

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0059.7
(1706 letters)
Database: uniref100
2,790,947 sequences; 848,049,833 total letters
Searching..................................................done
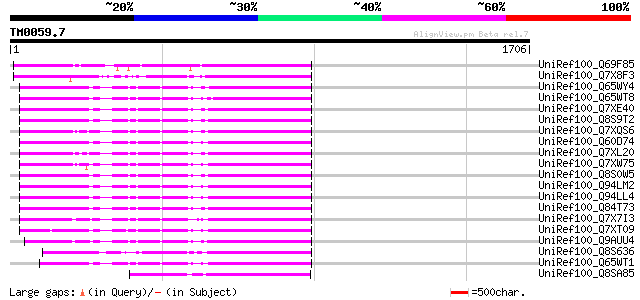
Score E
Sequences producing significant alignments: (bits) Value
UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris] 711 0.0
UniRef100_Q7X8F3 OSJNBa0093P23.5 protein [Oryza sativa] 471 e-130
UniRef100_Q65WY4 Putative polyprotein [Oryza sativa] 460 e-127
UniRef100_Q65WT8 Putative polyprotein [Oryza sativa] 458 e-127
UniRef100_Q7XE40 Putative polyprotein [Oryza sativa] 458 e-127
UniRef100_Q8S9T2 B1064G04.18 protein [Oryza sativa] 458 e-127
UniRef100_Q7XQS6 OSJNBa0043L09.21 protein [Oryza sativa] 457 e-126
UniRef100_Q60D74 Putative polyprotein [Oryza sativa] 456 e-126
UniRef100_Q7XL20 OSJNBa0079M09.11 protein [Oryza sativa] 456 e-126
UniRef100_Q7XW75 OSJNBa0019J05.19 protein [Oryza sativa] 455 e-126
UniRef100_Q8S0W5 Putative polyprotein [Oryza sativa] 455 e-126
UniRef100_Q94LM2 Putative polyprotein [Oryza sativa] 454 e-126
UniRef100_Q94LL4 Putative polyprotein [Oryza sativa] 452 e-125
UniRef100_Q84T73 Putative polyprotein [Oryza sativa] 452 e-125
UniRef100_Q7X7I3 OSJNBb0085F13.8 protein [Oryza sativa] 451 e-125
UniRef100_Q7XT09 OSJNBb0050O03.13 protein [Oryza sativa] 451 e-124
UniRef100_Q9AUU4 Putative polyprotein [Oryza sativa] 441 e-122
UniRef100_Q8S636 Putative polyprotein [Oryza sativa] 433 e-119
UniRef100_Q65WT1 Putative polyprotein [Oryza sativa] 419 e-115
UniRef100_Q8SA85 Prpol [Zea mays] 404 e-110
>UniRef100_Q69F85 Gag-pol polyprotein [Phaseolus vulgaris]
Length = 1859
Score = 711 bits (1835), Expect = 0.0
Identities = 391/1011 (38%), Positives = 587/1011 (57%), Gaps = 61/1011 (6%)
Query: 11 PIGGPLSPEIMAFPFPQGWARPKVKLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPM 70
P G P + +I+A P P W + LY+G +DP EH+N F M ++C+ FP
Sbjct: 207 PKGHPFTDDIIATPLPDKWRGLTINLYDGSTDPDEHLNIFRTQMTLYTTDRTVWCKVFPT 266
Query: 71 SLGKGPMNWFQNLPSNSLHDWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFL 130
SL +GP+ WF +LP NS+ ++ + F QY++ R ++ +L VKQ+ ESL+ F+
Sbjct: 267 SLREGPLGWFSDLPPNSIASFDALELKFTTQYATSRPHRTSSMSLLNVKQERGESLRTFM 326
Query: 131 NRFNKEAGDITGLLPDTRLVLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAV 190
NRF+K +I L P+ + +A+ PG F SL +P ++E RA KF+ +E+ +
Sbjct: 327 NRFSKVCMNIRNLNPEIAMHHLVSAILPGRFTESLIKRPPCNMDELRTRATKFMQIEEHI 386
Query: 191 TLRAASQTLAIKGPQKMKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILRE 250
+ + ++ P+ R + + R P+ + +YTPL ++L E
Sbjct: 387 DYHRKTYAENTDNSKGIRP----PTIPTDRERHRPNRGPR---FHNYTPLIVPRGKVLDE 439
Query: 251 KASTDLRDRPPPLLTRGDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVA 310
+L P T + SKR C++H + GH T+ C LKDK+EEL++AG L ++V
Sbjct: 440 ALQIELIPTLRPSQTPPNADTSKR-CQYHRNYGHTTEGCQALKDKIEELVQAGHLRKFVK 498
Query: 311 VSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPP-------RRRSPERRRRS 363
+ A PRSP RD P + RS R D+T R+RS RR+
Sbjct: 499 TTITA---PRSP----------QRDHDP-RERSGRRDDRTRDNHYRSSRRKRSESPIRRT 544
Query: 364 RERRRSPDRHGRDEVRRQHGSNIV-----------DVGSIAGGWAAGGPTNNSQKRSTRV 412
R + SP+R R + + + N + ++ IAGG+A GG +N+++K+ R
Sbjct: 545 RPKSESPERRSRTKQKVRTVINTIAGPVSLGQPPQEINYIAGGFAGGGCSNSARKKHLRA 604
Query: 413 IMSAAGRPRPGSLHPPRQKVAITFTEDDYGEDTGEEDDPIVIEALIGNGKIRRTLIDTGS 472
I S P H P ITFT++D+ +DDP+VI I I + L+D GS
Sbjct: 605 IQSVHSTPTQRRPHIP----PITFTDEDFTAIDPSQDDPMVITVEIDKFAIAKVLVDQGS 660
Query: 473 SADIMFYDAYKSLGLSVKDLLPYDHDLVGFTGDRVLPLGYFDTCLSLGDHRICRTIKARF 532
S DI++++ +K + + ++ PY+ +VGF+ +RV G+ D + GD + +TI R+
Sbjct: 661 SVDILYWETFKKMKIPEAEIQPYNEQIVGFSRERVDTKGFIDLYTTFGDDYLSKTINIRY 720
Query: 533 LVVECPTAYNAILGRPSLNTFRAIISTHHLMLKYPWA-GRAVPVRGNLEMARSCYNSSCR 591
L+V T+YN +LGRPS+N +AI+ST HL +K+P G V + + AR CY +S +
Sbjct: 721 LLVNANTSYNILLGRPSINRLKAIVSTPHLAMKFPSVNGDIATVHIDQKTARECYVASLK 780
Query: 592 LA-----------REERKRKKSKAHDQNGNYHVQHANLLTDLDPRVDPSRDDHRLKPDGE 640
+ R +R +S G +H L DLDPR+D D R++ +
Sbjct: 781 VEPTRRLYTTSAERTTERRGRSTERRSRGRESRRHLVALVDLDPRLD----DPRMEAGED 836
Query: 641 SHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPGIDPAFCSHRLSV 700
PI + + T + L D + + K L N LFAW++ DMPG+ +HRLSV
Sbjct: 837 LQPIFLRD-KDRKTYMGTSLKPDDRETIGKTLTKNADLFAWTAADMPGVKSDVITHRLSV 895
Query: 701 DRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANGKWRMC 760
+ +P+AQKKR++ E+++ +++T +L++AG I++ YTTWL+NVV+VKK NGKWRMC
Sbjct: 896 YTEARPIAQKKRKLGEERRKAAREETDKLIQAGFIQKAHYTTWLANVVMVKKTNGKWRMC 955
Query: 761 VDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITD 820
VDYTDLNKACPKD +PLP+ID LVD ++G++ LS +DAYSGYNQI M+ D EKTAF TD
Sbjct: 956 VDYTDLNKACPKDSYPLPTIDRLVDGAAGHQILSFLDAYSGYNQIQMYHRDREKTAFRTD 1015
Query: 821 RGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADLAVV 880
+ Y ++PFGLKNA ATYQR+M +F D++G++VEVY+DDI+VK+ H +DL V
Sbjct: 1016 SDNFFYEVMPFGLKNAGATYQRLMDHVFHDMIGRNVEVYVDDIVVKSDSCEQHVSDLKEV 1075
Query: 881 FEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMKSPRTVKEVQQL 940
F+ L+++ MRLNP+KC FGV GKFLG+MLT+RGIE NP+KC+AI M+SP+ +KE+Q+L
Sbjct: 1076 FQALRQYRMRLNPEKCAFGVEGGKFLGFMLTHRGIEANPEKCKAITEMRSPKGLKEIQRL 1135
Query: 941 AGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEILSSP 991
R+ ++ RF+PK A R P+ LLK + FEW+ E + F QLK L+SP
Sbjct: 1136 VSRLTSLSRFVPKLAERTRPIIKLLKKTSKFEWTDECEQNFQQLKAFLASP 1186
>UniRef100_Q7X8F3 OSJNBa0093P23.5 protein [Oryza sativa]
Length = 2074
Score = 471 bits (1211), Expect = e-130
Identities = 313/998 (31%), Positives = 482/998 (47%), Gaps = 120/998 (12%)
Query: 13 GGPLSPEIMAFPFPQGWARPKVKLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSL 72
G PLS I P P + P V Y G++DP+E ++ + A++ A + + ++L
Sbjct: 1089 GLPLSGGIKTRPIPPQFKFPPVPRYSGETDPKEFLSIYESAIEAAHGDENTKAKVIHLAL 1148
Query: 73 GKGPMNWFQNLPSNSLHDWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNR 132
+W+ NLP+NS++ W + F+ + PKT Q L ++Q+ ES++ + R
Sbjct: 1149 DGIARSWYFNLPANSIYSWEQLRDVFVLNFRGTYEEPKTQQHLLGIRQRPGESIRENMRR 1208
Query: 133 FNKEAGDITGLLPDTRLVLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTL 192
F++ + + + + A+A L + K TLE L + F E+
Sbjct: 1209 FSQARCQVQDITEASVINAASAGLLEDELTRKIVNKEPQTLEHPLRIIDGFARGEEDSKR 1268
Query: 193 RAASQT---------------LAIKGPQKMKEHKDQPSTR-ESRRKGQDERKPKRKKYDS 236
R A Q + I P + + QP+ + + R+GQ ++ + D
Sbjct: 1269 RQAIQAEYDKASVAAAQAQAQVQIAEPPPLSVRQSQPAIQGQPPRQGQAPMTWRKFRTDR 1328
Query: 237 YTPLNSSLSRILREKASTDLRDRPPPLLTRGDKLDSKRFCEFHDSPGHNTDECLNLKDKV 296
++ + + D + K+ +C FH H T++C N++ +
Sbjct: 1329 AGKAVMAVEEVQALRKEFDTQQASNHQQPARKKVRKDLYCAFHGRSSHTTEQCRNIRQR- 1387
Query: 297 EELIRAGRLSRYVAVSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRS 356
G + PRP+ P R A + TP ++R
Sbjct: 1388 ------GNVQD---------PRPQQGTTVEAP-------------REAVQEQTTPAKQRQ 1419
Query: 357 PERRRRSRERRRSPDRHGRDEVRRQHGSNIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSA 416
+RRR + +R H +I G G P +Q ++S
Sbjct: 1420 DTQRRRKMQ------------IRMVH--SITSAGE-------GAPQYVNQ------LISF 1452
Query: 417 AGRPRPGSLHPPRQKVAITFTEDDYGEDTGEEDDPIVIEALIGNGKIRRTLIDTGSSADI 476
G L P DP+VI A I ++RR L+D GSSAD+
Sbjct: 1453 GPEDAEGVLFP--------------------HQDPLVISAEIAGFEVRRILVDGGSSADV 1492
Query: 477 MFYDAYKSLGLSVKDLLPYDHDLVGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVE 536
++ +AY +GL + L P L GF+G+ V LG ++ G R + F VV
Sbjct: 1493 IYAEAYAKMGLPTQALTPALTSLRGFSGEAVQVLGQALLLIAFGSGENRREEQILFDVVN 1552
Query: 537 CPTAYNAILGRPSLNTFRAIISTHHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREE 596
P YNAI GR +LN AI ++L LK P + V+G A S C LA
Sbjct: 1553 IPYNYNAIFGRATLNKLEAISHHNYLKLKMPGPTGVIVVKGLQPSAAS----KCDLAIIN 1608
Query: 597 RK--RKKSKAHDQNGNYHVQHANLLTDLDPRVDPSRDDHRLKPDGESHPI*VGPL-PENT 653
R K++ H++ P+ P H G+ + + P
Sbjct: 1609 RAVHNVKTEPHER----------------PKHTPKPTPH-----GKVTKVQINDADPTKL 1647
Query: 654 TNLARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQ 713
+L + + + + ++L N +FAWS +++ G+ H L+V KP QK R+
Sbjct: 1648 VSLGGDMGEEEVESILEVLKKNIDIFAWSPDEVGGVPADLIMHHLAVKPNIKPRKQKLRK 1707
Query: 714 MSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKD 773
MSA++Q+ + + +LLRAG+I+E+ + WL+N VLV+K+NGKWRMCVD+TDLNKACPK
Sbjct: 1708 MSADRQEATKTEVQKLLRAGVIQEIDHPEWLANPVLVRKSNGKWRMCVDFTDLNKACPKH 1767
Query: 774 PFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGL 833
FPLP ID LVD ++G E +S +DAYSGY+QI M+ D KT+FIT GT+C+ +PFGL
Sbjct: 1768 DFPLPRIDQLVDLTAGCELMSFLDAYSGYHQIHMNPLDIPKTSFITPFGTFCHLRMPFGL 1827
Query: 834 KNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNP 893
+NA AT+ R++ ++ G +G++VE Y+DDI+VK+ K DHA DL F+ L ++LNP
Sbjct: 1828 RNAGATFARLVYKVLGKQLGRNVEAYVDDIVVKSRKAFDHAIDLQETFDNLCVAGIKLNP 1887
Query: 894 DKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPK 953
+KC F VR GK LG++++ GIE NP+K AI MK P ++ EVQ+LAGR+AA+ RFL K
Sbjct: 1888 EKCVFSVRVGKLLGFLVSETGIEANPEKIDAIQQMKPPSSIHEVQKLAGRIAALSRFLSK 1947
Query: 954 AALRALPLYTLLKNGATFEWSAEADAAFTQLKEILSSP 991
A R LP + L++ F W+ E AAF +LK+ L SP
Sbjct: 1948 AVERGLPFFKTLRDARKFNWTPECQAAFDELKQHLQSP 1985
>UniRef100_Q65WY4 Putative polyprotein [Oryza sativa]
Length = 2027
Score = 460 bits (1183), Expect = e-127
Identities = 315/974 (32%), Positives = 471/974 (48%), Gaps = 112/974 (11%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIK---GPQK 206
+ L + K T E A++ E+A T ++ +A G
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNSSGSGVAASAAPGSAA 417
Query: 207 MKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTR 266
D+ + S R G + + T S +E A T R+RP
Sbjct: 418 QTGRCDRRRKKRSDRSGDEGHVLAVEGAPRATRKGRPASDKKKE-AGTPSRERP------ 470
Query: 267 GDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRR 326
+ R+C H++ H+ +C +K+ E
Sbjct: 471 -----AGRWCSVHNTSLHDLADCRAVKNLAE----------------------------- 496
Query: 327 TPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GS 384
R + + R ERR+ P S +RR ++++ + D G D++ Q G+
Sbjct: 497 -------RTRKWEEDRRQERREGKSPAVPSGKRRSEAKQKAPAVDIDDGDDDLGFQEPGA 549
Query: 385 NIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGE 443
I V G A + S K R +++ A P ++ R +VA+TF + D+
Sbjct: 550 TIATVD----GGACAQVSRRSFKAMKRELLAMA--PTHEAMRRARWSEVALTFDQTDHPP 603
Query: 444 DTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDL 499
+V+ I N K+ R LID G++ +I+ +DA K+ G+ ++ P +
Sbjct: 604 CIARGGQVAMVVSPTICNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVLQPSQP----I 659
Query: 500 VGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIIST 559
+G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 660 IGVTPGHTWPLGHIDLPVTFGGPANFRTERVSFDVADLSLPYNAVLGRPALVKFMAAVHY 719
Query: 560 HHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLL 619
+L +K P G + VRG+L++A +C R +R SK
Sbjct: 720 AYLQMKMPGPGGPISVRGDLKVALACMEQ-----RADRLAAASK---------------- 758
Query: 620 TDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARG--LPRDLSKRMEKLLLSNGK 677
P D RL P + P +A G +P D + L +N
Sbjct: 759 --------PEGGDERLGPSAPTAP---------RQRMATGDEVPEDA---LVSFLRANAD 798
Query: 678 LFAWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIRE 737
+FAW DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IRE
Sbjct: 799 VFAWRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIRE 858
Query: 738 VKYTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMD 797
V + WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +D
Sbjct: 859 VIHPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLD 918
Query: 798 AYSGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVE 857
AYSGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE
Sbjct: 919 AYSGYHQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVE 978
Query: 858 VYIDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIEL 917
Y+DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE
Sbjct: 979 AYVDDLVVKTRNQETLLSDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEA 1038
Query: 918 NPDKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEA 977
NP+K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA
Sbjct: 1039 NPEKIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEA 1098
Query: 978 DAAFTQLKEILSSP 991
+ A TQLK LSSP
Sbjct: 1099 EHALTQLKAYLSSP 1112
>UniRef100_Q65WT8 Putative polyprotein [Oryza sativa]
Length = 2027
Score = 458 bits (1178), Expect = e-127
Identities = 309/973 (31%), Positives = 466/973 (47%), Gaps = 110/973 (11%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTIAEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMANFFPMTLKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++++ ESL++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRRDDESLRSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKE 209
+ L + K T E A++ E+A T + +A
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVAASAAPGSAA 417
Query: 210 HKDQPSTRESRR--KGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRG 267
+ R +R + DE + S + +++AS R+RP
Sbjct: 418 QAGRRDRRRKKRSVRSDDEGHVLAVEGASRATRKGRPASDKKKEASVPSRERP------- 470
Query: 268 DKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRT 327
+ ++C H++ H+ +C +K E
Sbjct: 471 ----TGKWCTVHNTSLHDLADCRAVKSLAE------------------------------ 496
Query: 328 PTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDR---HGRDEVRRQH-G 383
R + + R ERR+ P P RRRS ++++P G D++ Q G
Sbjct: 497 ------RTRKWEEERRQERREGKSP--AVPSGRRRSEAKQKAPAEDIDDGDDDLGFQEPG 548
Query: 384 SNIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYG 442
+ I V G A + S K R +++AA P + R +VA+TF + D+
Sbjct: 549 ATIATV----DGGACAHVSRRSLKAMKRELLAAA--PTHEATRRARWSEVALTFDQTDHP 602
Query: 443 EDTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIM---FYDAYKSLGLSVKDLLPYDHD 498
+V+ + N K+ R LID G++ +++ +DA K+ G+ ++ P
Sbjct: 603 PCVARGGQIAMVVSPTVCNVKLGRVLIDGGAALNVLSPAAFDAIKAPGMMLRPSQP---- 658
Query: 499 LVGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIIS 558
++G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 659 IIGVTPGHTWPLGHIDLPVTFGGPANFRTERVNFDVADLSLPYNAVLGRPALVKFMAAVH 718
Query: 559 THHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANL 618
+L +K P + V G+L +A +C R++R SK
Sbjct: 719 YAYLQMKMPCPDSPISVHGDLRVALACMEQ-----RKDRLAVVSK--------------- 758
Query: 619 LTDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKL 678
P D RL G S P P + +P D + L +N +
Sbjct: 759 ---------PEGGDERL---GTSAP----AAPRQRIATSDEVPED---ALVSFLRANADV 799
Query: 679 FAWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREV 738
FAW DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV
Sbjct: 800 FAWRPADMPGVPREVIEHRLAVRPSARPVRQKVRRQAPERQAFIREEVARLLEAGFIREV 859
Query: 739 KYTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDA 798
+ WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DA
Sbjct: 860 IHPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDA 919
Query: 799 YSGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEV 858
YSGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE
Sbjct: 920 YSGYHQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEA 979
Query: 859 YIDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELN 918
Y+DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE N
Sbjct: 980 YVDDLVVKTRNQETLLSDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEAN 1039
Query: 919 PDKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEAD 978
P+K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+
Sbjct: 1040 PEKIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAE 1099
Query: 979 AAFTQLKEILSSP 991
A TQLK SSP
Sbjct: 1100 RALTQLKAYFSSP 1112
>UniRef100_Q7XE40 Putative polyprotein [Oryza sativa]
Length = 2026
Score = 458 bits (1178), Expect = e-127
Identities = 311/972 (31%), Positives = 466/972 (46%), Gaps = 108/972 (11%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 237 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 296
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 297 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 356
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIK---GPQK 206
+ L + K T E A++ E+A T ++ +A G
Sbjct: 357 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNSSGSGVAASAAPGSAA 416
Query: 207 MKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTR 266
+D+ + S R G + + T S +E A T R+RP
Sbjct: 417 QTGRRDRRRKKRSDRSGDEGHVLAVEGAPRATRKGRPASDKKKE-AGTPSRERP------ 469
Query: 267 GDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRR 326
+ R+C H++ H+ +C +K+ E
Sbjct: 470 -----AGRWCSVHNTSLHDLADCRAVKNLAE----------------------------- 495
Query: 327 TPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GS 384
R + + R ERR+ P S +RR ++++ + D G D++ Q G+
Sbjct: 496 -------RTRKWEEDRRQERREGKSPADPSGKRRSEAKQKAPAVDIDDGDDDLGFQEPGA 548
Query: 385 NIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGE 443
I V G A + S K R ++ A P ++ R +VA+TF + D+
Sbjct: 549 TIATVD----GGACAHVSRRSFKAMKREFLAMA--PTHEAMRRARWSEVALTFDQTDHPP 602
Query: 444 DTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDL 499
+V+ I N K+ R L+D G++ +I+ +DA K+ G+ ++ P +
Sbjct: 603 CIARGGQVAMVVSPTICNVKLGRVLVDGGAALNILSPAAFDAIKAPGMVLRPSQP----I 658
Query: 500 VGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIIST 559
+G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 659 IGVTPGHTWPLGHIDLPVTFGGPANFRTERVSFDVADLSLPYNAVLGRPALVKFMAAVHY 718
Query: 560 HHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLL 619
+L +K P G + V G+L++A +C R +R SK
Sbjct: 719 AYLQMKMPGPGGPISVHGDLKVALACMEQ-----RADRLAAASK---------------- 757
Query: 620 TDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLF 679
P D RL P + P +P D + L +N +F
Sbjct: 758 --------PEGGDERLGPSAPT-------APRQRIVTCDEVPEDA---LVSFLRANADVF 799
Query: 680 AWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVK 739
AW DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV
Sbjct: 800 AWRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVI 859
Query: 740 YTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAY 799
+ WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAY
Sbjct: 860 HPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAY 919
Query: 800 SGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVY 859
SGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y
Sbjct: 920 SGYHQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEAY 979
Query: 860 IDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNP 919
+DD++VKT +DLA FE L+ M+LNPDKC FGV +GK LG++++ RGIE NP
Sbjct: 980 VDDLVVKTRNQETLLSDLAETFENLRSARMKLNPDKCVFGVPAGKLLGFLVSARGIEANP 1039
Query: 920 DKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADA 979
+K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+
Sbjct: 1040 EKIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAEN 1099
Query: 980 AFTQLKEILSSP 991
A QLK LSSP
Sbjct: 1100 ALAQLKAYLSSP 1111
>UniRef100_Q8S9T2 B1064G04.18 protein [Oryza sativa]
Length = 2030
Score = 458 bits (1178), Expect = e-127
Identities = 310/972 (31%), Positives = 467/972 (47%), Gaps = 105/972 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIK---GPQK 206
+ L + K T E A++ E+A T ++ +A G
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNSSGSGVAASAAPGSAA 417
Query: 207 MKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTR 266
+D+ + S R G + + T S +E A T ++RP
Sbjct: 418 QTGRRDRRRKKRSDRSGDEGHVLAVEGAPRATRKGRPASDKKKE-AGTPSKERP------ 470
Query: 267 GDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRR 326
+ R+C H++ H+ +C +K+ E
Sbjct: 471 -----AGRWCSVHNTSLHDLVDCRAVKNLAE----------------------------- 496
Query: 327 TPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GS 384
R + + R ERR+ P S +RR ++++ + D G D++ Q G+
Sbjct: 497 -------RTRKWEEDRRQERREGKSPAVPSGKRRSEAKQKAPAVDIDDGDDDLGFQEPGA 549
Query: 385 NIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGE 443
I V G A + S K R +++ A P ++ R +VA+TF + D+
Sbjct: 550 TIATVD----GGACAHVSRRSFKAMKRELLAMA--PTHEAMRRARWSEVALTFDQTDHPP 603
Query: 444 DTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDL 499
+V+ I N K+ R LID G++ +I+ +DA K+ G+ ++ P +
Sbjct: 604 CIARGGQVAMVVSPTICNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVLQPSQP----I 659
Query: 500 VGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIIST 559
+G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 660 IGVTPGHTWPLGHIDLPVTFGGPANFRTERVNFDVADLSLPYNAVLGRPALVKFMAAVHY 719
Query: 560 HHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLL 619
+L +K P G + V G+L++A +C R +R SK
Sbjct: 720 AYLQMKMPGPGGPISVHGDLKVALACMEQ-----RADRLAAASK---------------- 758
Query: 620 TDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLF 679
P D RL P + P +P + L +N +F
Sbjct: 759 --------PEGGDERLGPSAPT-------APRQRMATCDEVPVKEEDALVSFLRANADVF 803
Query: 680 AWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVK 739
AW DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV
Sbjct: 804 AWRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVI 863
Query: 740 YTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAY 799
+ WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAY
Sbjct: 864 HPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAY 923
Query: 800 SGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVY 859
SGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y
Sbjct: 924 SGYHQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEAY 983
Query: 860 IDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNP 919
+DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP
Sbjct: 984 VDDLVVKTRNQETLLSDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANP 1043
Query: 920 DKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADA 979
+K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+
Sbjct: 1044 EKIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAEH 1103
Query: 980 AFTQLKEILSSP 991
A TQLK LSSP
Sbjct: 1104 ALTQLKAYLSSP 1115
>UniRef100_Q7XQS6 OSJNBa0043L09.21 protein [Oryza sativa]
Length = 2028
Score = 457 bits (1177), Expect = e-126
Identities = 309/969 (31%), Positives = 468/969 (47%), Gaps = 102/969 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKE 209
+ L + K + T E A++ E+A T ++ +A
Sbjct: 358 YAFRGGVRHNRMLEKIASKESQTTAELFQLADRVARKEEAWTWNSSGSGVAASAAPGSAA 417
Query: 210 HKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRGDK 269
Q R+ RRK ++P R + + R R+ D+ T +
Sbjct: 418 ---QTGRRDRRRK----KRPDRSGDEGHVLAVEGAPRATRKGRPAS--DKKNEAGTPSRE 468
Query: 270 LDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRTPT 329
+ R+C H++ H+ +C +K+ E
Sbjct: 469 RLAGRWCSVHNTSLHDLADCRAVKNLAE-------------------------------- 496
Query: 330 PPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GSNIV 387
R + + R ERR+ P S +RR ++++ + D G D++ Q G+ I
Sbjct: 497 ----RTRKWEEDRRQERREGKSPAVPSGKRRSEAKQKAPAVDIDDGDDDLGFQEPGATIA 552
Query: 388 DVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGEDTG 446
V G A + S K R +++ A P ++ R +VA+TF + D+
Sbjct: 553 TVD----GGACAHVSRRSFKAMKRELLAMA--PTHEAMRRARWSEVALTFDQTDHPPCIA 606
Query: 447 EEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLVGF 502
+ + I N K+ R LID G++ +I+ +DA K+ G+ ++ P ++G
Sbjct: 607 RGGQVAMAVSPTICNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVLQPSQP----IIGV 662
Query: 503 TGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTHHL 562
T PLG+ D ++ G RT + F V + YNA+LGRP+L F A + +L
Sbjct: 663 TPGHTWPLGHIDLPVTFGGPANFRTERVSFDVADLSLPYNAVLGRPALVKFMAAVHYAYL 722
Query: 563 MLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLTDL 622
+K P G + VRG+L++A +C R +R SK
Sbjct: 723 QMKMPGPGGPISVRGDLKVALACMEQ-----RADRLAAASK------------------- 758
Query: 623 DPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWS 682
P D RL P + P +P D + L +N +FAW
Sbjct: 759 -----PEGGDERLGPSAPT-------APRQRMATCDEVPEDA---LVSFLRANADVFAWR 803
Query: 683 SEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTT 742
DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV +
Sbjct: 804 PADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVIHPE 863
Query: 743 WLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGY 802
WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAYSGY
Sbjct: 864 WLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAYSGY 923
Query: 803 NQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDD 862
+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y+DD
Sbjct: 924 HQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEAYVDD 983
Query: 863 IIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKC 922
++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+K
Sbjct: 984 LVVKTRNQETLLSDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPEKI 1043
Query: 923 QAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFT 982
+AI ++ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+ A T
Sbjct: 1044 RAIERIRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAEHALT 1103
Query: 983 QLKEILSSP 991
QLK LSSP
Sbjct: 1104 QLKAYLSSP 1112
>UniRef100_Q60D74 Putative polyprotein [Oryza sativa]
Length = 1869
Score = 456 bits (1174), Expect = e-126
Identities = 307/971 (31%), Positives = 462/971 (46%), Gaps = 103/971 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 111 RPTIAEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 170
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++++ ESL++++ RF + I + +
Sbjct: 171 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRRDDESLRSYIQRFCQVRNTIPCIPAHAVI 230
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKE 209
+ L + K T E A++ E+A T + +A
Sbjct: 231 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVAASAAPGSTA 290
Query: 210 HKDQPSTRESRRK--GQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRG 267
+ R +R DE + S + ++KA R+RP
Sbjct: 291 QTGRRDRRRKKRSVHSDDEGHVLAVEGASRATQKGRPASDKKKKAGAPSRERP------- 343
Query: 268 DKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRT 327
+ ++C H++ H+ +C +K E
Sbjct: 344 ----ASKWCTVHNTSLHDLADCRAVKSLAE------------------------------ 369
Query: 328 PTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GSN 385
R + + R ERR+ P S RR ++++ + D G D++ Q G+
Sbjct: 370 ------RTRKWEEERRQERREGKSPAVPSGNRRSEAKQKAPAEDIDDGDDDLGFQEPGAT 423
Query: 386 IVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGED 444
I V G A + S K R +++AA P + R +VA+TF + D+
Sbjct: 424 IATVD----GGACAHVSRRSLKAMKRELLAAA--PTHEATRRARWSEVALTFDQTDHPPC 477
Query: 445 TGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLV 500
+V+ + N K+ R LID G++ +I+ +DA K+LG+ ++ P ++
Sbjct: 478 VARGGQIAMVVSPTVCNVKLGRVLIDGGAALNILSPAAFDAIKALGMMLRPSQP----II 533
Query: 501 GFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTH 560
G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 534 GVTPGHTWPLGHIDLPVTFGGPANFRTERVNFDVADLSPPYNAVLGRPALVKFMAAVHYA 593
Query: 561 HLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLT 620
+L +K P + V G+L++A +C R +R SK
Sbjct: 594 YLQMKMPGPDGPISVHGDLKVALACMEQ-----RADRLAVVSK----------------- 631
Query: 621 DLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFA 680
P D RL G S P P +P + L +N +FA
Sbjct: 632 -------PEGGDERL---GTSAPA----TPRQRIVTCDEVPVKEEDALVSFLRANADVFA 677
Query: 681 WSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKY 740
W DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV +
Sbjct: 678 WRPADMPGVPREVIEHRLAVRPDARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVIH 737
Query: 741 TTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYS 800
WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAYS
Sbjct: 738 PEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAYS 797
Query: 801 GYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYI 860
GY+QI M R+DEEKT FIT GTYCYT +PFGL+NA T QR+ G +G++VE Y+
Sbjct: 798 GYHQIRMAREDEEKTTFITPIGTYCYTTMPFGLRNAGPTIQRITRISLGSQLGRNVEAYV 857
Query: 861 DDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPD 920
DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+
Sbjct: 858 DDLVVKTRNQETLLSDLAETFESLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPE 917
Query: 921 KCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAA 980
K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+ A
Sbjct: 918 KIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAERA 977
Query: 981 FTQLKEILSSP 991
TQLK LSSP
Sbjct: 978 LTQLKAYLSSP 988
>UniRef100_Q7XL20 OSJNBa0079M09.11 protein [Oryza sativa]
Length = 2028
Score = 456 bits (1173), Expect = e-126
Identities = 310/969 (31%), Positives = 465/969 (46%), Gaps = 102/969 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 239 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 298
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 299 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 358
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKE 209
+ L + K T E A++ E+A T ++ A
Sbjct: 359 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNSSGSGAAASAAPGSAA 418
Query: 210 HKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRGDK 269
Q R+ RRK + +R + + R R+ D+ T G +
Sbjct: 419 ---QTGRRDRRRKKRSDRSGD----EGHVLAVEGAPRATRKGRPAS--DKKKEAGTPGRE 469
Query: 270 LDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRTPT 329
+ R+C H++ H+ +C +K+ E
Sbjct: 470 RPAGRWCSVHNTSLHDLADCRAVKNLAE-------------------------------- 497
Query: 330 PPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GSNIV 387
R + + R ERR+ P S +RR ++++ D G D++ Q G+ I
Sbjct: 498 ----RTRKWEEDRRQERREGKSPVVPSGKRRGEAKQKAPVVDIDDGDDDLGFQEPGATIA 553
Query: 388 DVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGEDTG 446
V G A + S K R +++ A P ++ R +VA+TF + D+
Sbjct: 554 TVD----GGACAHVSRRSFKAMKRELLAMA--PTHEAMRRARWSEVALTFDQTDHPPCIA 607
Query: 447 EEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLVGF 502
+V+ I N K+ R LID G++ +I+ +DA K+ G+ ++ P ++G
Sbjct: 608 RGGQVAMVVSPTICNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVLQPSQP----IIGV 663
Query: 503 TGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTHHL 562
T PLG+ D ++ G RT + F V + YNA+LGRP+L F A + +L
Sbjct: 664 TPGHTWPLGHIDLPVTFGGPANFRTERVSFDVADLSLPYNAVLGRPALVKFMAAVHYAYL 723
Query: 563 MLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLTDL 622
+K P G + VRG+L++A +C R +R SK
Sbjct: 724 QMKMPGPGGPISVRGDLKVALACIEQ-----RADRLAAASK------------------- 759
Query: 623 DPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWS 682
P D RL P + P +P D + L +N +FAW
Sbjct: 760 -----PEGGDERLGPSAPT-------APRQRMATCDEVPEDA---LVSFLRANADVFAWR 804
Query: 683 SEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTT 742
DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV +
Sbjct: 805 PADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVIHPE 864
Query: 743 WLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGY 802
WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAYSGY
Sbjct: 865 WLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAYSGY 924
Query: 803 NQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDD 862
+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y+DD
Sbjct: 925 HQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEAYVDD 984
Query: 863 IIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKC 922
++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+K
Sbjct: 985 LVVKTRNQETLLSDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPEKI 1044
Query: 923 QAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFT 982
+AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+ A
Sbjct: 1045 RAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAENALA 1104
Query: 983 QLKEILSSP 991
QLK LSSP
Sbjct: 1105 QLKAYLSSP 1113
>UniRef100_Q7XW75 OSJNBa0019J05.19 protein [Oryza sativa]
Length = 1969
Score = 455 bits (1171), Expect = e-126
Identities = 316/980 (32%), Positives = 468/980 (47%), Gaps = 124/980 (12%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTIAEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++++ ESL++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRRDDESLRSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKE 209
+ L + K T E A++ E+A T + +A
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVAASAAPGSAA 417
Query: 210 HKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILRE---------KASTDLRDRP 260
Q R+ RRK ++ + + SR R+ +AS R+RP
Sbjct: 418 ---QAGRRDRRRK----KRSVHSDNEGHVLAVEGASRATRKGRPASDKKKEASVPSRERP 470
Query: 261 PPLLTRGDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPR 320
+ ++C H++ H+ +C +K E
Sbjct: 471 -----------TGKWCTVHNTSLHDLADCRAVKSLAE----------------------- 496
Query: 321 SPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRH---GRDE 377
R + + R ERR+ P P RRRS R+++P G D+
Sbjct: 497 -------------RTRKWEEERRQERREGKSPA--VPSGRRRSEARQKAPAEDIDDGDDD 541
Query: 378 VRRQH-GSNIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAIT 435
+ Q G+ I V G A + S K R +++AA P + R +VA+T
Sbjct: 542 LGFQEPGATIATVD----GGACAHVSRRSLKAMKRELLAAA--PTHEATRRARWSEVALT 595
Query: 436 FTEDDYGEDTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKD 491
F + D+ +V+ I N K+ R LID G++ +I+ +DA K+ G+ ++
Sbjct: 596 FDQTDHPPCVAWGGQIAMVVSPTICNVKLGRVLIDGGAALNILSPAAFDAIKAPGMMLRP 655
Query: 492 LLPYDHDLVGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLN 551
P ++G T PLG+ D ++ G RT + F V + YNA+LGRP+L
Sbjct: 656 SQP----IIGVTPGHTWPLGHIDLPVTFGGPANFRTERVNFDVADLSLPYNAVLGRPALV 711
Query: 552 TFRAIISTHHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNY 611
F A + +L +K P + V G+L++A +C R +R SK
Sbjct: 712 KFMAAVHYAYLQMKMPGPDGPISVHGDLKVALACMEQ-----RADRLAVVSK-------- 758
Query: 612 HVQHANLLTDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKL 671
P D RL G S P P + +P D +
Sbjct: 759 ----------------PEGGDERL---GTSAPA----APRQRIVTSDEVPEDA---LVSF 792
Query: 672 LLSNGKLFAWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLR 731
L +N +FAW DMPG+ HRL+V +PV QK R+ + E+Q I+ + LL
Sbjct: 793 LRANADVFAWRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIRVEVARLLE 852
Query: 732 AGIIREVKYTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYE 791
AG IREV + WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G +
Sbjct: 853 AGFIREVIHPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCD 912
Query: 792 YLSLMDAYSGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDL 851
L +DAYSGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G
Sbjct: 913 LLCFLDAYSGYHQIRMAREDEEKTAFITPIGTYCYTTMPFGLKNAGPTFQRTTRISLGSQ 972
Query: 852 MGKSVEVYIDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLT 911
+G++VE Y+DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++
Sbjct: 973 LGRNVEAYVDDLVVKTRNQETLLSDLAETFESLRSARIKLNPDKCVFGVPAGKLLGFLVS 1032
Query: 912 NRGIELNPDKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATF 971
RGIE NP+K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F
Sbjct: 1033 ARGIEANPEKIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPF 1092
Query: 972 EWSAEADAAFTQLKEILSSP 991
W+ EA+ A TQLK LSSP
Sbjct: 1093 TWTEEAERALTQLKAYLSSP 1112
>UniRef100_Q8S0W5 Putative polyprotein [Oryza sativa]
Length = 2030
Score = 455 bits (1171), Expect = e-126
Identities = 307/972 (31%), Positives = 463/972 (47%), Gaps = 105/972 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIK---GPQK 206
+ L + K T E A++ E+A T + +A G
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVAASAAPGSAA 417
Query: 207 MKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTR 266
+D+ + S G + + T S +E A T R+RP
Sbjct: 418 QTGRRDRRRKKRSAHSGDEGHVLAVESAPRATRKGRPASDKKKE-AGTPSRERPV----- 471
Query: 267 GDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRR 326
++C H++ H+ +C +K+ E
Sbjct: 472 ------SKWCSVHNTSLHDLADCRAVKNLAE----------------------------- 496
Query: 327 TPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVR--RQHGS 384
R + + R ERR+ P S +RR ++++ + D D+ ++ G+
Sbjct: 497 -------RTRKWEEDRRQERREGKSPAVPSGKRRSEAKQKAPAVDIDDGDDGLGFQEPGA 549
Query: 385 NIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGE 443
I V G A + S K R +++ A P ++ R +VA+TF + D+
Sbjct: 550 TIATVD----GGACAHVSRRSFKAMKRELLAMA--PTHEAMRRARWSEVALTFDQTDHPP 603
Query: 444 DTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDL 499
+V+ I N K+ R LID G++ +I+ +DA K+ G+ ++ P +
Sbjct: 604 CIARGGQVAMVVSPTICNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVLRPSQP----I 659
Query: 500 VGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIIST 559
+G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 660 IGVTPGHTWPLGHIDLPVTFGGPANFRTERVSFDVADLSLPYNAVLGRPALVKFMAAVHY 719
Query: 560 HHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLL 619
+L +K P G + V G+L++A +C R +R SK
Sbjct: 720 AYLQMKMPGPGGPISVHGDLKVALACMEQ-----RADRLAAASK---------------- 758
Query: 620 TDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLF 679
P D RL P + P +P + L +N +F
Sbjct: 759 --------PEGGDERLGPSAPT-------APRQRIATCDEVPVKEEDALVSFLRANADVF 803
Query: 680 AWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVK 739
AW DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV
Sbjct: 804 AWRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVI 863
Query: 740 YTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAY 799
+ WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAY
Sbjct: 864 HPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAY 923
Query: 800 SGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVY 859
SGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y
Sbjct: 924 SGYHQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEAY 983
Query: 860 IDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNP 919
+DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP
Sbjct: 984 VDDLVVKTRNQETLLSDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANP 1043
Query: 920 DKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADA 979
+K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+
Sbjct: 1044 EKIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAEH 1103
Query: 980 AFTQLKEILSSP 991
A TQLK LSSP
Sbjct: 1104 ALTQLKVYLSSP 1115
>UniRef100_Q94LM2 Putative polyprotein [Oryza sativa]
Length = 2027
Score = 454 bits (1169), Expect = e-126
Identities = 308/972 (31%), Positives = 463/972 (46%), Gaps = 108/972 (11%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIK---GPQK 206
+ L + K T E A++ E+A T + +A G
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVAASAAPGSAA 417
Query: 207 MKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTR 266
+D+ + S G + + T S +E A R+RP
Sbjct: 418 QTGRRDRRRKKRSVHSGDEGHVLAVEGAPRATRKGRPASDKKKE-AGAPSRERP------ 470
Query: 267 GDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRR 326
+ ++C H++ H+ +C +K+ E
Sbjct: 471 -----AGKWCSVHNTSLHDLADCRAVKNLAE----------------------------- 496
Query: 327 TPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVR--RQHGS 384
R + + R ERR+ P S RRR ++++ + D G D+ ++ G+
Sbjct: 497 -------RTKKWEEDRRQERRENKSPAVLSGNRRREAKQKAPAEDIDGDDDDLGFQEPGA 549
Query: 385 NIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGE 443
I V G A + S K R +++A P + R +VA+TF + D+
Sbjct: 550 TIATVD----GGACAHVSRRSFKAMKRELLAAV--PTHEATRRARWSEVALTFDQTDHPP 603
Query: 444 DTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDL 499
+V+ I N K+ R LID G++ +I+ +DA K+ G+ ++ P +
Sbjct: 604 CVARGGQIAMVVSPTICNVKLGRVLIDGGTALNILSPAAFDAIKAPGMVLRPSQP----I 659
Query: 500 VGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIIST 559
+G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 660 IGVTPGHTWPLGHIDLPVTFGGSANFRTERVNFDVADLSLPYNAVLGRPALVKFMAAVHY 719
Query: 560 HHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLL 619
+L +K P G + V G+L++A +C R + SK
Sbjct: 720 AYLQMKMPGPGGPISVHGDLKVALACMEQ-----RADHLAAASK---------------- 758
Query: 620 TDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLF 679
P D RL G S P P + +P D + L +N +F
Sbjct: 759 --------PEGGDERL---GTSAPT----APRQRIDTCDDVPEDA---LVSFLRANADVF 800
Query: 680 AWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVK 739
AW DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV
Sbjct: 801 AWRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVI 860
Query: 740 YTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAY 799
+ WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAY
Sbjct: 861 HPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAY 920
Query: 800 SGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVY 859
SGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+Q G +G++VE Y
Sbjct: 921 SGYHQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQHTTRISLGSQIGRNVEAY 980
Query: 860 IDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNP 919
+DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP
Sbjct: 981 VDDLVVKTRNQETFLSDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANP 1040
Query: 920 DKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADA 979
+K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+
Sbjct: 1041 EKIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAER 1100
Query: 980 AFTQLKEILSSP 991
A QLK LSSP
Sbjct: 1101 ALAQLKAYLSSP 1112
>UniRef100_Q94LL4 Putative polyprotein [Oryza sativa]
Length = 1992
Score = 452 bits (1164), Expect = e-125
Identities = 308/971 (31%), Positives = 463/971 (46%), Gaps = 106/971 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTIAEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++++ E+L++++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRRDDETLRSYIQRFCQARNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKE 209
+ L + K T E A++ E+A T + +A
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVAASAASGSAA 417
Query: 210 HKDQPSTRESRRK--GQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRG 267
+ R +R DE + S + ++KA R+RP
Sbjct: 418 QTGRRDRRRKKRSVHSDDEGHVLAVEGASRATQKGRPASDKKKKAGAPSRERP------- 470
Query: 268 DKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRT 327
+ ++C H++ H+ +C +K E
Sbjct: 471 ----AGKWCTVHNTSLHDLMDCRAVKSLAE------------------------------ 496
Query: 328 PTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GSN 385
R + + R ERR+ P S RR ++++ + D G D++ Q G+
Sbjct: 497 ------RTRKWEEERRQERREGKSPAVPSGNRRSEAKQKAPAEDIDDGDDDLGFQEPGAT 550
Query: 386 IVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGED 444
I V G A + S K R +++AA P + R +VA+TF + D+
Sbjct: 551 IATVD----GGACAHVSRRSLKAMKRELLAAA--PTHEATRRARWSEVALTFDQTDHPPC 604
Query: 445 TGEEDDPI-VIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLV 500
+ + V+ + N K+ LID G++ +I+ +DA K+ G+ ++ P ++
Sbjct: 605 VAQGGQIVMVVSPTVCNVKLGHVLIDGGAALNILSPAAFDAIKAPGMMLRPSQP----II 660
Query: 501 GFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTH 560
G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 661 GVTPGHTWPLGHIDLPVTFGGPVNFRTERVNFDVADLSLPYNAVLGRPALVKFMAAVHYA 720
Query: 561 HLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLT 620
+L +K P + V G+L++A +C R +R SK
Sbjct: 721 YLQMKMPGPDGPISVHGDLKVALACMEQ-----RADRLAVVSK----------------- 758
Query: 621 DLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFA 680
P D RL G S P P +P D + L +N +FA
Sbjct: 759 -------PEGGDERL---GTSAPA----APRQRIVTCDEVPEDA---LVSFLRANADVFA 801
Query: 681 WSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKY 740
W DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV +
Sbjct: 802 WRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQALERQAFIREEVARLLEAGFIREVIH 861
Query: 741 TTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYS 800
WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAYS
Sbjct: 862 PEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAYS 921
Query: 801 GYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYI 860
GY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G +VE Y+
Sbjct: 922 GYHQIRMAREDEEKTAFITPIGTYCYTTMPFGLKNAGPTFQRTTRISLGSQIGCNVEAYV 981
Query: 861 DDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPD 920
DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+
Sbjct: 982 DDLVVKTRNQETLLSDLAETFESLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPE 1041
Query: 921 KCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAA 980
K +AI M+ P +++VQ +AG MAA+ RF+ + +ALPL+ LLK F W+ EA+ A
Sbjct: 1042 KIRAIERMRPPSKLRDVQCVAGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAERA 1101
Query: 981 FTQLKEILSSP 991
TQLK LSSP
Sbjct: 1102 LTQLKAYLSSP 1112
>UniRef100_Q84T73 Putative polyprotein [Oryza sativa]
Length = 2026
Score = 452 bits (1163), Expect = e-125
Identities = 310/972 (31%), Positives = 466/972 (47%), Gaps = 108/972 (11%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NL S+H
Sbjct: 237 RPTITEKYDGSVNPTEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLSPASVH 296
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++ + ESL++++ RF + I + +
Sbjct: 297 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVI 356
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIK---GPQK 206
+ L + K T E A++ E+A T ++ +A G
Sbjct: 357 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNSSGSGVAASAAPGSAA 416
Query: 207 MKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTR 266
+D+ + S R G + + T S +E A T R+RP
Sbjct: 417 QTGRRDRRRKKRSDRSGDEGHVLAVEGAPRATRKGRPASDKKKE-AGTPSRERP------ 469
Query: 267 GDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRR 326
+ R+C H++ H+ +C +K+ E
Sbjct: 470 -----AGRWCSVHNTSLHDLADCRAVKNLAE----------------------------- 495
Query: 327 TPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GS 384
R + + R ERR+ P S +RR ++++ + D G D++ Q G+
Sbjct: 496 -------RTRKWEEDRRQERREGKFPAVPSGKRRSEAKQKAPAVDIDDGDDDLGFQEPGA 548
Query: 385 NIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGE 443
I V G A + S K R +++ A P ++ R +VA+TF + D+
Sbjct: 549 TIATVD----GGACAHVSRRSFKAMKRELLAMA--PTHEAMRRARWSEVALTFDQTDHPP 602
Query: 444 DTGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDL 499
+V+ I N K+ R LID G++ +I+ +DA K+ G+ ++ P +
Sbjct: 603 CIARGGQVAMVVSPTICNVKLGRVLIDGGAALNILSPVAFDAIKAPGMVLRPSQP----I 658
Query: 500 VGFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIIST 559
+G T PLG+ D ++ G RT + F V + YNA+LGRP+L F A +
Sbjct: 659 IGVTPGHTWPLGHIDLPVTFGGPANFRTERVSFDVADLSLPYNAVLGRPALVKFMAAVHY 718
Query: 560 HHLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLL 619
+L +K P G + V G+L++A +C R +R SK
Sbjct: 719 AYLQMKMPGPGGPISVHGDLKVALACMEQ-----RTDRLAAASK---------------- 757
Query: 620 TDLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLF 679
P D RL P + P +P D + L +N +F
Sbjct: 758 --------PEGGDERLGPSAPT-------APRQRIATYDEVPEDA---LVSFLRANADVF 799
Query: 680 AWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVK 739
AW DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV
Sbjct: 800 AWRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVI 859
Query: 740 YTTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAY 799
+ WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAY
Sbjct: 860 HPEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAY 919
Query: 800 SGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVY 859
SGY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y
Sbjct: 920 SGYHQIRMAREDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEAY 979
Query: 860 IDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNP 919
+DD++VKT DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP
Sbjct: 980 VDDLVVKTRNQETLLLDLAETFENLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANP 1039
Query: 920 DKCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADA 979
+K +AI ++ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+
Sbjct: 1040 EKIRAIERIRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAEH 1099
Query: 980 AFTQLKEILSSP 991
A TQLK LSSP
Sbjct: 1100 ALTQLKAYLSSP 1111
>UniRef100_Q7X7I3 OSJNBb0085F13.8 protein [Oryza sativa]
Length = 1932
Score = 451 bits (1160), Expect = e-125
Identities = 309/969 (31%), Positives = 464/969 (46%), Gaps = 102/969 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTIAEKYDGSVNPAEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++++ ESL +++ RF + I + +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRRDDESLHSYIQRFCQVRNTIPCIPAHAVI 357
Query: 150 VLATAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKE 209
+ L + K T E A++ E+A T + +A
Sbjct: 358 YAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEACTWNPSGSGVAASAAPGS-- 415
Query: 210 HKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRGDK 269
+ R RR + +++ + + SR R+ + + +R ++
Sbjct: 416 -----AARTGRRDRRRKKRSAHSDDEGHVLAVEGASRATRKGRPASDKKKEAGAPSR-ER 469
Query: 270 LDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRTPT 329
L K +C H++ H+ +C +K E
Sbjct: 470 LTGK-WCIVHNTSLHDLADCRAVKSLAE-------------------------------- 496
Query: 330 PPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GSNIV 387
R + + R ERR+ P S RR ++++ + D G D++ Q G+ I
Sbjct: 497 ----RTRKWEEERRQERREGKSPAVPSGNRRSEAKQKAPAEDIDDGDDDLGFQEPGATIA 552
Query: 388 DVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGEDTG 446
V G A + S K R +++AA P + R +VA+TF + D+
Sbjct: 553 TVD----GGACAHVSRWSLKAMKRELLAAA--PTHEATRRARWSEVALTFDQTDHPPCVA 606
Query: 447 EEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLVGF 502
+V+ + N K+ R LID G++ +I+ +DA K+ G+ ++ P ++G
Sbjct: 607 RGGQIAMVVSPTVCNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVLRPSQP----IIGV 662
Query: 503 TGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTHHL 562
T PLG+ D ++ G RT + F V + YNA+LGRP+L F A + +L
Sbjct: 663 TPGHTWPLGHIDLPVTFGGSANFRTERVNFDVADLSLPYNAVLGRPALVKFMAAVHYAYL 722
Query: 563 MLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLTDL 622
+K P G + V G+L++A +C Q AN LT
Sbjct: 723 QMKMPGPGGPISVHGDLKVALACME--------------------------QRANHLTAA 756
Query: 623 DPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWS 682
P D RL G S P P +P D + L +N +FAW
Sbjct: 757 SK---PKGGDERL---GTSAPA----APRQRIVTCDEVPEDA---LVSFLRANAHVFAWR 803
Query: 683 SEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTT 742
DMPG+ HRL+V + V QK R+ + E+Q I+++ LL AG IREV +
Sbjct: 804 PADMPGVPREVIEHRLAVRSGARLVRQKVRRQAPERQAFIREEVARLLEAGFIREVIHPE 863
Query: 743 WLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGY 802
WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID VD+++G + L +DAYSGY
Sbjct: 864 WLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQFVDSTAGCDLLCFLDAYSGY 923
Query: 803 NQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDD 862
+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y+DD
Sbjct: 924 HQIRMAREDEEKTAFITPIGTYCYTTMPFGLKNAGPTFQRTTRISLGSQIGRNVEAYVDD 983
Query: 863 IIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKC 922
++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+K
Sbjct: 984 LVVKTRNQETLLSDLAETFESLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPEKI 1043
Query: 923 QAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFT 982
+AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+ A T
Sbjct: 1044 RAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAERALT 1103
Query: 983 QLKEILSSP 991
QLK LSSP
Sbjct: 1104 QLKAYLSSP 1112
>UniRef100_Q7XT09 OSJNBb0050O03.13 protein [Oryza sativa]
Length = 2030
Score = 451 bits (1159), Expect = e-124
Identities = 307/971 (31%), Positives = 465/971 (47%), Gaps = 103/971 (10%)
Query: 31 RPKV-KLYEGDSDPQEHVNFFVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLH 89
RP + + Y+G +P E + + ++ AG D + FPM+L W NLP S+H
Sbjct: 238 RPTIAEKYDGSINPAEFLQVYTTGIEAAGGDDRVMANFFPMALKGQARGWLMNLPPASVH 297
Query: 90 DWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRL 149
W + F + P L V++++ ESL++++ RF + + +P +
Sbjct: 298 SWEDLCQQFTMNFQGTYPRPGEEADLHAVQRRDDESLRSYIQRFC-QVRNTKPCIPAHAV 356
Query: 150 VLA-TAALAPGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMK 208
+ A + L + K T E A++ E+A T + +A
Sbjct: 357 IYAFRGGVRHNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSGSGVAASAAPGST 416
Query: 209 EHKDQPSTRESRRK--GQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTR 266
+ R +R DE + S + ++KA R+RP
Sbjct: 417 AQTGRRDRRRKKRSVHSDDEGHVLAVEGASRATQKGRPASDKKKKAGAPSRERP------ 470
Query: 267 GDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRR 326
+ ++C H++ H+ +C +K E
Sbjct: 471 -----AGKWCAVHNTSLHDLADCRAVKSLAE----------------------------- 496
Query: 327 TPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQHGSN 385
R + + R ERR+ P S RR ++++ + D G D++ Q
Sbjct: 497 -------RTRKWEEERRQERREGKSPAVPSGNRRSEAKQKAPAEDIDDGDDDLGFQEPG- 548
Query: 386 IVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGED 444
V ++ GG A + S K R +++AA P + R +VA+TF + D+
Sbjct: 549 -ATVATVDGG-ACAHVSRRSLKAMKRELLAAA--PTHEATRRARWSEVALTFDQTDHPPC 604
Query: 445 TGEEDD-PIVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLV 500
+V+ + N K+ R LID G++ +I+ +DA K+ G+ ++ P ++
Sbjct: 605 VARGGQIAMVVSPTVCNVKLGRVLIDGGAALNILSPAAFDAIKAPGMMLRPSQP----II 660
Query: 501 GFTGDRVLPLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTH 560
G T PLG+ D ++ G T + F V + YNA+LGRP+L F A +
Sbjct: 661 GVTPGHTWPLGHIDLPVTFGGPANFCTERVNFDVADLSLPYNAVLGRPALVKFMAAVHYA 720
Query: 561 HLMLKYPWAGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLT 620
+L +K P + V G+L++A +C R +R SK
Sbjct: 721 YLQMKMPGPDGPISVHGDLKVALACMEQ-----RADRLAVVSK----------------- 758
Query: 621 DLDPRVDPSRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFA 680
P D RL G S P P + +P + L +N +FA
Sbjct: 759 -------PEGGDERL---GTSAPA----APRQRIVTSDEVPVKEEDALISFLRANADVFA 804
Query: 681 WSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKY 740
W DMPG+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV +
Sbjct: 805 WRPADMPGVPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVIH 864
Query: 741 TTWLSNVVLVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYS 800
WL+N V+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAYS
Sbjct: 865 PEWLANPVVVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAYS 924
Query: 801 GYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYI 860
GY+QI M R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y+
Sbjct: 925 GYHQIRMAREDEEKTAFITPIGTYCYTTMPFGLKNAGPTFQRTTRISLGSQLGRNVEAYV 984
Query: 861 DDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPD 920
DD++VKT +DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+
Sbjct: 985 DDLVVKTRNQETLLSDLAETFESLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPE 1044
Query: 921 KCQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAA 980
K +AI M+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ EA+ A
Sbjct: 1045 KIRAIERMRPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFTWTEEAERA 1104
Query: 981 FTQLKEILSSP 991
TQLK LSSP
Sbjct: 1105 LTQLKAYLSSP 1115
>UniRef100_Q9AUU4 Putative polyprotein [Oryza sativa]
Length = 1867
Score = 441 bits (1134), Expect = e-122
Identities = 297/951 (31%), Positives = 452/951 (47%), Gaps = 102/951 (10%)
Query: 50 FVGAMQYAGASDPLYCRCFPMSLGKGPMNWFQNLPSNSLHDWNGVVTCFLAQYSSVRSIP 109
FV +++ + FPM+L W NLP S+H W + F + P
Sbjct: 224 FVASLRNVRWPPRVMANFFPMALKGQARGWLMNLPPASVHSWEDLCQQFTMNFQGTYPRP 283
Query: 110 KTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRLVLATAALAPGPFLTSLDGKP 169
L V++ + ESL++++ RF + I + + + L + K
Sbjct: 284 GEEADLHAVQRGDDESLRSYIQRFCQVRNTIPCIPAHAVIYAFRGGVRHNRMLEKIASKE 343
Query: 170 ANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKEHKDQPSTRESRRKGQDERKP 229
T E A++ E+A T ++ + P+ R RR + +++
Sbjct: 344 PQTTSELFQLADRVARKEEAWTWNSSGSGVTAPAAPG-------PAARSGRRDRRRKKRS 396
Query: 230 KRKKYDSYTPLNSSLSRILRE--KASTDLRDRPPPLLTRGDKLDSKRFCEFHDSPGHNTD 287
+ + SR+ ++ AS ++ P R + ++C H++ H+
Sbjct: 397 AHSDDEGHVLAVEGASRVTQKGRPASDKKKEAGAPSRER----PTGKWCTVHNTSLHDLA 452
Query: 288 ECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERR 347
+C +K E R + + R ERR
Sbjct: 453 DCRAVKSLAE------------------------------------RTRKWEEERRQERR 476
Query: 348 DKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GSNIVDVGSIAGGWAAGGPTNNS 405
+ P S RR ++++ + D G D++ Q G+ I V G A + S
Sbjct: 477 EGKAPAAPSGNRRSEAKQKAPAEDIDDGDDDLGFQEPGATIATVD----GGACAHASRRS 532
Query: 406 QKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGEDTGEEDD-PIVIEALIGNGKI 463
K R +++AA P + R +VA+TF + D+ +V+ + N K+
Sbjct: 533 LKAMKRELLAAA--PTHEATRRARWSEVALTFDQTDHPPCVARGGQIAMVVSPTVCNVKL 590
Query: 464 RRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLVGFTGDRVLPLGYFDTCLSLG 520
R LID G++ +I+ +DA K+ G+ ++ P ++G T PLG+ D ++ G
Sbjct: 591 GRVLIDGGAALNILSPAAFDAIKAPGMVLRPSQP----IIGVTPRHTWPLGHIDLPVTFG 646
Query: 521 DHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTHHLMLKYPWAGRAVPVRGNLE 580
RT + F V + YNA+LGRP+L F + +L +K P G + V G+L+
Sbjct: 647 GSANFRTERVNFDVADLSLPYNAVLGRPALVKFMVAVHYAYLQMKMPGPGGPITVHGDLK 706
Query: 581 MARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLTDLDPRVDPSRDDHRLKPDGE 640
+A +C R E SK P+ D RL
Sbjct: 707 VALACMEQ-----RAEHLAAASK------------------------PAGGDERLSTS-- 735
Query: 641 SHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPGIDPAFCSHRLSV 700
V P +P + L +N +FAW DMPG+ HRL++
Sbjct: 736 -----VPAAPRQRMITCDEVPIKQEDALVSFLRANADVFAWRPADMPGVPREVIEHRLAI 790
Query: 701 DRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANGKWRMC 760
+PV QK R+ + E+Q I+++ LL AG IREV + WL+N V+V KANGK RMC
Sbjct: 791 RPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVIHPEWLANPVVVPKANGKLRMC 850
Query: 761 VDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITD 820
+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAYSGY+QI M R+DEEKTAFIT
Sbjct: 851 IDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAYSGYHQIRMAREDEEKTAFITP 910
Query: 821 RGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADLAVV 880
GTYCYT +PFGLKNA T+QR G +G++VE Y+DD++VKT +DLA
Sbjct: 911 IGTYCYTTMPFGLKNAGPTFQRTTRISLGSQIGRNVEAYVDDLVVKTRNQETLLSDLAET 970
Query: 881 FEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMKSPRTVKEVQQL 940
FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+K +AI M+ P +++VQ +
Sbjct: 971 FESLRTARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPEKIRAIERMRPPSKLRDVQCV 1030
Query: 941 AGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEILSSP 991
G MAA+ RF+ + +ALPL+ LLK F W+ EA+ A TQLK LSSP
Sbjct: 1031 TGCMAALSRFISRLGEKALPLFKLLKRSGPFIWTEEAERALTQLKAYLSSP 1081
>UniRef100_Q8S636 Putative polyprotein [Oryza sativa]
Length = 1722
Score = 433 bits (1113), Expect = e-119
Identities = 289/903 (32%), Positives = 446/903 (49%), Gaps = 104/903 (11%)
Query: 109 PKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRLVLATAALAPGPFLTSLDGK 168
PKT Q L ++Q+ ES++ ++ RF++ + + + + A+A L G + K
Sbjct: 235 PKTQQHLLGIRQRLGESIREYMRRFSQARCQVQDITEASVINAASAGLLEGEVTRKIANK 294
Query: 169 PANTLEEFL------ARAEKFINMEDAVTLR------AASQTLA---IKGPQKMKEHKDQ 213
TLE L AR E+ A+ AA+Q A + P + + Q
Sbjct: 295 EPQTLEHLLRIIDGFARGEEDSKRRQAIQAEYDKASVAAAQAQAQAQVAEPPPLAVRQPQ 354
Query: 214 PSTR-ESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRGDKLDS 272
P+ + + R+GQ ++ + D + ++ + + D + K+
Sbjct: 355 PAIQGQPPRQGQAPMTWRKFRTDRASKAVMAVEEVQALRKEFDAQQASNHQQPVRKKVRK 414
Query: 273 KRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRTPTPPR 332
+C FH H ++C N++ + G + PR P T PR
Sbjct: 415 DLYCAFHGRSLHTMEQCRNIRQR------------------GNVQDPR-PQQGATVEAPR 455
Query: 333 H--RDQTP--PKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVRRQHGSNIVD 388
++QTP +R+ A+RR R P + R+++ ++R H
Sbjct: 456 EAFQEQTPLAEQRQDAQRRVIQVITRADPPSQLSKRQKKM--------QIRMVHS----- 502
Query: 389 VGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPRQKVAITFTEDDYGEDTGEE 448
+++AG P L+ I F +D
Sbjct: 503 -------------------------ITSAGEGAPQYLNQ-----LIYFGPEDAEGVMFPH 532
Query: 449 DDPIVIEALIGNGKIRRTLIDTGSSADIMFYDAYKSLGLSVKDLLPYDHDLVGFTGDRVL 508
DP+VI A I ++RR L+D GSSAD++F +AY +GL+ + L P L GF G+ V
Sbjct: 533 QDPLVISAEIAGFEVRRILVDGGSSADVIFAEAYAKMGLTSQALTPAPASLRGFGGEAVQ 592
Query: 509 PLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTHHLMLKYPW 568
LG ++ G R + F VV+ P YN I GR +LN F I ++L LK P
Sbjct: 593 VLGQALLLIAFGSGENRREEQVLFDVVDIPYNYNTIFGRATLNMFEVISHHNYLELKMPG 652
Query: 569 AGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLTDLDPRVDP 628
+ V+G A S + + K HD+ +HA T +
Sbjct: 653 PTGVIVVKGLQPSAAS--KGDLAIINRAVHNVEVKLHDRP-----KHAPKPTPHGKIIKV 705
Query: 629 SRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPG 688
DD +P +L + ++ + + K+ N +FAWS +++ G
Sbjct: 706 QIDD---------------AVPTKLVSLGGDMGKEEVESILKVPKKNIDIFAWSPDEVGG 750
Query: 689 IDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVV 748
+ H L+V + KP QK R+MSA++Q+ + + +LLRAG+I+E+ + WL+N V
Sbjct: 751 VSIVLIMHHLAVKPEAKPRKQKLRKMSADRQEAAKAEVQKLLRAGVIQEIDHPEWLANPV 810
Query: 749 LVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMH 808
LV+K+NGKWRMCVD+TDLNKACPKD FPLP ID LVD+++G E +S ++AYSGY+QI M+
Sbjct: 811 LVQKSNGKWRMCVDFTDLNKACPKDDFPLPRIDQLVDSTAGCELMSFLNAYSGYHQIHMN 870
Query: 809 RDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTP 868
D KT+FIT T+C+ +PFGL+NA AT+ R++ ++ +G++VE Y+DDI+VK
Sbjct: 871 PLDIPKTSFITPFDTFCHLRMPFGLRNAGATFARLVYKVLCKQLGRNVEAYVDDIVVKIR 930
Query: 869 KGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNM 928
K DHA DL F+ L+ ++LNP+KC FGVR+GK LG++++ RGIE NP+K AI M
Sbjct: 931 KAFDHAIDLQETFDNLRAAGIKLNPEKCVFGVRAGKLLGFLVSERGIEANPEKIDAIQQM 990
Query: 929 KSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEIL 988
K P +V EVQ+LAGR+ A+ RFL +AA R LP + L+ F W+ E AF +LK+ L
Sbjct: 991 KPPSSVHEVQKLAGRIVALSRFLSRAAERGLPFFKTLRGAGKFNWTPECQVAFDELKQYL 1050
Query: 989 SSP 991
SP
Sbjct: 1051 QSP 1053
>UniRef100_Q65WT1 Putative polyprotein [Oryza sativa]
Length = 1992
Score = 419 bits (1076), Expect = e-115
Identities = 290/903 (32%), Positives = 433/903 (47%), Gaps = 105/903 (11%)
Query: 98 FLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRFNKEAGDITGLLPDTRLVLATAALA 157
F + P L V++++ ESL++++ RF + I + + +
Sbjct: 271 FTMNFQGTYPRPGEEADLHAVQRRDDESLRSYIQRFCQVRNTIPCIPAHAVIYAFRGGVR 330
Query: 158 PGPFLTSLDGKPANTLEEFLARAEKFINMEDAVTLRAASQTLAIKGPQKMKEHKDQPSTR 217
L + K T E A++ E+A T +S +A + R
Sbjct: 331 HNRMLEKIASKEPQTTAELFQLADRVARKEEAWTWNPSSSGVAASAAPGSAAQTGRRDRR 390
Query: 218 ESRR--KGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRGDKLDSKRF 275
+R + DE + S + +++AS R+RP + ++
Sbjct: 391 RKKRSVRSDDEGHVLAVEGASRATRKGRPASDKKKEASAPSRERP-----------TGKW 439
Query: 276 CEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGALPRPRSPPPRRTPTPPRHRD 335
C H++ H+ +C +K E R
Sbjct: 440 CTVHNTSLHDLADCRAVKSLAE------------------------------------RT 463
Query: 336 QTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPD-RHGRDEVRRQH-GSNIVDVGSIA 393
+ + R ERR+ P S RR ++++ + D G D++ Q G+ I V
Sbjct: 464 RKWEEERRQERREGKSPAVPSGHRRSEAKQKAPAEDIDDGDDDLGFQEPGATIATVD--- 520
Query: 394 GGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPR-QKVAITFTEDDYGEDTGEEDD-P 451
G A + S K R +++AA P + R +VA+TF + D+
Sbjct: 521 -GGACAHVSRQSLKAMKRELLAAA--PTHEATRRARWSEVALTFDQTDHPPCVARGGQIA 577
Query: 452 IVIEALIGNGKIRRTLIDTGSSADIMF---YDAYKSLGLSVKDLLPYDHDLVGFTGDRVL 508
+V+ I N K+ R LID G++ +I+ +DA K+ G+ ++ P ++G T
Sbjct: 578 MVVSPTICNVKLGRVLIDGGAALNILSPAAFDAIKAPGMVLRPSQP----IIGVTPGHTW 633
Query: 509 PLGYFDTCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTHHLMLKYPW 568
PLG+ D ++ G RT + F V + YNA+LGRP+L F A + +L +K P
Sbjct: 634 PLGHIDLPVTFGGPANFRTERVNFDVADLSLPYNAVLGRPALVKFMAAVHYAYLQMKMPG 693
Query: 569 AGRAVPVRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLTDLDPRVDP 628
+ V G+L++A +C R +R SK P
Sbjct: 694 PDGPISVHGDLKVALACMEQ-----RADRLAVVSK------------------------P 724
Query: 629 SRDDHRLKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPG 688
D RL G S P P +P D + L +N +FAW DMPG
Sbjct: 725 EGGDERL---GTSAPA----APRQRIVTGDEVPEDA---LVSFLRANADVFAWRPADMPG 774
Query: 689 IDPAFCSHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVV 748
+ HRL+V +PV QK R+ + E+Q I+++ LL AG IREV + WL+N V
Sbjct: 775 VPREVIEHRLAVRPGARPVRQKVRRQAPERQAFIREEVARLLEAGFIREVIHPEWLANPV 834
Query: 749 LVKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMH 808
+V KANGK RMC+DYTDLNKACPKDP+PLP ID +VD+++G + L +DAYSGY+QI M
Sbjct: 835 VVPKANGKLRMCIDYTDLNKACPKDPYPLPRIDQIVDSTAGCDLLCFLDAYSGYHQIRMA 894
Query: 809 RDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTP 868
R+DEEKTAFIT GTYCYT +PFGLKNA T+QR G +G++VE Y+DD++VKT
Sbjct: 895 REDEEKTAFITPVGTYCYTSMPFGLKNAGPTFQRTTRISLGSQIGRNVEAYVDDLVVKTR 954
Query: 869 KGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNM 928
+DLA FE L+ ++LNPDKC FGV +GK LG++++ RGIE NP+K +AI M
Sbjct: 955 NQETLLSDLAETFESLRSARIKLNPDKCVFGVPAGKLLGFLVSARGIEANPEKIRAIERM 1014
Query: 929 KSPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEIL 988
+ P +++VQ + G MAA+ RF+ + +ALPL+ LLK F W+ E + A TQLK L
Sbjct: 1015 RPPSKLRDVQCVTGCMAALSRFISRLGEKALPLFKLLKRSGPFAWTEEVERALTQLKAYL 1074
Query: 989 SSP 991
SSP
Sbjct: 1075 SSP 1077
>UniRef100_Q8SA85 Prpol [Zea mays]
Length = 1317
Score = 404 bits (1037), Expect = e-110
Identities = 219/594 (36%), Positives = 334/594 (55%), Gaps = 21/594 (3%)
Query: 395 GWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPRQKVAITFTEDDYGEDTGEEDDPIVI 454
G ++ P N QK+ + + G P + + ITF+++D +D +VI
Sbjct: 22 GGSSSEPANKKQKKEAQRRVQHVGVQGP-FIKSRWSHIPITFSQEDLQLKDYPHNDAMVI 80
Query: 455 EALIGNGKIRRTLIDTGSSADIMFYDAYKSLGLSVKDLLPYDHDLVGFTGDRVLPLGYFD 514
+I + L+DTGS+ADI+F A++ + + H L GF G +++ LG
Sbjct: 81 SCVIKGFLVHNVLVDTGSAADIIFAKAFRQMQEPEDKIHDATHPLCGFGGRQIVALGKIT 140
Query: 515 TCLSLGDHRICRTIKARFLVVECPTAYNAILGRPSLNTFRAIISTHHLMLKYPWAGRAVP 574
++ G RT + F +V+ YNAI+GR +LN F AI+ +L +K P +
Sbjct: 141 MSVTFGFINNTRTEQVVFDIVDMEYPYNAIIGRGTLNAFEAILHPAYLCMKIPSDQGPIA 200
Query: 575 VRGNLEMARSCYNSSCRLAREERKRKKSKAHDQNGNYHVQHANLLTDLDPRVDPSRDDHR 634
+ G+ E AR R + D +++ A R + + +
Sbjct: 201 IHGSQEAAR---------------RAEGNWTDSKAIHNIDGAEACEQYKFRREKAASADQ 245
Query: 635 LKPDGESHPI*VGPLPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPGIDPAFC 694
KP + + E L L + K + + L +N +FAWS+ D+ G++
Sbjct: 246 PKP-----MLLCEDIAEQKVLLGSQLSEEQEKTLIRFLFNNKDVFAWSANDLCGVNRDVI 300
Query: 695 SHRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKAN 754
H L+VD ++P Q+ R+MS +K + + + LL AG+IREVKY WL+N V+VKKAN
Sbjct: 301 EHSLNVDPSFRPRKQRLRKMSDDKAEGARNEVKRLLSAGVIREVKYPEWLANTVMVKKAN 360
Query: 755 GKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEK 814
GKWRMC+D+TDLNKACPKD FPLP ID+LVD ++ E +SL+D YSGY+QI M ++DE K
Sbjct: 361 GKWRMCIDFTDLNKACPKDEFPLPRIDSLVDAAASSELMSLLDCYSGYHQIWMKKEDEPK 420
Query: 815 TAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHA 874
T+FIT GTYCY +P GLKNA ++ RM ++ +G++V Y+DDIIVK+ K +H
Sbjct: 421 TSFITPSGTYCYLRMPEGLKNAGGSFSRMTAKVLQSQIGRNVLTYVDDIIVKSTKQENHI 480
Query: 875 ADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMKSPRTV 934
ADL F ++ ++LNP+KC FGV+ GKFLG +++ +GIE NP K +AI+ M+ P T
Sbjct: 481 ADLQETFASFRQAGLKLNPEKCVFGVKKGKFLGCLVSTKGIEANPSKIEAILRMEPPTTK 540
Query: 935 KEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEIL 988
K Q+L GR+A++ RF+ ++A R LP + +LK+ F+W AF +LK+ L
Sbjct: 541 KGAQRLTGRLASLNRFISRSAERNLPFFEVLKSAEVFQWGPIQQKAFEELKQYL 594
Database: uniref100
Posted date: Jan 5, 2005 1:24 AM
Number of letters in database: 848,049,833
Number of sequences in database: 2,790,947
Lambda K H
0.325 0.138 0.443
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,944,354,994
Number of Sequences: 2790947
Number of extensions: 130884096
Number of successful extensions: 672486
Number of sequences better than 10.0: 30372
Number of HSP's better than 10.0 without gapping: 2771
Number of HSP's successfully gapped in prelim test: 27738
Number of HSP's that attempted gapping in prelim test: 579078
Number of HSP's gapped (non-prelim): 62468
length of query: 1706
length of database: 848,049,833
effective HSP length: 141
effective length of query: 1565
effective length of database: 454,526,306
effective search space: 711333668890
effective search space used: 711333668890
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 82 (36.2 bits)
Lotus: description of TM0059.7


