

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0400a.3
(419 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
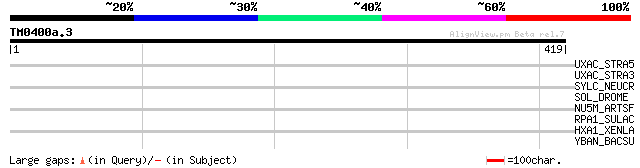
Score E
Sequences producing significant alignments: (bits) Value
UXAC_STRA5 (Q8E0M9) Uronate isomerase (EC 5.3.1.12) (Glucuronate... 32 2.3
UXAC_STRA3 (Q8E6A3) Uronate isomerase (EC 5.3.1.12) (Glucuronate... 32 2.3
SYLC_NEUCR (P10857) Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.... 32 2.3
SOL_DROME (P27398) Small optic lobes protein (EC 3.4.22.-) 32 2.3
NU5M_ARTSF (Q37710) NADH-ubiquinone oxidoreductase chain 5 (EC 1... 32 4.0
RPA1_SULAC (P11512) DNA-directed RNA polymerase subunit A' (EC 2... 31 5.2
HXA1_XENLA (Q08821) Homeobox protein Hox-A1 (Hox.lab2) (Fragment) 31 6.8
YBAN_BACSU (P50865) Hypothetical protein ybaN precursor (ORFX) 30 8.8
>UXAC_STRA5 (Q8E0M9) Uronate isomerase (EC 5.3.1.12) (Glucuronate
isomerase) (Uronic isomerase)
Length = 466
Score = 32.3 bits (72), Expect = 2.3
Identities = 34/117 (29%), Positives = 47/117 (40%), Gaps = 19/117 (16%)
Query: 62 TRIMFKPIDEPHV-DLIATV-SGPLDHKPEESITGNALFRWQRILRMRSCAYYPRYGFGA 119
T I+F+ DE + DL V G + ++ E S +WQ + M C Y +YGF
Sbjct: 238 TEIVFEQTDELELNDLFNKVCEGYIPNQSEIS-------KWQTAVFMELCRLYKKYGFVT 290
Query: 120 FGVFPLLLRKREFSEDYGLMGLRYGSGNLSFGVTLM--------PFAKKDELPKSAW 168
F L + S + +G G +L V L KKD LPK W
Sbjct: 291 QVHFGAL--RNNHSTIFEKLGADVGVDSLGDQVALTVNMNRLLDSLVKKDSLPKMIW 345
>UXAC_STRA3 (Q8E6A3) Uronate isomerase (EC 5.3.1.12) (Glucuronate
isomerase) (Uronic isomerase)
Length = 466
Score = 32.3 bits (72), Expect = 2.3
Identities = 34/117 (29%), Positives = 47/117 (40%), Gaps = 19/117 (16%)
Query: 62 TRIMFKPIDEPHV-DLIATV-SGPLDHKPEESITGNALFRWQRILRMRSCAYYPRYGFGA 119
T I+F+ DE + DL V G + ++ E S +WQ + M C Y +YGF
Sbjct: 238 TEIVFEQTDELELNDLFNKVCEGYIPNQSEIS-------KWQTAVFMELCRLYKKYGFVT 290
Query: 120 FGVFPLLLRKREFSEDYGLMGLRYGSGNLSFGVTLM--------PFAKKDELPKSAW 168
F L + S + +G G +L V L KKD LPK W
Sbjct: 291 QVHFGAL--RNNHSTIFEKLGADVGVDSLGDQVALTVNMNRLLDSLVKKDSLPKMIW 345
>SYLC_NEUCR (P10857) Leucyl-tRNA synthetase, cytoplasmic (EC
6.1.1.4) (Leucine--tRNA ligase) (LeuRS)
Length = 1123
Score = 32.3 bits (72), Expect = 2.3
Identities = 15/39 (38%), Positives = 21/39 (53%), Gaps = 2/39 (5%)
Query: 194 LMNWSCAMAYGVGSGSPLSPSFNFSLELVKSSQFVASFY 232
L W+CA YG+GS P P NF +E + S ++Y
Sbjct: 608 LKQWACARTYGLGSKLPWDP--NFLVESLSDSTVYMAYY 644
>SOL_DROME (P27398) Small optic lobes protein (EC 3.4.22.-)
Length = 1594
Score = 32.3 bits (72), Expect = 2.3
Identities = 28/117 (23%), Positives = 48/117 (40%), Gaps = 7/117 (5%)
Query: 219 LELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYIDFGFELQTSVDDAIATNNIS- 277
L+L + Q + H + Q++ + P +E +++ + V +IS
Sbjct: 678 LQLQQQQQQQQQHHHHHLQQQQAEAPRDEPWTCKKCTLVNYSTAMACVVCGGSKLKSISS 737
Query: 278 --DSTFRIGASWQANKNFLLKAKVGPRISSMALAFKSWWKPSFTFSISATRDRADGQ 332
D T R G W + L + P S A KS +P + ++ A R+R DGQ
Sbjct: 738 IEDMTLRKGEFWTCSHCTLKNSLHSPVCS----ACKSHRQPQLSMAMEAVRERPDGQ 790
>NU5M_ARTSF (Q37710) NADH-ubiquinone oxidoreductase chain 5 (EC
1.6.5.3)
Length = 541
Score = 31.6 bits (70), Expect = 4.0
Identities = 26/87 (29%), Positives = 39/87 (43%), Gaps = 12/87 (13%)
Query: 132 FSEDYGLMGLRYGSGNLSFGVTLMPFAKKDELPKSAWLVSKMGRLTAGVQYEPQHGNAKL 191
+S+ + M L Y S ++ L F K +LP SAWL + M T +
Sbjct: 151 YSKSWDYMFLSYFSLSIMLLFILSSFTKSAQLPFSAWLPAAMAAPT------------PV 198
Query: 192 SNLMNWSCAMAYGVGSGSPLSPSFNFS 218
S+L++ S + G+ LSPSF S
Sbjct: 199 SSLVHSSTLVTAGIYLMIRLSPSFEES 225
>RPA1_SULAC (P11512) DNA-directed RNA polymerase subunit A' (EC
2.7.7.6)
Length = 880
Score = 31.2 bits (69), Expect = 5.2
Identities = 20/57 (35%), Positives = 32/57 (56%), Gaps = 5/57 (8%)
Query: 143 YGSGNLSFGVTLMPFAKKDEL-PKSAWLVSKMGRLTAGVQYEPQHGNAKLSNLMNWS 198
+G N+S G P A KDE+ P +++V K G L GV + GN + ++++WS
Sbjct: 563 HGPANISKG----PRACKDEICPHDSFIVIKNGLLLEGVFDKKAIGNQQPESMLHWS 615
>HXA1_XENLA (Q08821) Homeobox protein Hox-A1 (Hox.lab2) (Fragment)
Length = 240
Score = 30.8 bits (68), Expect = 6.8
Identities = 20/80 (25%), Positives = 35/80 (43%)
Query: 198 SCAMAYGVGSGSPLSPSFNFSLELVKSSQFVASFYQHMVVQRRVKNPLEENGVIGITNYI 257
SC YG+ + SP F E SS F S Y + V++ ++ + G +YI
Sbjct: 6 SCGSNYGMQNFSPGYSHFPIHQETEVSSGFPQSVYSGNIASSVVQHQQHQSYIEGSAHYI 65
Query: 258 DFGFELQTSVDDAIATNNIS 277
+ + ++ A NN++
Sbjct: 66 HHSYGPEQNLSVANYNNNVA 85
>YBAN_BACSU (P50865) Hypothetical protein ybaN precursor (ORFX)
Length = 254
Score = 30.4 bits (67), Expect = 8.8
Identities = 19/65 (29%), Positives = 32/65 (49%), Gaps = 2/65 (3%)
Query: 251 IGITNYIDFGFELQTSVDDAIATNNISDSTFRIGASWQANKNFLLK--AKVGPRISSMAL 308
I +T I +G E + + + N I ++TF + ASW + K G +I SM
Sbjct: 59 ISLTFDISWGDERAEPILNTLKANGIKNATFFLSASWAERHPDTVARIVKDGHQIGSMGY 118
Query: 309 AFKSW 313
A+K++
Sbjct: 119 AYKNY 123
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,293,832
Number of Sequences: 164201
Number of extensions: 2138890
Number of successful extensions: 4864
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 4860
Number of HSP's gapped (non-prelim): 8
length of query: 419
length of database: 59,974,054
effective HSP length: 113
effective length of query: 306
effective length of database: 41,419,341
effective search space: 12674318346
effective search space used: 12674318346
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Lotus: description of TM0400a.3


