--------KDQKKPALVVVDSSVFPT 537
++ W + + GS + S+ S+ T +Q +++ PA +++S V+P
Sbjct: 416 QRTEWAAAPAASQRGSFLTTISTTLSSPFTQLQAAAAPAAPPRNEPAPA-ALLNSGVYPD 474
Query: 538 GPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
GPPVP+Q++KLLKFVD S++ KGP D PGHWLVT A+L+ D GKI L KF+LLN
Sbjct: 475 GPPVPLQSRKLLKFVDMSEVVKGPHDVPGHWLVTAAKLVKDGGKIGLNVKFALLN 529
>UniRef100_Q9SGN6 F3M18.18 [Arabidopsis thaliana]
Length = 612
Score = 462 bits (1188), Expect = e-128
Identities = 270/602 (44%), Positives = 359/602 (58%), Gaps = 31/602 (5%)
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGE---ERLVILNETERRELTVPGFGPVTDVSVDIKCD 63
EKA++ +G G+DL SD R CK RLV ++ T R+L PG V +VS IKCD
Sbjct: 16 EKAVSVIGLGYDLCSDVRFSACKTTPDGSRLVEIDPTRNRDLIFPGGIVVNNVSSSIKCD 75
Query: 64 KGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLD 123
KG+ TR +SDILSF QMSE FNQ + GKIPSG FN +F F W DA++ K L D
Sbjct: 76 KGERTRLRSDILSFNQMSEKFNQDMCLSGKIPSGMFNNMFAFSK-CWPKDASSVKTLAYD 134
Query: 124 GYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLV 183
G+ I L++V I R L L +V VPSSWD ALA FIEK+GTH++VG+++GGKD++ V
Sbjct: 135 GWFISLYSVEIVRKQLTLRDEVKREVPSSWDSAALAGFIEKYGTHVVVGVTMGGKDVIHV 194
Query: 184 KQDVSSNLEPSELKKNLDELGDQLFT-------GTCNFLPKTKDQKHKVPPAFDVFGPQI 236
KQ SN EP E++K L GD+ F + +++ + FG +
Sbjct: 195 KQMRKSNHEPEEIQKMLKHWGDERFCVDPVESKSPASVYSGKPKEENLLQWGLQPFGTSV 254
Query: 237 VAFNNSTCVCAK-DGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAA 295
+++ + K + I +C +RGG +H WL TV P+ + F+P TSLL
Sbjct: 255 ---SSAVVMHTKNEEIMRVCIRRGGVDLGQSHERWLSTVSQAPNVISMCFVPITSLLSGL 311
Query: 296 PGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLN 355
PG GFLSHA+NLYLRYKPP+ +L FL++Q R WAP++ DLPLG + SPSL +
Sbjct: 312 PGTGFLSHAVNLYLRYKPPIEELHQFLEFQLPRQWAPVYGDLPLG-LRRSKQSSPSLQFS 370
Query: 356 LMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWS 415
LMGPKLYVNT+KV G+RPVTG+R FLEG K N LAIHLQ+L P L+ +DT
Sbjct: 371 LMGPKLYVNTSKVDSGERPVTGLRFFLEGKKGNHLAIHLQHLSACPPSLHLSHDDTYEPI 430
Query: 416 EEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLR 475
EE ++ + + FSHVCT PV+YN + S D A IVT L VK R VL LR
Sbjct: 431 EEPVEKGYYVPVKWGIFSHVCTYPVQYNGARSD--DTASIVTKAWLEVKGMGMRKVLFLR 488
Query: 476 LLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPAL------VV 529
L FS ++A+ KS W ST SG VFS IST + PA +
Sbjct: 489 LGFSLDASAVTRKSCWDNLSTNSRKSG------VFSMISTRLSTGLSPNPATTKPQSKID 542
Query: 530 VDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFS 589
++S+V+P GP PV+ KLL VDT ++ +GP++ PG+W+VTGA+L ++ GKI + AK+S
Sbjct: 543 INSAVYPRGPSPPVK-PKLLSLVDTKEVMRGPEEQPGYWVVTGAKLCVEAGKISIKAKYS 601
Query: 590 LL 591
LL
Sbjct: 602 LL 603
>UniRef100_Q8S2D5 P0401G10.6 protein [Oryza sativa]
Length = 573
Score = 430 bits (1105), Expect = e-119
Identities = 253/602 (42%), Positives = 357/602 (59%), Gaps = 58/602 (9%)
Query: 3 EGIVEKALNSLGKGFDLTSDFRLKFCKG----EERLVILNETERRELTVPGFGPVTDVSV 58
+ + E A+ S+G G+D+ +D RLK CK + L+ L+ + +++ +PG VT VS
Sbjct: 8 QSVSESAIRSIGLGYDIANDIRLKNCKQRGSPDPLLIELDHDKVQDIVLPGNLTVTGVSK 67
Query: 59 DIKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTK 118
IKCDKG+ R++SD+LSF QMSE FN++ S+ GKIPSG+FN +F F G W DA+ TK
Sbjct: 68 SIKCDKGERMRFRSDVLSFQQMSEQFNRELSLSGKIPSGFFNAMFEF-TGCWQKDASITK 126
Query: 119 FLGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGK 178
L DG+ I L+ V + + ++L V +AVPS+W+P ALARFI+KFGTHI+VG+ +GGK
Sbjct: 127 SLAFDGWCITLYTVALSKAHIILKDHVKQAVPSTWEPAALARFIKKFGTHIVVGVKMGGK 186
Query: 179 DLVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNF----LPKTKDQKHKVPPAFDVFGP 234
D++ +KQ SS+L+ +++K L E+ DQ F L + + +KV
Sbjct: 187 DVIYLKQQHSSSLQAVDVQKRLKEMSDQRFLDANGHSDISLADSYAKDNKVEAREQRL-- 244
Query: 235 QIVAFNNSTCVCAKDGITVICAKRGG-DTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLK 293
+ V N + + + ++ +RGG D + +HSEWL TV +PD + SFIP TSLL
Sbjct: 245 RFVESNPLNSYSSNEELVMMPKRRGGRDKDIISHSEWLNTVQAEPDVISMSFIPITSLLN 304
Query: 294 AAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLS 353
PG GFL+HAINLYLRYKP + +L FL++Q R WAP+++DLPLGP R + S SL
Sbjct: 305 GVPGCGFLNHAINLYLRYKPRVEELHQFLEFQLPRQWAPVYSDLPLGPQRKRQS-SASLP 363
Query: 354 LNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTT 413
+NL+GPKLYV C + I L+ P +I
Sbjct: 364 VNLIGPKLYV-----------------------CTNMIIQLEDDTYNPQTPEAEIR---- 396
Query: 414 WSEEIIDDRFLEAI-SGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVL 472
++ E I S K+FSHVCTAPV D D + IVTG L V H + +L
Sbjct: 397 --------KYYEPIGSWKRFSHVCTAPV--------DSDDSSIVTGAHLEVVSHGFKKIL 440
Query: 473 HLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDS 532
LRL FSKV NA VK+ GS G SG S++ S ++ K +PA V ++S
Sbjct: 441 FLRLHFSKVCNATSVKNPEWDGS-PNLGQKSGLISTLISTHFSTAALKPAPRPAEVNINS 499
Query: 533 SVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
+V+P GPPVPVQ KLL+FVDT+++ +GPQD PG+W+V+GA+L L++GKI L K+SLL
Sbjct: 500 AVYPGGPPVPVQTPKLLRFVDTTEMLRGPQDLPGYWVVSGAKLHLERGKISLRVKYSLLT 559
Query: 593 TS 594
+
Sbjct: 560 VN 561
>UniRef100_Q6I604 Hypothetical protein OJ1214_E03.15 [Oryza sativa]
Length = 628
Score = 429 bits (1104), Expect = e-119
Identities = 256/603 (42%), Positives = 351/603 (57%), Gaps = 18/603 (2%)
Query: 3 EGIVEKALNSLGKGFDLTSDFRLKFCKGEERLVILNETE---RRELTVPGFGPVTDVSVD 59
+ E A+ ++G G+DLTSD RL K RLV ++ RREL +P V V V
Sbjct: 21 QAAAEAAVGAIGCGYDLTSDLRLSRVKAGGRLVDIDGASGAARRELVLPWGAVVGGVPVG 80
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKF 119
I DKG+ TR++SD+LSF QM+E NQ S+ GKIPSG FN +F + G W DAA T
Sbjct: 81 IVADKGERTRFRSDVLSFAQMAEQVNQTMSVAGKIPSGAFNAMFDYH-GCWHKDAAATGS 139
Query: 120 LGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKD 179
L DG I+L+ V R L L +V VP WDP ALA FI+K+GTH++ G+ +GGKD
Sbjct: 140 LCFDGRFIELYAVEAPRAHLALLDRVKRDVPPFWDPAALAEFIDKYGTHVIAGVKMGGKD 199
Query: 180 LVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVF-GPQIVA 238
+V +KQ SNL S+++ L +L D +D K + + GP A
Sbjct: 200 VVCIKQLKGSNLTQSDVQSRLKKLSDDKLAQDSPESLTARDDKFLLGLNGSLLLGPGSAA 259
Query: 239 FNN--STCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAP 296
+ + + V KD I I +RGG HS WL T+ PD + +F+P TSLL
Sbjct: 260 WRSFRPSVVSHKDDILSIHIRRGGVDNGQGHSNWLSTISGSPDVISMAFVPITSLLTGVR 319
Query: 297 GRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLP--LGPVSNRTTHSPSLSL 354
G GFL+HA+NLYLRYKPP+ +L FL++Q R WAP +LP LGP + + PSL
Sbjct: 320 GCGFLNHAVNLYLRYKPPIEELHQFLEFQVPRQWAPEFGELPLALGPRKKKNS-LPSLQF 378
Query: 355 NLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTW 414
LMGPKL+V TAK G RPVTG+RLFLEG K NRL +HLQ+L TP + E +
Sbjct: 379 TLMGPKLHVTTAKADSGNRPVTGIRLFLEGKKNNRLGVHLQHLSATPGTITIAGEVASAE 438
Query: 415 SEEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHL 474
+ + ++E I SHVCTAPV+YN + D A IVT L V++ + VL L
Sbjct: 439 DATVRERDYIEPIKSPLLSHVCTAPVQYN--GARIDDCAAIVTRAWLEVQETCLKKVLFL 496
Query: 475 RLLFSKVSNAIVVKSNWTQG--STQRSGSGSGSGSSVFSAIST--SIQGKDQKKPA--LV 528
RL FS V++ + +S W +++SGS S S+ SA S Q Q++P V
Sbjct: 497 RLGFSGVASTKIRRSEWDGPFVVSRKSGSLSALFSARLSAAGAGGSAQMMQQQQPVGEKV 556
Query: 529 VVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKF 588
V+S++FP GPPVP+ Q++ ++VDT+++ +GP D PG+W+VTGA+L ++ GK+ L K+
Sbjct: 557 EVNSAIFPKGPPVPLPVQRMARYVDTTEVMRGPADLPGYWVVTGAKLCIEGGKVALKVKY 616
Query: 589 SLL 591
SLL
Sbjct: 617 SLL 619
>UniRef100_Q9C7N2 Hypothetical protein F15D2.24 [Arabidopsis thaliana]
Length = 561
Score = 401 bits (1031), Expect = e-110
Identities = 240/596 (40%), Positives = 335/596 (55%), Gaps = 65/596 (10%)
Query: 6 VEKALNSLGKGFDLTSDFRLKFCKGE--ERLVILNETERRELTVPGFGPVTDVSVDIKCD 63
+ A+ +LG+GFD+TSD RL +CKG RLV + E + R+L + + +V DI C
Sbjct: 21 LRNAIQALGRGFDVTSDVRLLYCKGAPGSRLVRIEEGQNRDLELSHGFLLPNVPADIDCS 80
Query: 64 KGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLD 123
+G+ + + SF +M+E FN +S + G IP G FN +F + GSW DAA+TK L L
Sbjct: 81 RGNSGTQRISVCSFHEMAEEFNVRSGVKGNIPLGCFNAMFNY-TGSWQVDAASTKSLALV 139
Query: 124 GYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLV 183
GY I L++V + + LVL ++ AVPSSWDP +LA FIE +GTHI+ ++IGG+D+V +
Sbjct: 140 GYFIPLYDVKLAKLTLVLHNEIRRAVPSSWDPASLASFIENYGTHIVTSVTIGGRDVVYI 199
Query: 184 KQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVFGPQIVAFNNST 243
+Q SS L SE++ ++++ K + H+ + + GP
Sbjct: 200 RQHQSSPLPVSEIENYVNDM--------------IKHRFHEA-ESQSITGP--------- 235
Query: 244 CVCAKD-GITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLS 302
+ KD ITVI +RGGD +H+ W TVP PD ++ +F P SLL+ PG L+
Sbjct: 236 -LKYKDKDITVIFRRRGGDDLEQSHARWAETVPAAPDIINMTFTPIVSLLEGVPGLRHLT 294
Query: 303 HAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLY 362
AI LYL YKPP+ DL YFLDYQ R WAP ++L + SL +LMGPKL+
Sbjct: 295 RAIELYLEYKPPIEDLQYFLDYQIARAWAPEQSNL-----QRKEPVCSSLQFSLMGPKLF 349
Query: 363 VNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIED-----TTTW-SE 416
++ +VTVG++PVTG+RL LEG K NRL+IHLQ+L++ P +L + W
Sbjct: 350 ISADQVTVGRKPVTGLRLSLEGSKQNRLSIHLQHLVSLPKILQPHWDSHVPIGAPKWQGP 409
Query: 417 EIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRL 476
E D R+ E I K FSHV T+P+++ + D IVTG QL V S++VLHL+L
Sbjct: 410 EEQDSRWFEPIKWKNFSHVSTSPIEHTETHIGDLSGVHIVTGAQLGVWDFGSKNVLHLKL 469
Query: 477 LFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFP 536
LFSKV + +S W SG G S S+
Sbjct: 470 LFSKVPGCTIRRSVWDHTPVASSGRLEPGGPSTSSSTE---------------------- 507
Query: 537 TGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
V Q+ KL K VD+S++ KGPQD PGHWLVTGA+L ++KGKI L K+SLLN
Sbjct: 508 ---EVSGQSGKLAKIVDSSEMLKGPQDLPGHWLVTGAKLGVEKGKIVLRVKYSLLN 560
>UniRef100_Q94J22 P0481E12.26 protein [Oryza sativa]
Length = 553
Score = 375 bits (963), Expect = e-102
Identities = 234/607 (38%), Positives = 331/607 (53%), Gaps = 82/607 (13%)
Query: 6 VEKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDIKCDKG 65
+E AL ++G+G D D RL +CKG RL++L+E+ R+LT+ G G + V D+ ++G
Sbjct: 8 LEAALQAVGRGLDAAGDHRLLYCKGTGRLLMLDESRARDLTING-GVLRGVPPDVVVEEG 66
Query: 66 D--LTRYQSD---------ILSFTQMSEMFNQKSSI-PGKIPSGYFNTVFGFEAGSWASD 113
L R + + SF +M+E FN+K+ + +P G FN++F F GSW +D
Sbjct: 67 HGILERIRQVPGPPTDEPVVCSFPKMAECFNRKAGLLETTVPLGSFNSLFSF-TGSWKND 125
Query: 114 AANTKFLGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGL 173
A TK L +DGY + LF V I L L + V A+P SWDP+ALA FIE +GTHI+ +
Sbjct: 126 EAATKSLAIDGYSVPLFKVKITSGELFLHESVKRAIPHSWDPSALASFIENYGTHIITSV 185
Query: 174 SIGGKDLVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVFG 233
++GGKD V +KQ SS L E + + E+G + F+ D K P
Sbjct: 186 TVGGKDEVYIKQHSSSQLSELEFRNYVKEIGSERFS--------DGDSKLNATP------ 231
Query: 234 PQIVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLK 293
N S + +TVI +RGG V N ++W+ TV + PD + +F+P SL+
Sbjct: 232 -----INYS-----EKDMTVIFRRRGGCDLVQNFNDWIKTVQSAPDVIGMTFLPIVSLVG 281
Query: 294 AAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTH--SPS 351
PG+ L+ AI LYL+YKP + +L YFLD+Q +WAP+ P G PS
Sbjct: 282 DMPGKKHLARAIELYLKYKPQIEELQYFLDFQVQLVWAPV----PPGIAGQHRKEPVCPS 337
Query: 352 LSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDT 411
L +LMGPKL+V+T +++VG+RPVTG++L LEG K NRLAIHLQ+L + P + +
Sbjct: 338 LQFSLMGPKLFVSTEQISVGRRPVTGLKLCLEGAKQNRLAIHLQHLGSLPKIFVPHWDSH 397
Query: 412 TT-----W-SEEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKK 465
T W E D R+ E I + F+HV TAP++Y + +D +IVTG QL V
Sbjct: 398 ITIGPPKWQGPEEQDSRWFEPIKWRNFAHVSTAPIEYTETSITDLSGVYIVTGAQLGVWD 457
Query: 466 HESRSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKP 525
++SVLHL+LLFS+V + +S W S S S V S D
Sbjct: 458 FGAKSVLHLKLLFSRVPGCTIRRSVWDH---------SPSSSLVHRTDEASSSSSDN--- 505
Query: 526 ALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLW 585
KL+K VD ++ KGPQD+PGHWLVTGA+L ++KGKI +
Sbjct: 506 --------------------AKLVKIVDMTETLKGPQDAPGHWLVTGAKLGVEKGKIVVR 545
Query: 586 AKFSLLN 592
AK+SLLN
Sbjct: 546 AKYSLLN 552
>UniRef100_Q6K740 Hypothetical protein P0419C03.10-2 [Oryza sativa]
Length = 406
Score = 354 bits (908), Expect = 5e-96
Identities = 186/377 (49%), Positives = 252/377 (66%), Gaps = 21/377 (5%)
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGP-------VTDVSVD 59
E+A +LG GFDLTSDFRLKF K E RLV L+E R++ VPG G + V D
Sbjct: 5 ERAAMALGAGFDLTSDFRLKFAK-EGRLVELDEAGARDVPVPGGGVGGGAAAVLRGVPRD 63
Query: 60 IKCDKGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKF 119
+ DKGD R++SD+L F QMSE+ NQKSS+ GK+PSGYFNT+F +G+W +DA TK
Sbjct: 64 VGVDKGDRIRFRSDVLEFNQMSELLNQKSSVQGKVPSGYFNTLFDL-SGAWMTDAKETKH 122
Query: 120 LGLDGYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKD 179
L DGY I L+ +H+ PLVL +V AVP WDP AL+RFI+ +GTHI+V +++GG+D
Sbjct: 123 LAFDGYFISLYKLHLKTSPLVLRDEVRSAVPPKWDPAALSRFIKTYGTHIIVEMAVGGQD 182
Query: 180 LVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLP---KTKDQKHKVPPAFDVFGPQ- 235
++ VKQ SS + ++LK +L++LGD LF+ N P KT+D K KVP F Q
Sbjct: 183 VICVKQSPSSTISSADLKLHLEDLGDFLFSDGRNHSPIHRKTRDGKSKVPDVFVRMEQQP 242
Query: 236 ----IVAFNNSTCVCAKDGITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSL 291
+ +++ S+ KDG+T+ C+KRGGD +++HS+WL TVP PDA+ F F+P TSL
Sbjct: 243 NNLHLSSYSESS---TKDGLTITCSKRGGDASIASHSKWLQTVPRVPDAIMFKFVPITSL 299
Query: 292 LKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPS 351
L PG G+LSHAINLYLRYKP DL +FL++Q WAP+ N+L LGP + ++ PS
Sbjct: 300 LTGIPGSGYLSHAINLYLRYKPDPEDLQHFLEFQVPLQWAPLFNELILGPQKRKGSY-PS 358
Query: 352 LSLNLMGPKLYVNTAKV 368
L +GPKL V+T++V
Sbjct: 359 LQFRFLGPKLQVSTSQV 375
>UniRef100_Q940R0 At1g29690/F15D2_24 [Arabidopsis thaliana]
Length = 383
Score = 279 bits (714), Expect = 1e-73
Identities = 158/384 (41%), Positives = 227/384 (58%), Gaps = 34/384 (8%)
Query: 6 VEKALNSLGKGFDLTSDFRLKFCKGE--ERLVILNETERRELTVPGFGPVTDVSVDIKCD 63
+ A+ +LG+GFD+TSD RL +CKG RLV + E + R+L + + +V DI C
Sbjct: 21 LRNAIQALGRGFDVTSDVRLLYCKGAPGSRLVRIEEGQNRDLELSHGFLLPNVPADIDCS 80
Query: 64 KGDLTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLD 123
+G+ + + SF +M+E FN +S + G IP G FN +F + GSW DAA+TK L L
Sbjct: 81 RGNSGTQRISVCSFHEMAEEFNVRSGVKGNIPLGCFNAMFNY-TGSWQVDAASTKSLALV 139
Query: 124 GYVIKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLV 183
GY I L++V + + LVL ++ AVPSSWDP +LA FIE +GTHI+ ++IGG+D+V +
Sbjct: 140 GYFIPLYDVKLAKLTLVLHNEIRRAVPSSWDPASLASFIENYGTHIVTSVTIGGRDVVYI 199
Query: 184 KQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVFGPQIVAFNNST 243
+Q SS L SE++ ++++ K + H+ + + GP
Sbjct: 200 RQHQSSPLPVSEIENYVNDM--------------IKHRFHEA-ESQSITGP--------- 235
Query: 244 CVCAKD-GITVICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLS 302
+ KD ITVI +RGGD +H+ W TVP PD ++ +F P SLL+ PG L+
Sbjct: 236 -LKYKDKDITVIFRRRGGDDLEQSHARWAETVPAAPDIINMTFTPIVSLLEGVPGLWHLT 294
Query: 303 HAINLYLRYKPPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLY 362
AI LYL YKPP+ DL YFLDYQ R WAP ++L + SL +LMGPKL+
Sbjct: 295 RAIELYLEYKPPIEDLQYFLDYQIARAWAPEQSNL-----QRKEPVCSSLQFSLMGPKLF 349
Query: 363 VNTAKVTVGKRPVTGMRLFLEGMK 386
++ +VTVG++PVTG+RL LEG K
Sbjct: 350 ISADQVTVGRKPVTGLRLSLEGSK 373
>UniRef100_UPI0000439A00 UPI0000439A00 UniRef100 entry
Length = 546
Score = 47.8 bits (112), Expect = 0.001
Identities = 44/186 (23%), Positives = 84/186 (44%), Gaps = 27/186 (14%)
Query: 146 IEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVKQDVSSNLEPSELKKN----LD 201
+ A+P +++P+A FI+++GTH + S+GG+ L++ D + +E S + + +
Sbjct: 79 LSALPITYEPSAYRLFIQRYGTHYMEEGSLGGQYRALLELDANYMMEMSRTETDFHQCIT 138
Query: 202 ELGDQLF----TGTCNFLPKT-----KDQKHKVPPAFDVFGPQIVAFNNSTCVCAKDGIT 252
+ +LF T C L KT +++ HK+P D+ G NS + G++
Sbjct: 139 RVKRRLFYKKKTTKCVKLMKTIENFSENRNHKMPIKTDIIG------GNSAYIA---GLS 189
Query: 253 VICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGF----LSHAINLY 308
++ + D +++W +V P + P L+K G L A+ Y
Sbjct: 190 LLDLE-NPDNNKQMYTKWAGSVKEFPKVIKQKLRPLHELVKEVACAGLKKVHLKRALEAY 248
Query: 309 LRYKPP 314
L + P
Sbjct: 249 LEEQSP 254
>UniRef100_UPI00003AFDA5 UPI00003AFDA5 UniRef100 entry
Length = 934
Score = 40.0 bits (92), Expect = 0.19
Identities = 74/314 (23%), Positives = 119/314 (37%), Gaps = 38/314 (12%)
Query: 13 LGKGFDLTSD------FRLKFCKGEERLVILNETERRELTVPG-----FGPVTDVSVDIK 61
+G GF + S F GE R V NET R+ VP V D D+K
Sbjct: 193 IGSGFHILSGESRGEVLGNSFNGGECRTVRRNET-RKSYRVPANLEAVSFQVIDEEDDVK 251
Query: 62 CD-KGDLTRYQSDILSFTQMSEMFNQKSSIPG---KIPSGYFNTVFGFEAGSWASDAANT 117
D DLT + + S ++S IPG K + F+ AS N+
Sbjct: 252 SDFYRDLTPLSDGDVGSSTSSHSSQRRSGIPGLFSKKRKVQITSSSSFKKAIEASHEKNS 311
Query: 118 KFLGLDGYVIKLFNVHIDRYPLVLSKQVIEAV---PSSWDPTALARFIEKFGTHILVGLS 174
F+ + VI + N + L LS ++A+ P ++ +R + FGTH
Sbjct: 312 NFIRIHK-VISVANFTMKESNLQLSDVFLKALNQLPLEYNYALYSRIFDDFGTHYYTSGK 370
Query: 175 IGGKDLVLVKQDVSSNLEPSELKKNLDELGDQLFTGTCNFLPKTKDQKHKVPPAFDVFGP 234
+GG +L Q S L+ S L ++DE + + T T + +K K +
Sbjct: 371 MGGSYDILY-QYSSEELKNSGL--SVDESMECIRTETTR---RVFFRKKKKVSTECITNK 424
Query: 235 QIVAFNNSTCVCAKDGITVI------------CAKRGGDTQVSNHSEWLLTVPNKPDAVD 282
V + S A+ ++++ K+G + + WL + + P +D
Sbjct: 425 MTVKHDGSILESAERSVSLVKGGRSEYAAALAWEKKGAFPGNTIFTNWLESTKDNPVVID 484
Query: 283 FSFIPFTSLLKAAP 296
F L+K P
Sbjct: 485 FEVSSIVDLVKNMP 498
>UniRef100_UPI0000363E83 UPI0000363E83 UniRef100 entry
Length = 794
Score = 38.1 bits (87), Expect = 0.74
Identities = 52/202 (25%), Positives = 82/202 (39%), Gaps = 33/202 (16%)
Query: 130 FNVHIDRYPLVLSKQVIEAV---PSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVKQD 186
F + RY + LS+++ +A+ PS +D A + +E+FGTH L S+GG V+ K D
Sbjct: 237 FQSNSPRY-IPLSEELWKALVKLPSVYDYAAYRQLLERFGTHYLSEGSLGGSFKVVAKID 295
Query: 187 VSSNLEPSELKKNLDELGDQLFTGTCNFLPK----TKDQKHKVPPAFDVFGPQIVAFNNS 242
+ SE + NFL + T +H+ P FD + N+
Sbjct: 296 EETEQYMSEDSST-----------SINFLREKQIFTTQNQHRFSPLFDCDRKR----NHL 340
Query: 243 TCVCAK-DGITVICAKRGGDTQVSN-----HSEWLLTVPNKPDAVDFSFIPFTSLLKAAP 296
+ V G+ I A + + N +S W +V + P P + L+K
Sbjct: 341 SRVDVDGGGVQHIAALKTMNLDDPNRNWEMYSNWADSVRSFPQVTKQKLRPLSELVKEVQ 400
Query: 297 GRG----FLSHAINLYLRYKPP 314
G +L AI YL P
Sbjct: 401 CAGVKKIYLRRAIEQYLSESAP 422
>UniRef100_UPI00003624A1 UPI00003624A1 UniRef100 entry
Length = 174
Score = 36.6 bits (83), Expect = 2.1
Identities = 26/79 (32%), Positives = 37/79 (45%), Gaps = 8/79 (10%)
Query: 469 RSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALV 528
+ ++ LR++ +K+ NA + Q S+ GSGSGSGS T +D A V
Sbjct: 78 QQIMALRVMTNKLRNAYNGNDIYFQDSSDE-GSGSGSGSGCTDVCPTDADAEDLTTEAPV 136
Query: 529 V-------VDSSVFPTGPP 540
V VD S P+ PP
Sbjct: 137 VEADRGGPVDGSELPSAPP 155
>UniRef100_Q9HG18 PH-regulated protein 2 [Candida dubliniensis]
Length = 556
Score = 35.8 bits (81), Expect = 3.7
Identities = 20/37 (54%), Positives = 23/37 (62%)
Query: 494 GSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVV 530
GS SGSGSGSGSS S+ S+S G KK A +V
Sbjct: 496 GSGSGSGSGSGSGSSSSSSSSSSSSGSSGKKSAASIV 532
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.318 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,056,829,843
Number of Sequences: 2790947
Number of extensions: 46525973
Number of successful extensions: 120260
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 120122
Number of HSP's gapped (non-prelim): 37
length of query: 595
length of database: 848,049,833
effective HSP length: 133
effective length of query: 462
effective length of database: 476,853,882
effective search space: 220306493484
effective search space used: 220306493484
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 78 (34.7 bits)
Lotus: description of TM0069.5


