

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
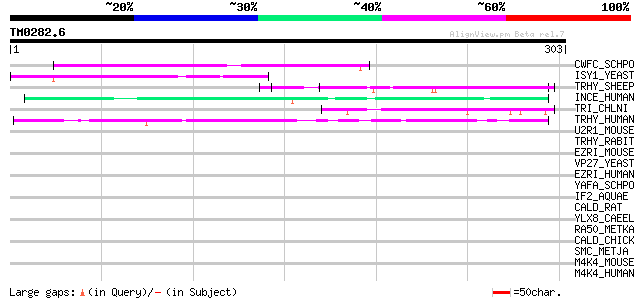
Query= TM0282.6
(303 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CWFC_SCHPO (O74370) Cell cycle control protein cwf12 108 2e-23
ISY1_YEAST (P21374) Splicing factor ISY1 (Unknown transcript 3 p... 72 1e-12
TRHY_SHEEP (P22793) Trichohyalin 55 2e-07
INCE_HUMAN (Q9NQS7) Inner centromere protein 48 3e-05
TRI_CHLNI (Q7M3Y3) Troponin I (TnI) 45 2e-04
TRHY_HUMAN (Q07283) Trichohyalin 44 5e-04
U2R1_MOUSE (Q64707) U2 small nuclear ribonucleoprotein auxiliary... 42 0.002
TRHY_RABIT (P37709) Trichohyalin 42 0.002
EZRI_MOUSE (P26040) Ezrin (p81) (Cytovillin) (Villin 2) 41 0.004
VP27_YEAST (P40343) Vacuolar protein sorting-associated protein ... 40 0.005
EZRI_HUMAN (P15311) Ezrin (p81) (Cytovillin) (Villin 2) 40 0.005
YAFA_SCHPO (Q09863) Hypothetical protein C29E6.10c in chromosome I 40 0.007
IF2_AQUAE (O67825) Translation initiation factor IF-2 40 0.007
CALD_RAT (Q62736) Non-muscle caldesmon (CDM) (L-caldesmon) 40 0.007
YLX8_CAEEL (P46504) Hypothetical protein F23F12.8 in chromosome ... 40 0.009
RA50_METKA (Q8TXI4) DNA double-strand break repair rad50 ATPase 40 0.009
CALD_CHICK (P12957) Caldesmon (CDM) 40 0.009
SMC_METJA (Q59037) Chromosome partition protein smc homolog 39 0.016
M4K4_MOUSE (P97820) Mitogen-activated protein kinase kinase kina... 39 0.016
M4K4_HUMAN (O95819) Mitogen-activated protein kinase kinase kina... 39 0.016
>CWFC_SCHPO (O74370) Cell cycle control protein cwf12
Length = 217
Score = 108 bits (269), Expect = 2e-23
Identities = 64/183 (34%), Positives = 107/183 (57%), Gaps = 18/183 (9%)
Query: 25 KPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQNEGLGEHRLRDLNDEINKLIRE 84
KP E+RP + + +K R ++++I RK++ IQ+ L E+++RDLND IN+L+RE
Sbjct: 11 KPTEKRPKDIKSIKSVPICEKHRASVVKDISRKISRIQSATLPEYQIRDLNDAINRLMRE 70
Query: 85 KSHWERRIVELGGPNYTRHSAKMTDLDG-NIVDVPNPSGRGPGYRYFGAAKKLPGVRELF 143
K WE +I +LGG NY + AK+ + +G I D+ + YRY+G A++LPGV+ELF
Sbjct: 71 KHEWEVQIRDLGGINYLYNKAKLFEDEGEQISDIDD-------YRYYGRARELPGVKELF 123
Query: 144 DKPPE-LRKRRTRYDIYK-RIDAAYYGY-RDDEDGILERLEPSAEAAMRR-------EAA 193
+ + +R+ + ++ K R+DA Y+GY ++ +LE E E + E
Sbjct: 124 EADMSFIPERQRKQEMQKRRLDAWYFGYIPPAQESLLEDFEAKIEEQQHKHLENLGDEVE 183
Query: 194 EEW 196
++W
Sbjct: 184 QDW 186
>ISY1_YEAST (P21374) Splicing factor ISY1 (Unknown transcript 3
protein)
Length = 235
Score = 72.4 bits (176), Expect = 1e-12
Identities = 47/145 (32%), Positives = 76/145 (52%), Gaps = 9/145 (6%)
Query: 1 MARNEEKAQSMLNRFITMKAEE----KKKPKERRPFLATECRDLSEADKWRQQIMREIGR 56
M+RN +KA S+L RF +AE K + +RP ++ + + EA++W++Q+ +EI +
Sbjct: 1 MSRNVDKANSVLVRFQEQQAESAGGYKDYSRYQRPRNVSKVKSIKEANEWKRQVSKEIKQ 60
Query: 57 KVAEIQNEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVD 116
K I + L E ++ +LNDE+N L +E W+ I K L+ + V
Sbjct: 61 KSTRIYDPSLNEMQIAELNDELNNLFKEWKRWQWHI----DHTLMEKKTKRKRLEDSHV- 115
Query: 117 VPNPSGRGPGYRYFGAAKKLPGVRE 141
+ N G RYFG A +LP V+E
Sbjct: 116 LMNSGKLINGKRYFGRALELPEVKE 140
>TRHY_SHEEP (P22793) Trichohyalin
Length = 1549
Score = 55.5 bits (132), Expect = 2e-07
Identities = 42/165 (25%), Positives = 82/165 (49%), Gaps = 8/165 (4%)
Query: 137 PGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEW 196
P RE + +LR + + KR YR+ E E E ++RE E+
Sbjct: 384 PAQREQVREEEQLRLKEEKLQREKRRQERERQYREVELQREEERLQREEEQLQREEREKR 443
Query: 197 YRLDRIRQEARKGVKSGEVAEVSAAVREILREEEE-------DVVEEERMRGREMKERLE 249
R +R +Q K V+ E ++ RE R+E E ++ EEE+++ +E ++R +
Sbjct: 444 RRQEREKQYLEK-VELWEEEQLQREEREKRRQEREKQYLEKVELREEEQLQRQEREKRRQ 502
Query: 250 ENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSE 294
E ER+++ V L +E+++++ EK++ E +Y+ + + E+ +
Sbjct: 503 ERERQYLEKVELQEEEQLQREEREKRRQERERQYLEKVELQEEEQ 547
Score = 52.8 bits (125), Expect = 1e-06
Identities = 33/135 (24%), Positives = 76/135 (55%), Gaps = 9/135 (6%)
Query: 170 RDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREE 229
++ E LE++E E ++R+ E+ R +R +Q K V+ E ++ R+ R+E
Sbjct: 530 QERERQYLEKVELQEEEQLQRQEREK-RRQEREKQYLEK-VELQEEEQLQRQERQKRRQE 587
Query: 230 EE-------DVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNK 282
E ++ EEE+++ +E ++R +E ER+++ V L +E+++++ EK++ E +
Sbjct: 588 REKQYLEKVELQEEEQLQRQEREKRRQERERQYLEKVELQEEEQVQRQEREKRRQERERQ 647
Query: 283 YVSEDLMDEQSEAKD 297
Y+ ++L ++ ++
Sbjct: 648 YLEKELQRQEERLQE 662
Score = 52.4 bits (124), Expect = 1e-06
Identities = 38/138 (27%), Positives = 72/138 (51%), Gaps = 15/138 (10%)
Query: 170 RDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREE 229
++ E LE++E E ++R+ E+ R +R RQ K V+ E +V RE R+E
Sbjct: 586 QEREKQYLEKVELQEEEQLQRQEREK-RRQERERQYLEK-VELQEEEQVQRQEREKRRQE 643
Query: 230 -------------EEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKK 276
EE + EEE++ E ++R +E ER+++ V L +E+++++ EK++
Sbjct: 644 RERQYLEKELQRQEERLQEEEQLLREEREKRRQERERQYLEKVELQEEEQLQREEREKRR 703
Query: 277 MELLNKYVSEDLMDEQSE 294
E +Y+ ++ + Q E
Sbjct: 704 QERERQYLEKEELQRQEE 721
Score = 51.6 bits (122), Expect = 2e-06
Identities = 35/132 (26%), Positives = 73/132 (54%), Gaps = 9/132 (6%)
Query: 170 RDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREE 229
++ E LE++E E ++RE E+ R +R +Q K V+ E ++ RE R+E
Sbjct: 446 QEREKQYLEKVELWEEEQLQREEREK-RRQEREKQYLEK-VELREEEQLQRQEREKRRQE 503
Query: 230 EE-------DVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNK 282
E ++ EEE+++ E ++R +E ER+++ V L +E+++++ EK++ E +
Sbjct: 504 RERQYLEKVELQEEEQLQREEREKRRQERERQYLEKVELQEEEQLQRQEREKRRQEREKQ 563
Query: 283 YVSEDLMDEQSE 294
Y+ + + E+ +
Sbjct: 564 YLEKVELQEEEQ 575
Score = 45.8 bits (107), Expect = 1e-04
Identities = 38/155 (24%), Positives = 79/155 (50%), Gaps = 16/155 (10%)
Query: 144 DKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWY---RLD 200
D+ + K+ RYD R ++ +E+ R A+ A R + EE + +
Sbjct: 349 DQGRQRLKQEQRYDQNWR-------WQLEEESQRRRYTLYAKPAQREQVREEEQLRLKEE 401
Query: 201 RIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREM-KERLEENEREFVVHV 259
++++E R+ + + EV E+ REEE EEE+++ E K R +E E++++ V
Sbjct: 402 KLQREKRRQERERQYREV-----ELQREEERLQREEEQLQREEREKRRRQEREKQYLEKV 456
Query: 260 PLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSE 294
L +E+++++ EK++ E +Y+ + + E+ +
Sbjct: 457 ELWEEEQLQREEREKRRQEREKQYLEKVELREEEQ 491
Score = 40.0 bits (92), Expect = 0.007
Identities = 35/159 (22%), Positives = 73/159 (45%), Gaps = 7/159 (4%)
Query: 134 KKLPGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAA 193
++L +L + E R++ ++++ E+ R E + + E
Sbjct: 658 ERLQEEEQLLREEREKRRQERERQYLEKVELQEEEQLQREEREKRRQERERQYLEKEELQ 717
Query: 194 EEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKERLEENER 253
+ RL R +++ ++ E E VRE EEE EE+R++ + R + +R
Sbjct: 718 RQEERLQREKEQLQR-----EDREKRRQVRERKYLEEELQQEEDRLQREKQLLREDREKR 772
Query: 254 EFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQ 292
+++ V L ++E E++ EK++ E +Y E+L+ E+
Sbjct: 773 QYLEKVEL--QREEEQLQREKRRQERERQYREEELLREE 809
Score = 36.6 bits (83), Expect = 0.079
Identities = 62/312 (19%), Positives = 131/312 (41%), Gaps = 34/312 (10%)
Query: 1 MARNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAE 60
+ R E + + + EE+ + +ER E + L ++ QQ+ +E RK E
Sbjct: 1039 LRRQERDRKFREEEQLLQEREEQLRRQERDRKFREEEQQLRLLER-EQQLRQERNRKFRE 1097
Query: 61 IQ--NEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSA--KMTDLDGNIVD 116
Q E + RL++ ++ + K H E ++++ R K + + +
Sbjct: 1098 EQLLREREEQLRLQEGEPQLRQKRDRKFHEEEQLLQEREEQLRRQERDRKFREEAQILKE 1157
Query: 117 VPNPSGRGPGYRYFGAAKKLPGVRELF---DKPPELRKRRTRYDIYKRIDAAYYGYRDDE 173
R R F ++L RE ++ P+LR+ R R +R++E
Sbjct: 1158 REEQLRRQERDRKFREEEQLLQEREELRRQEREPQLRQERDRK------------FREEE 1205
Query: 174 DGILERLE---PSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEE 230
+ ER + E +R+E +++ +++ QE + ++ E ++L+E E
Sbjct: 1206 QLLQEREKLRRQEREPQLRQERDRKFHEEEQLLQEREEQLRRQERDRKFREEAQLLQERE 1265
Query: 231 EDVVEEERMRG--------REMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNK 282
E + +ER R +E +E+L ER+ +E+ +++ + ++ E K
Sbjct: 1266 EQLRRQERDRKFREEEQLLQEREEQLRRQERDRKFR---EEEQLLQEREEQLRRQERDRK 1322
Query: 283 YVSEDLMDEQSE 294
+ E+ + ++SE
Sbjct: 1323 FREEEQLLKESE 1334
Score = 34.7 bits (78), Expect = 0.30
Identities = 56/299 (18%), Positives = 111/299 (36%), Gaps = 22/299 (7%)
Query: 6 EKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQNEG 65
EK + + + EK++ + R +L E + E + +Q++RE K + +
Sbjct: 622 EKVELQEEEQVQRQEREKRRQERERQYLEKELQRQEERLQEEEQLLREEREKRRQERERQ 681
Query: 66 LGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSGRGP 125
E +++ + REK ER L R ++ + R
Sbjct: 682 YLEKVELQEEEQLQREEREKRRQERERQYLEKEELQRQEERLQREKEQLQREDREKRRQV 741
Query: 126 GYRYFGAAKKLPGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAE 185
R + + L + LR+ R + ++++ L+R E +
Sbjct: 742 RERKYLEEELQQEEDRLQREKQLLREDREKRQYLEKVE-------------LQREEEQLQ 788
Query: 186 AAMRREAAEEWYRLDR-IRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREM 244
RR+ E YR + +R+E R K ++ R R+E E +EEE ++ +
Sbjct: 789 REKRRQERERQYREEELLREEERLHRKEQQLQREECEKRR--RQELERQLEEEELQRLDR 846
Query: 245 KERLEENE------REFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAKD 297
K + +++ R V+ + KE + + + E + L DEQ E ++
Sbjct: 847 KRQFRDDDQHQNEVRNSRVYSKHRENKEKSRQLDDSWVRESQFQQDLRPLQDEQEEKRE 905
Score = 33.5 bits (75), Expect = 0.67
Identities = 29/128 (22%), Positives = 55/128 (42%), Gaps = 11/128 (8%)
Query: 181 EPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMR 240
E + E +R + RL R QE G + + + E+L E+ +E+R R
Sbjct: 139 ELAEEEELREKQVRREQRLQRREQEEYGGEEELQQRPKGRELEELLNREQRFERQEQRER 198
Query: 241 GR-----------EMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLM 289
R E++ER EE + + L E+ E+ L+K+K L + + ++
Sbjct: 199 QRLQVEQQQRQRGELRERQEEVQLQKRETQELQRERLEEEQQLQKQKRGLEERLLEQERR 258
Query: 290 DEQSEAKD 297
+++ K+
Sbjct: 259 EQELRRKE 266
Score = 32.0 bits (71), Expect = 1.9
Identities = 27/134 (20%), Positives = 64/134 (47%), Gaps = 22/134 (16%)
Query: 169 YRDDEDGILERLEPSAEAAMRREAAEEWYRLDRI-RQEARKGVKSGEVAEVSAAVREILR 227
+R++E + L+ S E R+E +++ + + R+ + ++ E+ V + ++ R
Sbjct: 1323 FREEE----QLLKESEEQLRRQERDRKFHEKEHLLREREEQQLRRQELEGVFSQEEQLRR 1378
Query: 228 EEEEDVVEEERMRGR--------------EMKERLEENEREFVVHVPLPDEKEIEKMVLE 273
E+E+ +R R R E K R++E +R+F+ +E E++
Sbjct: 1379 AEQEEEQRRQRQRDRKFLEEEQSLQREREEEKRRVQEQDRKFLEQEEQLHREEQEEL--- 1435
Query: 274 KKKMELLNKYVSED 287
+++ +L +Y +E+
Sbjct: 1436 RRRQQLDQQYRAEE 1449
Score = 31.6 bits (70), Expect = 2.5
Identities = 23/85 (27%), Positives = 46/85 (54%), Gaps = 11/85 (12%)
Query: 200 DRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMK--ERLEENEREFVV 257
DR R++ R+ E + R + R+E E++ EEE +R ++++ +RL+ E+E
Sbjct: 111 DRRREDQRRF----EPQDRQLEERRLKRQELEELAEEEELREKQVRREQRLQRREQE--- 163
Query: 258 HVPLPDEKEIEKMVLEKKKMELLNK 282
E+E+++ ++ ELLN+
Sbjct: 164 --EYGGEEELQQRPKGRELEELLNR 186
Score = 30.8 bits (68), Expect = 4.3
Identities = 25/114 (21%), Positives = 50/114 (42%), Gaps = 4/114 (3%)
Query: 169 YRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVRE---- 224
+R++ + ER E R+ EE L ++ R+ + + E ++E
Sbjct: 1254 FREEAQLLQEREEQLRRQERDRKFREEEQLLQEREEQLRRQERDRKFREEEQLLQEREEQ 1313
Query: 225 ILREEEEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKME 278
+ R+E + EE +E +E+L ER+ H +E E+ L ++++E
Sbjct: 1314 LRRQERDRKFREEEQLLKESEEQLRRQERDRKFHEKEHLLREREEQQLRRQELE 1367
Score = 30.0 bits (66), Expect = 7.4
Identities = 30/128 (23%), Positives = 57/128 (44%), Gaps = 16/128 (12%)
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEE 237
ER E R+E ++ +++ QE + ++ E ++L+E EE + +E
Sbjct: 984 ERDRKFREELSRQERDRKFREEEQLLQEREEQLRRQERDRKFREEEQLLQEREEQLRRQE 1043
Query: 238 RMRG--------REMKERLEENEREFVVHVPLPDEKEIEKMVLEKK---KMELLNKYVSE 286
R R +E +E+L ER+ E+E + +LE++ + E K+ E
Sbjct: 1044 RDRKFREEEQLLQEREEQLRRQERDRKFR-----EEEQQLRLLEREQQLRQERNRKFREE 1098
Query: 287 DLMDEQSE 294
L+ E+ E
Sbjct: 1099 QLLREREE 1106
Score = 30.0 bits (66), Expect = 7.4
Identities = 34/140 (24%), Positives = 62/140 (44%), Gaps = 20/140 (14%)
Query: 169 YRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILRE 228
+R++E + ER E R+ EE L ++ R+ + + E +LRE
Sbjct: 1300 FREEEQLLQEREEQLRRQERDRKFREEEQLLKESEEQLRRQERDRKFHEKE----HLLRE 1355
Query: 229 EEEDVV----------EEERMRGREMKE---RLEENEREFV-VHVPLPDEKEIEKMVLEK 274
EE + +EE++R E +E R + +R+F+ L E+E EK +++
Sbjct: 1356 REEQQLRRQELEGVFSQEEQLRRAEQEEEQRRQRQRDRKFLEEEQSLQREREEEKRRVQE 1415
Query: 275 KKMELLNKYVSEDLMDEQSE 294
+ + L + E L E+ E
Sbjct: 1416 QDRKFLEQ--EEQLHREEQE 1433
>INCE_HUMAN (Q9NQS7) Inner centromere protein
Length = 919
Score = 48.1 bits (113), Expect = 3e-05
Identities = 60/289 (20%), Positives = 117/289 (39%), Gaps = 25/289 (8%)
Query: 9 QSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQNEGLGE 68
+S++ FI + PKE+ R EA++ R+Q + E R+
Sbjct: 513 RSVMKSFIKRNTPLRMDPKEKERQRLENLRRKEEAEQLRRQKVEEDKRR----------- 561
Query: 69 HRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSGRGPGYR 128
RL ++ + + +R+ R+ ++ + K +D +
Sbjct: 562 -RLEEVKLKREERLRKVLQARERVEQMKEEKKKQIEQKFAQIDEKTEKAKEERLAEEKAK 620
Query: 129 YFGAAKKLPGVRELFDKPPELRKRR--TRYDIYKRIDAAYYGYRDDEDGILERLEPSAEA 186
AAKK+ V + + R+ R + + +R +++E ERL +AEA
Sbjct: 621 KKAAAKKMEEVEARRKQEEDARRLRWLQQEEEERRHQELLQKKKEEEQ---ERLRKAAEA 677
Query: 187 AMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGRE-MK 245
E E+ +R QE R+ + + + RE+E + E+ER R +E ++
Sbjct: 678 KRLAEQREQ----ERREQERREQERREQERREQERREQERREQERQLAEQERRREQERLQ 733
Query: 246 ERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSE 294
E ERE + + ++++++ + EKKK E + L +EQ +
Sbjct: 734 AERELQEREKALRL---QKEQLQRELEEKKKKEEQQRLAERQLQEEQEK 779
>TRI_CHLNI (Q7M3Y3) Troponin I (TnI)
Length = 292
Score = 45.1 bits (105), Expect = 2e-04
Identities = 47/143 (32%), Positives = 68/143 (46%), Gaps = 23/143 (16%)
Query: 171 DDEDGILERLEPS--AEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILRE 228
D+ED EP+ AEA RR +E +EA E +R +E
Sbjct: 41 DEEDSYSAPAEPAYDAEAENRRRQQQE-------EEEAAARAAEEEYNRQQEELRRQRQE 93
Query: 229 EEEDVVEEERMRGREMKERL----EENEREFVVHVPLPDEKEIEKMVL-----EKKKM-- 277
EE EEE+ R +E +ERL EE ERE ++K+ +K L EKKKM
Sbjct: 94 EERQRREEEQRRQQEEEERLRLEREEQEREEEARRMAEEQKKKKKKGLGGLSPEKKKMLK 153
Query: 278 ELLNKYVSEDLMDE---QSEAKD 297
+L+ + +EDL +E ++EAK+
Sbjct: 154 KLIMQKAAEDLANEAKAKAEAKE 176
>TRHY_HUMAN (Q07283) Trichohyalin
Length = 1898
Score = 43.9 bits (102), Expect = 5e-04
Identities = 67/297 (22%), Positives = 123/297 (40%), Gaps = 52/297 (17%)
Query: 3 RNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQ 62
+ E+AQ + ++ +++++ +E+R RD KWR Q+ E R+ +
Sbjct: 859 QRRERAQQLQEEEDGLQEDQERRRQEQR-------RD----QKWRWQLEEERKRRRHTLY 907
Query: 63 NEGLGEHRLRD----LNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVP 118
+ + +LR L +E +L RE+ RR + Y + + + +
Sbjct: 908 AKPALQEQLRKEQQLLQEEEEELQREEREKRRRQEQ--ERQYREEEQLQQEEEQLLREER 965
Query: 119 NPSGRGPGYRYFGAAKKLPGVRE-LFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGIL 177
R R + KKL E L + PE R+R+ R YR++E+
Sbjct: 966 EKRRRQERERQYRKDKKLQQKEEQLLGEEPEKRRRQEREK----------KYREEEE--- 1012
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEE 237
L+ E +R E R R RQE + + + E+ ++LREE E +E
Sbjct: 1013 --LQQEEEQLLREE------REKRRRQEWERQYRKKD--ELQQEEEQLLREEREKRRLQE 1062
Query: 238 RMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSE 294
R R +E L++ E + L +E+E +++ EL +Y E+ + ++ E
Sbjct: 1063 RERQYREEEELQQEEEQL-----LGEERE------TRRRQELERQYRKEEELQQEEE 1108
Score = 40.0 bits (92), Expect = 0.007
Identities = 60/287 (20%), Positives = 125/287 (42%), Gaps = 36/287 (12%)
Query: 21 EEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQNEGLGEHRL--RDLNDEI 78
++++ +++R F + + E + R+Q +E R++AE + + + RL RD
Sbjct: 111 QDRRTEEDQRRFEPRDRQLEEEPGQRRRQKRQEQERELAEGEEQSEKQERLEQRDRQRRD 170
Query: 79 NKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSGRGPGYRYFGAAKKLPG 138
+L R++ W+ R A+ L S +G F ++L
Sbjct: 171 EELWRQRQEWQER---------EERRAEEEQLQ---------SCKGHETEEFPDEEQLRR 212
Query: 139 VRELFD-----KPPELRKRRTRYD-IYKRIDAAYYGYRDD----EDGILERLEPSAEAAM 188
REL + + + ++RR R D +++ + + R+ E+ L+ EP +
Sbjct: 213 -RELLELRRKGREEKQQQRRERQDRVFQEEEEKEWRKRETVLRKEEEKLQEEEPQRQ--- 268
Query: 189 RREAAEEWYRLDRI-RQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKER 247
RE EE +L ++ RQE R+ + E + + LR ++E+ E++ RE +ER
Sbjct: 269 -RELQEEEEQLRKLERQELRRERQEEEQQQQRLRREQQLRRKQEEERREQQEERREQQER 327
Query: 248 LEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSE 294
E+ E + E+ E+ + +++ E + + + +E+ E
Sbjct: 328 REQQEERREQQLRREQEERREQQLRREQEEERREQQLRREQEEERRE 374
Score = 38.5 bits (88), Expect = 0.021
Identities = 39/157 (24%), Positives = 73/157 (45%), Gaps = 5/157 (3%)
Query: 141 ELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRLD 200
EL+ + E ++R R +++ + G+ +E E+L +RR+ EE +
Sbjct: 172 ELWRQRQEWQEREERRAEEEQLQSCK-GHETEEFPDEEQLRRRELLELRRKGREEKQQQR 230
Query: 201 RIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKERLEENEREFVVHVP 260
R RQ+ V E + +LR+EEE + EEE R RE++E EE R+
Sbjct: 231 RERQDR---VFQEEEEKEWRKRETVLRKEEEKLQEEEPQRQRELQEE-EEQLRKLERQEL 286
Query: 261 LPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAKD 297
+ +E E+ ++ + L + E+ ++Q E ++
Sbjct: 287 RRERQEEEQQQQRLRREQQLRRKQEEERREQQEERRE 323
Score = 35.4 bits (80), Expect = 0.18
Identities = 37/147 (25%), Positives = 66/147 (44%), Gaps = 33/147 (22%)
Query: 169 YRDDEDGILERLEPSA------------EAAMRREAAEEWYRLDRIRQ--EARKGVKSGE 214
+R+DE + ER E E +RR+ E+ R DR R+ E + ++ GE
Sbjct: 1576 FREDEQLLQEREEQQLHRQERDRKFLEEEPQLRRQEREQQLRHDRDRKFREEEQLLQEGE 1635
Query: 215 VAEVSAAVREILREEEEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEK 274
+ LR +E D + R E + R +E ER+F L +E+++ + LE+
Sbjct: 1636 EQQ--------LRRQERD----RKFREEEQQLRRQERERKF-----LQEEQQLRRQELER 1678
Query: 275 K--KMELLNKYVSEDLMDEQSEAKDML 299
K + E L + ++ + Q + +L
Sbjct: 1679 KFREEEQLRQETEQEQLRRQERYRKIL 1705
Score = 35.4 bits (80), Expect = 0.18
Identities = 29/122 (23%), Positives = 59/122 (47%), Gaps = 13/122 (10%)
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEE 237
ER + + E E W +L+ +E R+ + E +++ RE+EE E+
Sbjct: 489 ERRKQQLKRDQEEERRERWLKLEE--EERREQQERRE--------QQLRREQEER--REQ 536
Query: 238 RMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAKD 297
R++ +E +ERL++ R + E+ +E+++ +++ L + + L EQ E +D
Sbjct: 537 RLKRQEEEERLQQRLRS-EQQLRREQEERLEQLLKREEEKRLEQERREQRLKREQEERRD 595
Query: 298 ML 299
L
Sbjct: 596 QL 597
Score = 33.5 bits (75), Expect = 0.67
Identities = 34/135 (25%), Positives = 61/135 (45%), Gaps = 8/135 (5%)
Query: 149 LRKRRTRYDIYKRIDAAYYGYRDDED--GILERLEPSAEAAMRREAAEEWYRLDRIRQEA 206
LR+ R + +R+ YR++E+ E+L RR+ E YR + Q+
Sbjct: 1051 LREEREK----RRLQERERQYREEEELQQEEEQLLGEERETRRRQELERQYRKEEELQQE 1106
Query: 207 RKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKE--RLEENEREFVVHVPLPDE 264
+ + E + RE EEE++ +EE RE +E R +E ER++ L +
Sbjct: 1107 EEQLLREEPEKRRRQERERQCREEEELQQEEEQLLREEREKRRRQELERQYREEEELQRQ 1166
Query: 265 KEIEKMVLEKKKMEL 279
K ++ E ++ +L
Sbjct: 1167 KRKQRYRDEDQRSDL 1181
Score = 33.1 bits (74), Expect = 0.87
Identities = 57/294 (19%), Positives = 118/294 (39%), Gaps = 58/294 (19%)
Query: 3 RNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQ 62
R EE+ Q + + + E++++ + R +CR+ E + +Q++RE
Sbjct: 1098 RKEEELQQEEEQLLREEPEKRRRQERER-----QCREEEELQQEEEQLLRE--------- 1143
Query: 63 NEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSG 122
R + E+ + RE+ +R+ + R + +DL P
Sbjct: 1144 ------EREKRRRQELERQYREEEELQRQKRK----QRYRDEDQRSDLKWQWE--PEKEN 1191
Query: 123 RGPGYRYFGAAKKLPGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEP 182
+ + ++ R+L D ++R R+++ D+ + G + + D ER
Sbjct: 1192 AVRDNKVYCKGRENEQFRQLEDS--QVRDRQSQQDLQHLL-----GEQQERDREQERRR- 1243
Query: 183 SAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGR 242
+ A R EE + ++ R+ KS E +++LREE R
Sbjct: 1244 -WQQANRHFPEEEQLEREEQKEAKRRDRKSQEE-------KQLLREE------------R 1283
Query: 243 EMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAK 296
E K R +E +R+F L E+E + ++ +++ K+ E+L+ ++ K
Sbjct: 1284 EEKRRRQETDRKFREEEQLLQEREEQPLLRQERD----RKFREEELLHQEQGRK 1333
Score = 33.1 bits (74), Expect = 0.87
Identities = 37/135 (27%), Positives = 61/135 (44%), Gaps = 22/135 (16%)
Query: 170 RDDEDGILERLEPSAEAAMRREAAE---------EWYRLDRIRQEARKGVKSGEVAEVSA 220
+++E+ + +RL +E +RRE E E RL++ R+E R + E +
Sbjct: 541 QEEEERLQQRLR--SEQQLRREQEERLEQLLKREEEKRLEQERREQRLKREQEERRDQLL 598
Query: 221 AVREILREEEEDVVEEERMRG---REMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKM 277
E R++ +EER+ RE ERLE+ ER DE+ + E+++
Sbjct: 599 KREEERRQQRLKREQEERLEQRLKREEVERLEQEERR--------DERLKREEPEEERRH 650
Query: 278 ELLNKYVSEDLMDEQ 292
ELL E+ EQ
Sbjct: 651 ELLKSEEQEERRHEQ 665
Score = 32.3 bits (72), Expect = 1.5
Identities = 31/123 (25%), Positives = 60/123 (48%), Gaps = 13/123 (10%)
Query: 141 ELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRLD 200
+L + E R++R + + +R++ + +E LE+ E + ++RE EE R +
Sbjct: 596 QLLKREEERRQQRLKREQEERLEQRL---KREEVERLEQ-EERRDERLKREEPEEERRHE 651
Query: 201 RIRQEARKGVKSGEVAEVSAAVRE--ILREEEEDVVE-------EERMRGREMKERLEEN 251
++ E ++ + ++ RE + REEEE+ +E EE R +E+ E +E
Sbjct: 652 LLKSEEQEERRHEQLRREQQERREQRLKREEEEERLEQRLKREHEEERREQELAEEEQEQ 711
Query: 252 ERE 254
RE
Sbjct: 712 ARE 714
Score = 32.3 bits (72), Expect = 1.5
Identities = 34/155 (21%), Positives = 72/155 (45%), Gaps = 13/155 (8%)
Query: 149 LRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARK 208
L+++ + +R+ + R+ E+ + + L+ E + +E E+ RL R ++E R
Sbjct: 538 LKRQEEEERLQQRLRSEQQLRREQEERLEQLLKREEEKRLEQERREQ--RLKREQEERRD 595
Query: 209 GV-KSGEVAEVSAAVRE--------ILREEEEDVVEEERMRGREMKERLEENEREFVVHV 259
+ K E RE + REE E + +EER R +E EE R ++
Sbjct: 596 QLLKREEERRQQRLKREQEERLEQRLKREEVERLEQEERRDERLKREEPEEERRHELLKS 655
Query: 260 PLPDEKEIEKMVLE--KKKMELLNKYVSEDLMDEQ 292
+E+ E++ E +++ + L + E+ ++++
Sbjct: 656 EEQEERRHEQLRREQQERREQRLKREEEEERLEQR 690
Score = 32.0 bits (71), Expect = 1.9
Identities = 56/301 (18%), Positives = 124/301 (40%), Gaps = 27/301 (8%)
Query: 4 NEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWR------QQIMR-EIGR 56
++E+ + L ++ E ++K + L E R+ D+ R QQ+ R E R
Sbjct: 1327 HQEQGRKFLEEEQRLREERERKFLKEEQQLRLEEREQLRQDRDRKFREEEQQLSRQERDR 1386
Query: 57 KVAEIQNEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVD 116
K E + + + R R +E +L R++ H + R E + D ++
Sbjct: 1387 KFREEEQQVRRQERERKFLEEEQQL-RQERHRKFREEEQLLQEREEQQLHRQERDRKFLE 1445
Query: 117 VPNPSGRGPGYRYFGAAKKLPGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGI 176
R R F +EL + PE + ++++ + + +
Sbjct: 1446 EEQQLRRQERDRKFRE-------QELRSQEPERKFLEEEQQLHRQQRQRKFLQEEQQLRR 1498
Query: 177 LERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEE 236
ER + + R+ EE R +R Q+ + + + VR +E+E +E+
Sbjct: 1499 QERGQQRRQDRDRKFREEEQLRQEREEQQLSRQERDRKFRLEEQKVRR--QEQERKFMED 1556
Query: 237 ER-MRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEA 295
E+ +R +E +++L + +R+F +E E+++ E+++ +L + ++E+ +
Sbjct: 1557 EQQLRRQEGQQQLRQEDRKF---------REDEQLLQEREEQQLHRQERDRKFLEEEPQL 1607
Query: 296 K 296
+
Sbjct: 1608 R 1608
Score = 30.8 bits (68), Expect = 4.3
Identities = 35/174 (20%), Positives = 77/174 (44%), Gaps = 19/174 (10%)
Query: 140 RELFDKPPELRKR----RTRYDIYKRIDAAYYGYRDDEDGILERLEPSA-----EAAMRR 190
R+ ++ P+LR++ + R+D ++ ++ E+ L R E E +RR
Sbjct: 1598 RKFLEEEPQLRRQEREQQLRHDRDRKFREEEQLLQEGEEQQLRRQERDRKFREEEQQLRR 1657
Query: 191 EAAEEWYRLDRI---RQEARKGVKSGEVAEVSAAVREILREEE-EDVVEEERMR--GREM 244
+ E + + RQE + + E ++ R+E ++EEE++R E
Sbjct: 1658 QERERKFLQEEQQLRRQELERKFREEEQLRQETEQEQLRRQERYRKILEEEQLRPEREEQ 1717
Query: 245 KERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAKDM 298
+ R +E +R+F L +E +++ + E K+ E+ + ++ E + +
Sbjct: 1718 QLRRQERDRKFREEEQLRQGREEQQL----RSQESDRKFREEEQLRQEREEQQL 1767
Score = 30.0 bits (66), Expect = 7.4
Identities = 27/118 (22%), Positives = 52/118 (43%), Gaps = 11/118 (9%)
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEE 237
E + ++ + R+ EE R +R Q+ R + G+ EEE+ +EE+
Sbjct: 1739 EEQQLRSQESDRKFREEEQLRQEREEQQLRPQQRDGKYRW----------EEEQLQLEEQ 1788
Query: 238 RMRGREMKERLEENEREFVV-HVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSE 294
R R+ ++R E +F +E+E+ + +K++ E K E + +Q E
Sbjct: 1789 EQRLRQERDRQYRAEEQFATQEKSRREEQELWQEEEQKRRQERERKLREEHIRRQQKE 1846
Score = 29.6 bits (65), Expect = 9.6
Identities = 28/108 (25%), Positives = 47/108 (42%), Gaps = 5/108 (4%)
Query: 143 FDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREA---AEEWYRL 199
F + +LR+ R + R + +R++E ER E R EE +L
Sbjct: 1728 FREEEQLRQGREEQQL--RSQESDRKFREEEQLRQEREEQQLRPQQRDGKYRWEEEQLQL 1785
Query: 200 DRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKER 247
+ Q R+ AE A +E R EE+++ +EE + R+ +ER
Sbjct: 1786 EEQEQRLRQERDRQYRAEEQFATQEKSRREEQELWQEEEQKRRQERER 1833
Score = 29.6 bits (65), Expect = 9.6
Identities = 27/136 (19%), Positives = 61/136 (44%), Gaps = 9/136 (6%)
Query: 140 RELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRL 199
REL ++ +LRK R ++ + R +E+ +RL + ++E +
Sbjct: 269 RELQEEEEQLRKLE-RQELRRE--------RQEEEQQQQRLRREQQLRRKQEEERREQQE 319
Query: 200 DRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKERLEENEREFVVHV 259
+R Q+ R+ + + +E RE++ +EE R ++++ EE RE +
Sbjct: 320 ERREQQERREQQEERREQQLRREQEERREQQLRREQEEERREQQLRREQEEERREQQLRR 379
Query: 260 PLPDEKEIEKMVLEKK 275
+E+ +++ E++
Sbjct: 380 EQEEERREQQLRREQQ 395
>U2R1_MOUSE (Q64707) U2 small nuclear ribonucleoprotein auxiliary
factor 35 kDa subunit related-protein 1 (U2(RNU2) small
nuclear RNA auxillary factor 1-like 1) (SP2)
Length = 428
Score = 41.6 bits (96), Expect = 0.002
Identities = 26/89 (29%), Positives = 47/89 (52%), Gaps = 3/89 (3%)
Query: 177 LERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVRE---ILREEEEDV 233
+ER E +R E A+E +R+ + ++EA + K + ++ A E REEEE
Sbjct: 62 IERQRLHEEWLLREEKAQEEFRIKKKKEEAARKQKEEQERQIKAEWEEQQKKQREEEEQK 121
Query: 234 VEEERMRGREMKERLEENEREFVVHVPLP 262
++E+R R +++ L++ E E + P P
Sbjct: 122 LQEKREREEAVQKMLDQAENERIWQNPEP 150
>TRHY_RABIT (P37709) Trichohyalin
Length = 1407
Score = 41.6 bits (96), Expect = 0.002
Identities = 44/169 (26%), Positives = 86/169 (50%), Gaps = 21/169 (12%)
Query: 140 RELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILE----RLEPSAEAAMRREAAEE 195
REL ++ + R+RR +++ +++E+ +L R EP + +RRE E
Sbjct: 216 RELREEEQQRRERREQHE---------RALQEEEEQLLRQRRWREEPREQQQLRRELEEI 266
Query: 196 WYRLDRIRQEARK--GVKSGEVAEVSAAVREILREEEEDVVE-EERMRGREMKE-RLEEN 251
R R+ QE R+ ++ + E + LR E E++ E E+R+ E +E RLE+
Sbjct: 267 REREQRLEQEERREQQLRREQRLEQEERREQQLRRELEEIREREQRLEQEERREQRLEQE 326
Query: 252 E-REFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAKDML 299
E RE + L + +E E+ + ++++ E L ++E++ ++ E + L
Sbjct: 327 ERREQQLKRELEEIREREQRLEQEERREQL---LAEEVREQARERGESL 372
Score = 38.9 bits (89), Expect = 0.016
Identities = 71/296 (23%), Positives = 130/296 (42%), Gaps = 35/296 (11%)
Query: 3 RNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQ 62
R EEK Q + E++K+ ++ R E +L + ++ R + +RE + + E +
Sbjct: 567 REEEKLQ---------REEDEKRRRQERERQYRELEELRQEEQLRDRKLREEEQLLQERE 617
Query: 63 NEGLG-EHRLRDLNDEINKLIREKSHW----ERRIVELGGPNYTRHSAKMTDLDGNIVDV 117
E L + R R L +E L +E+ ER++ E + + + +
Sbjct: 618 EERLRRQERERKLREEEQLLRQEEQELRQERERKLREEEQLLRREEQELRQERERKLREE 677
Query: 118 PN--PSGRGPGYRYFGAAKKLPGVRELF-DKPPELRKRRTRY-----DIYKRIDAAYYGY 169
R A+KL +L + ELR+ R R + +R +
Sbjct: 678 EQLLQEREEERLRRQERARKLREEEQLLRQEEQELRQERERKLREEEQLLRREEQLLRQE 737
Query: 170 RDDEDGILERL-EPSAEAAMRREAAEEWYRLDRIRQ--EARKGVKSGEVAEVSAAVREI- 225
RD + E+L + S E +RR+ E+ R +R R+ E + ++ E + RE
Sbjct: 738 RDRKLREEEQLLQESEEERLRRQEREQQLRRERDRKFREEEQLLQEREEERLRRQERERK 797
Query: 226 LREEEEDVV--EEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMEL 279
LREEE+ + EEER+R +E + +L E E+ L E+E E++ ++++ +L
Sbjct: 798 LREEEQLLQEREEERLRRQERERKLREEEQ-------LLQEREEERLRRQERERKL 846
Score = 38.5 bits (88), Expect = 0.021
Identities = 43/185 (23%), Positives = 78/185 (41%), Gaps = 43/185 (23%)
Query: 137 PGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEW 196
PG +E + EL++ + R + + YR++E +L+ + RR+ E
Sbjct: 541 PGQQEQLREEEELQREKRRQERERE-------YREEE-----KLQREEDEKRRRQERERQ 588
Query: 197 YR-LDRIRQEA----------------------RKGVKSGEVAEVSAAVR----EILREE 229
YR L+ +RQE R+ + ++ E +R E+ +E
Sbjct: 589 YRELEELRQEEQLRDRKLREEEQLLQEREEERLRRQERERKLREEEQLLRQEEQELRQER 648
Query: 230 EEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLM 289
E + EEE++ RE +E +E ER+ L E+E E++ ++ E K E+ +
Sbjct: 649 ERKLREEEQLLRREEQELRQERERKLREEEQLLQEREEERL----RRQERARKLREEEQL 704
Query: 290 DEQSE 294
Q E
Sbjct: 705 LRQEE 709
Score = 37.0 bits (84), Expect = 0.060
Identities = 31/129 (24%), Positives = 61/129 (47%), Gaps = 6/129 (4%)
Query: 170 RDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVR----EI 225
R++E + ER E R E +L R ++ + ++ ++ E +R E+
Sbjct: 1171 REEEQLLQEREEERLRRQERARKLREEEQLLRQEEQELRQERARKLREEEQLLRQEEQEL 1230
Query: 226 LREEEEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVS 285
+E + EEE++ RE +E E +R+F L E+E E++ +++ +L +
Sbjct: 1231 RQERDRKFREEEQLLRREEQELRRERDRKFREEEQLLQEREEERLRRQERARKLREE--E 1288
Query: 286 EDLMDEQSE 294
E L+ E+ E
Sbjct: 1289 EQLLFEEQE 1297
Score = 37.0 bits (84), Expect = 0.060
Identities = 65/298 (21%), Positives = 118/298 (38%), Gaps = 39/298 (13%)
Query: 21 EEKKKPKERRPFLATECRDLSEADKWRQQIMREIG-----RKVAEIQNEGLGEHRLRDLN 75
E +++ +E+RP + EA + R + + G R+ E+Q E + R R+
Sbjct: 508 ERRRRQQEQRPGQTWRWQLQEEAQRRRHTLYAKPGQQEQLREEEELQREKRRQEREREYR 567
Query: 76 DEINKLIRE------KSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSGRGPGYRY 129
+E KL RE + ER+ EL R ++ D + R
Sbjct: 568 EE-EKLQREEDEKRRRQERERQYREL---EELRQEEQLRDRKLREEEQLLQEREEERLRR 623
Query: 130 FGAAKKLPGVRELF-DKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAM 188
+KL +L + ELR+ R R +E+ +L R E
Sbjct: 624 QERERKLREEEQLLRQEEQELRQERERK-------------LREEEQLLRREEQELRQER 670
Query: 189 RREAAEEWYRL-----DRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGRE 243
R+ EE L +R+R++ R E + +E+ +E E + EEE++ RE
Sbjct: 671 ERKLREEEQLLQEREEERLRRQERARKLREEEQLLRQEEQELRQERERKLREEEQLLRRE 730
Query: 244 MKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNK-----YVSEDLMDEQSEAK 296
+ +E +R+ L E E E++ ++++ +L + E L+ E+ E +
Sbjct: 731 EQLLRQERDRKLREEEQLLQESEEERLRRQEREQQLRRERDRKFREEEQLLQEREEER 788
Score = 36.2 bits (82), Expect = 0.10
Identities = 66/307 (21%), Positives = 127/307 (40%), Gaps = 37/307 (12%)
Query: 3 RNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKW----RQQIMREIGRKV 58
R +E+A+ + ++ EE++ +ER L E + L ++ R + +RE + +
Sbjct: 690 RRQERARKLREEEQLLRQEEQELRQERERKLREEEQLLRREEQLLRQERDRKLREEEQLL 749
Query: 59 AEIQNEGLG-EHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDV 117
E + E L + R + L E ++ RE+ E+ + E R + + +
Sbjct: 750 QESEEERLRRQEREQQLRRERDRKFREE---EQLLQEREEERLRRQERERKLREEEQLLQ 806
Query: 118 PNPSGRGPGYRYFGAAKKLPGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGIL 177
R R +KL +L + E R RR + R +E+ +L
Sbjct: 807 EREEER---LRRQERERKLREEEQLLQEREEERLRRQERERKLR----------EEEQLL 853
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVRE---ILREEEEDVV 234
+ E R+ EE L + QE R+ E +RE +LR+EE+++
Sbjct: 854 RQEEQELRQERARKLREEEQLLRQEEQELRQ--------ERDRKLREEEQLLRQEEQELR 905
Query: 235 EEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYV-----SEDLM 289
+E + RE ++ L+E+E E + + E+ +L +++ EL + E L+
Sbjct: 906 QERDRKLREEEQLLQESEEERLRRQERERKLREEEQLLRREEQELRRERARKLREEEQLL 965
Query: 290 DEQSEAK 296
E+ E +
Sbjct: 966 QEREEER 972
Score = 34.7 bits (78), Expect = 0.30
Identities = 36/139 (25%), Positives = 61/139 (42%), Gaps = 24/139 (17%)
Query: 180 LEPSAEAAMRREAAEEWYR-----LDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVV 234
L+ S E +RR+ E R L R QE R+ E A ++L+E EE+ +
Sbjct: 919 LQESEEERLRRQERERKLREEEQLLRREEQELRR-----ERARKLREEEQLLQEREEERL 973
Query: 235 ----------EEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYV 284
EEE++ RE +E +E +R+F L E+E E++ ++ E K+
Sbjct: 974 RRQERARKLREEEQLLRREEQELRQERDRKFREEEQLLQEREEERL----RRQERDRKFR 1029
Query: 285 SEDLMDEQSEAKDMLNIHR 303
E+ + E ++ R
Sbjct: 1030 EEERQLRRQELEEQFRQER 1048
Score = 33.5 bits (75), Expect = 0.67
Identities = 66/335 (19%), Positives = 125/335 (36%), Gaps = 46/335 (13%)
Query: 3 RNEEKAQSMLNRFIT--MKAEEKKKPKERRPFLATECRDLSEADKWRQQIMRE---IGRK 57
R E + Q L R + + E++ + +ERR + L + ++ QQ+ RE I +
Sbjct: 250 REEPREQQQLRRELEEIREREQRLEQEERREQQLRREQRLEQEERREQQLRRELEEIRER 309
Query: 58 VAEIQNEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDV 117
++ E E RL +L RE R L + + L + +
Sbjct: 310 EQRLEQEERREQRLEQEERREQQLKRELEEIREREQRL-----EQEERREQLLAEEVREQ 364
Query: 118 PNPSGRGPGYRYFGAAKKLPGVRE--LFDKP-----------PELRKRRTRYDIYKRIDA 164
G R+ + G R+ ++ +P E R+R+ R +
Sbjct: 365 ARERGESLTRRWQRQLESEAGARQSKVYSRPRRQEEQSLRQDQERRQRQERERELEEQAR 424
Query: 165 AYYGYRDDEDGILERLEPSAEAAMRRE---AAEEWYRLDRIRQEARKGVKSGEVAEVSAA 221
++ +E+ R SA ++R A E + R R+E + + + +
Sbjct: 425 RQQQWQAEEESERRRQRLSARPSLRERQLRAEERQEQEQRFREEEEQRRERRQELQFLEE 484
Query: 222 VREILREE------EEDVVEEERMRGREMKER----------LEENEREFVVHVPLPDE- 264
++ R E EED +E+R R R +E+ EE +R P +
Sbjct: 485 EEQLQRRERAQQLQEEDSFQEDRERRRRQQEQRPGQTWRWQLQEEAQRRRHTLYAKPGQQ 544
Query: 265 ---KEIEKMVLEKKKMELLNKYVSEDLMDEQSEAK 296
+E E++ EK++ E +Y E+ + + + K
Sbjct: 545 EQLREEEELQREKRRQEREREYREEEKLQREEDEK 579
Score = 33.1 bits (74), Expect = 0.87
Identities = 68/294 (23%), Positives = 122/294 (41%), Gaps = 50/294 (17%)
Query: 5 EEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQN- 63
+E+A+ + ++ EE++ +ER L E + L + + Q++ +E RK+ E +
Sbjct: 862 QERARKLREEEQLLRQEEQELRQERDRKLREEEQLLRQEE---QELRQERDRKLREEEQL 918
Query: 64 -EGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSG 122
+ E RLR E + +RE+ RR + R A+ + ++
Sbjct: 919 LQESEEERLR--RQERERKLREEEQLLRREEQ----ELRRERARKLREEEQLLQEREEER 972
Query: 123 RGPGYRYFGAAKKLPGVRELFDKPP-ELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLE 181
R A+KL +L + ELR+ R R +R++E + ER E
Sbjct: 973 ----LRRQERARKLREEEQLLRREEQELRQERDRK------------FREEEQLLQEREE 1016
Query: 182 P------------SAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREE 229
E +RR+ EE +R +R R+ + E E LR +
Sbjct: 1017 ERLRRQERDRKFREEERQLRRQELEEQFRQERDRKFRLEEQIRQEKEEKQ------LRRQ 1070
Query: 230 EED--VVEEERMRGREMKER--LEENEREFVVHVPLPDEKEIEKMVLEKKKMEL 279
E D EEE+ R R+ +E+ E +R+F L E+E E++ +++ +L
Sbjct: 1071 ERDRKFREEEQQRRRQEREQQLRRERDRKFREEEQLLQEREEERLRRQERARKL 1124
Score = 31.6 bits (70), Expect = 2.5
Identities = 62/294 (21%), Positives = 120/294 (40%), Gaps = 37/294 (12%)
Query: 3 RNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQ 62
R +E+ + + ++ EE++ +ER L E + L E ++ R + +E RK+ E
Sbjct: 928 RRQERERKLREEEQLLRREEQELRRERARKLREEEQLLQEREEERLR-RQERARKLRE-- 984
Query: 63 NEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSG 122
E L ++L E ++ RE+ E+ + E R
Sbjct: 985 EEQLLRREEQELRQERDRKFREE---EQLLQEREEERLRRQERD---------------- 1025
Query: 123 RGPGYRYFGAAKKLPGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEP 182
R F ++ +EL ++ + R R+ R + R + R E +R
Sbjct: 1026 -----RKFREEERQLRRQELEEQFRQERDRKFRLEEQIRQEKEEKQLRRQER---DRKFR 1077
Query: 183 SAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGR 242
E RR+ E+ R R+ RK + ++ + R +E + EEE++ R
Sbjct: 1078 EEEQQRRRQEREQQLR----RERDRKFREEEQLLQEREEERLRRQERARKLREEEQLLRR 1133
Query: 243 EMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAK 296
E + +E +R+F L E E E++ ++++ +L + E L+ E+ E +
Sbjct: 1134 EEQLLRQERDRKFREEEQLLQESEEERLRRQERERKLREE---EQLLQEREEER 1184
Score = 30.8 bits (68), Expect = 4.3
Identities = 69/321 (21%), Positives = 121/321 (37%), Gaps = 39/321 (12%)
Query: 3 RNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRK----- 57
R E + L I + EEK+ ++ R E + QQ+ RE RK
Sbjct: 1045 RQERDRKFRLEEQIRQEKEEKQLRRQERDRKFREEEQQRRRQEREQQLRRERDRKFREEE 1104
Query: 58 --VAEIQNEGLG-EHRLRDLNDEINKLIRE----------KSHWERRIVELGGPNYTRHS 104
+ E + E L + R R L +E L RE K E ++++ R
Sbjct: 1105 QLLQEREEERLRRQERARKLREEEQLLRREEQLLRQERDRKFREEEQLLQESEEERLRRQ 1164
Query: 105 AKMTDLDGNIVDVPNPSGRGPGYRYFGAAKKLPGVRELF-DKPPELRKRRTRY-----DI 158
+ L + R A+KL +L + ELR+ R R +
Sbjct: 1165 ERERKLREE--EQLLQEREEERLRRQERARKLREEEQLLRQEEQELRQERARKLREEEQL 1222
Query: 159 YKRIDAAYYGYRD----DEDGILERLEPSAEAAMRREAAEEWYRL-DRIRQEARKGVKSG 213
++ + RD +E+ +L R E R+ EE L +R + R+ ++
Sbjct: 1223 LRQEEQELRQERDRKFREEEQLLRREEQELRRERDRKFREEEQLLQEREEERLRRQERAR 1282
Query: 214 EVAEVSAAVREILREEEEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLE 273
++ E + + E+EE + +ER R +E+ E+ + L E+E +
Sbjct: 1283 KLREEEEQL--LFEEQEEQRLRQERDRRYRAEEQFAREEKSRRLERELRQEEE------Q 1334
Query: 274 KKKMELLNKYVSEDLMDEQSE 294
+++ E K+ E L +Q E
Sbjct: 1335 RRRRERERKFREEQLRRQQEE 1355
Score = 30.4 bits (67), Expect = 5.6
Identities = 34/153 (22%), Positives = 64/153 (41%), Gaps = 6/153 (3%)
Query: 150 RKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKG 209
RK R + ++ + RD + E+L E +R+E + +++ QE+ +
Sbjct: 866 RKLREEEQLLRQEEQELRQERDRKLREEEQLLRQEEQELRQERDRKLREEEQLLQESEEE 925
Query: 210 VKSGEVAEVSAAVREILREEEEDVVEEERMRG-REMKERLEENEREFVVHVPLPDEKEIE 268
+ E E L EE + ER R RE ++ L+E E E + + E
Sbjct: 926 RLRRQERERKLREEEQLLRREEQELRRERARKLREEEQLLQEREEERLRRQERARKLREE 985
Query: 269 KMVLEKKKMELLNK-----YVSEDLMDEQSEAK 296
+ +L +++ EL + E L+ E+ E +
Sbjct: 986 EQLLRREEQELRQERDRKFREEEQLLQEREEER 1018
Score = 30.4 bits (67), Expect = 5.6
Identities = 29/114 (25%), Positives = 52/114 (45%), Gaps = 23/114 (20%)
Query: 140 RELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRL 199
REL ++ + +KR Y R YRD E +RL+ R+E E
Sbjct: 144 RELAEEEEQRKKRERFEQHYSR------QYRDKE----QRLQ-------RQELEERRAEE 186
Query: 200 DRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKERLEENER 253
+++R+ + G AE ++ R E++++ E R ++ +ER E++ER
Sbjct: 187 EQLRR------RKGRDAEEFIEEEQLRRREQQELKRELREEEQQRRERREQHER 234
>EZRI_MOUSE (P26040) Ezrin (p81) (Cytovillin) (Villin 2)
Length = 585
Score = 40.8 bits (94), Expect = 0.004
Identities = 37/121 (30%), Positives = 59/121 (48%), Gaps = 16/121 (13%)
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSA--AVREILREEEEDVVE 235
ERLE AA+R + E D+I+ + + + E+AE +A A+ E R +ED VE
Sbjct: 386 ERLEADRMAALRAKEELERQAQDQIKSQEQ---LAAELAEYTAKIALLEEARRRKEDEVE 442
Query: 236 EERMRGREMKERLEENERE--FVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQS 293
E + R +E ++ L + + E V+ P P + E +N +V E L DE +
Sbjct: 443 EWQHRAKEAQDDLVKTKEELHLVMTAPPPPPPPV---------YEPVNYHVQEGLQDEGA 493
Query: 294 E 294
E
Sbjct: 494 E 494
>VP27_YEAST (P40343) Vacuolar protein sorting-associated protein
VPS27
Length = 622
Score = 40.4 bits (93), Expect = 0.005
Identities = 34/150 (22%), Positives = 74/150 (48%), Gaps = 6/150 (4%)
Query: 151 KRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEA-RKG 209
K+ ++ ++ D Y D+E+ I + +E S + + ++E + + E R+
Sbjct: 238 KKSKKHRHKRKKDRDYSTPEDEEELIRKAIELSLKESRNSASSEPIVPVVESKNEVKRQE 297
Query: 210 VKSGEVAEVSAAVREILREEEEDVVEEERMR-GREMKERLEENEREFVVHVPLPDEKE-- 266
++ E ++ AA++E LRE EE V ER + R+M+ + + + + V L DE++
Sbjct: 298 IEEEEDPDLKAAIQESLREAEEANVRSERQKASRQMQPQQPSPQPQPIHSVDLSDEEKDS 357
Query: 267 --IEKMVLEKKKMELLNKYVSEDLMDEQSE 294
+ ++EK K LN+ + + + ++
Sbjct: 358 IYMFASLVEKMKSRPLNEILEDSKLQNLAQ 387
>EZRI_HUMAN (P15311) Ezrin (p81) (Cytovillin) (Villin 2)
Length = 585
Score = 40.4 bits (93), Expect = 0.005
Identities = 36/121 (29%), Positives = 60/121 (48%), Gaps = 16/121 (13%)
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSA--AVREILREEEEDVVE 235
ERLE AA+R + E +D+I+ + + + E+AE +A A+ E R +ED VE
Sbjct: 386 ERLEADRMAALRAKEELERQAVDQIKSQEQ---LAAELAEYTAKIALLEEARRRKEDEVE 442
Query: 236 EERMRGREMKERLEENERE--FVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQS 293
E + R +E ++ L + + E V+ P P + E ++ +V E L DE +
Sbjct: 443 EWQHRAKEAQDDLVKTKEELHLVMTAPPPPPPPV---------YEPVSYHVQESLQDEGA 493
Query: 294 E 294
E
Sbjct: 494 E 494
Score = 30.0 bits (66), Expect = 7.4
Identities = 31/132 (23%), Positives = 57/132 (42%), Gaps = 7/132 (5%)
Query: 148 ELRKRRTRYDIYKRIDAAYYGYRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEAR 207
EL RR + D + + LER + E R E ++ R ++E
Sbjct: 288 ELYMRRRKPDTIEVQQMKAQAREEKHQKQLERQQLETEKKRRETVEREKEQMMREKEELM 347
Query: 208 KGVKSGEVAEVSAAVREILREEEEDV-VEEERMRGREMKERLEENEREFVVHVPLPDEKE 266
++ E + A RE+ + + + +EEER R +E ERLE + L ++E
Sbjct: 348 LRLQDYE-EKTKKAERELSEQIQRALQLEEERKRAQEEAERLEADRM-----AALRAKEE 401
Query: 267 IEKMVLEKKKME 278
+E+ +++ K +
Sbjct: 402 LERQAVDQIKSQ 413
>YAFA_SCHPO (Q09863) Hypothetical protein C29E6.10c in chromosome I
Length = 1085
Score = 40.0 bits (92), Expect = 0.007
Identities = 34/156 (21%), Positives = 81/156 (51%), Gaps = 19/156 (12%)
Query: 148 ELRKRRTRYDIYKRIDAAYYGYRD--DEDGILERLEPSAEAAMRREAAEEWYRLDRIRQE 205
++R++ + D K++ A R + + + E+ A A R+E A + R+++E
Sbjct: 590 KIREKEKKRDKKKQLKLAKEEERQRREAERLAEQAAQKALEAKRQEEARKKREEQRLKRE 649
Query: 206 ARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKERLEENEREFVVHVPLPDEK 265
K +E+ R++ E+ ++++ R +++K++ +E +RE + E+
Sbjct: 650 QEK------------KQQELERQKREEK-QKQKEREKKLKKQQQEADREKMAREQRLREE 696
Query: 266 EIEKMVLEKKKMELLNKYVSE----DLMDEQSEAKD 297
E ++++ E+K+ E L+K E +L++++SE K+
Sbjct: 697 EEKRILEERKRREKLDKEEEERRRRELLEKESEEKE 732
Score = 34.3 bits (77), Expect = 0.39
Identities = 32/112 (28%), Positives = 52/112 (45%), Gaps = 11/112 (9%)
Query: 1 MARNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMRE------- 53
+ R +EK Q L R K EEK+K KER L + ++ R+Q +RE
Sbjct: 646 LKREQEKKQQELER---QKREEKQKQKEREKKLKKQQQEADREKMAREQRLREEEEKRIL 702
Query: 54 IGRKVAEIQNEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSA 105
RK E ++ E R R+L ++ ++ +E+ E +I PN T+ +
Sbjct: 703 EERKRREKLDKEEEERRRRELLEKESE-EKERRLREAKIAAFFAPNQTKEGS 753
Score = 30.8 bits (68), Expect = 4.3
Identities = 29/131 (22%), Positives = 57/131 (43%), Gaps = 19/131 (14%)
Query: 170 RDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREE 229
++D +E +E AE M+RE ++ E+ E E E+
Sbjct: 474 KNDGKKFIEMMEQLAERRMQREDNSNFHE--------------PELYESGLEYDEDEEED 519
Query: 230 EEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEI----EKMVLEKKKMELLNKYVS 285
EED V+E+ + ++R+EE R F + E+ + + V ++++ +LL +
Sbjct: 520 EED-VDEDELDLMTDEQRMEEGRRMFQIFAARLFEQRVLQAYREKVAQQRQAKLLEEIEE 578
Query: 286 EDLMDEQSEAK 296
E+ ++ E K
Sbjct: 579 ENKRKQERELK 589
>IF2_AQUAE (O67825) Translation initiation factor IF-2
Length = 805
Score = 40.0 bits (92), Expect = 0.007
Identities = 27/93 (29%), Positives = 49/93 (52%), Gaps = 8/93 (8%)
Query: 176 ILERLEPSAEAA-MRREAAEEWYRLDRIRQEARKGVKSGEVAE-------VSAAVREILR 227
ILE+ E E + +E EE R+ +++E RK K E E +S REI+R
Sbjct: 135 ILEKKEKEKEKKKVEKERKEEKVRVVEVKKEERKEEKKEEKKEEEKPKIKMSKKEREIMR 194
Query: 228 EEEEDVVEEERMRGREMKERLEENEREFVVHVP 260
+ E V +E++ + + KE+ ++ E ++++P
Sbjct: 195 KLEHAVEKEKKKQEKREKEKKKKEEEVKIIYIP 227
Score = 37.7 bits (86), Expect = 0.035
Identities = 31/114 (27%), Positives = 54/114 (47%), Gaps = 11/114 (9%)
Query: 176 ILERLEPSAEAAMRRE------AAEEWYR--LDRIRQEARKGVKSGEVAEVSAAVREILR 227
I+E +E E ++E + EE + L++ +E K E E V E+ +
Sbjct: 105 IVEEIEEKKEEEEKKEEEKPKKSVEELIKEILEKKEKEKEKKKVEKERKEEKVRVVEVKK 164
Query: 228 EEEEDVVEEERMRGREMKERLEENEREF---VVHVPLPDEKEIEKMVLEKKKME 278
EE ++ +EE+ + K ++ + ERE + H ++K+ EK EKKK E
Sbjct: 165 EERKEEKKEEKKEEEKPKIKMSKKEREIMRKLEHAVEKEKKKQEKREKEKKKKE 218
Score = 37.7 bits (86), Expect = 0.035
Identities = 26/105 (24%), Positives = 51/105 (47%), Gaps = 7/105 (6%)
Query: 188 MRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKER 247
++ E +E ++ + A K E + V E++ E++ +V+ EE +E +E+
Sbjct: 59 IKEEEEKEEVVTEQAQAPAEVEEKKEEEKKEEVIVEEVVEEKKPEVIVEEIEEKKEEEEK 118
Query: 248 LEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQ 292
EE + P +E+ K +LEKK+ E K V ++ +E+
Sbjct: 119 KEEEK-------PKKSVEELIKEILEKKEKEKEKKKVEKERKEEK 156
Score = 36.6 bits (83), Expect = 0.079
Identities = 26/78 (33%), Positives = 43/78 (54%), Gaps = 8/78 (10%)
Query: 228 EEEEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMV--------LEKKKMEL 279
EE+E+VV E+ E++E+ EE ++E V+ + +EK+ E +V E+KK E
Sbjct: 63 EEKEEVVTEQAQAPAEVEEKKEEEKKEEVIVEEVVEEKKPEVIVEEIEEKKEEEEKKEEE 122
Query: 280 LNKYVSEDLMDEQSEAKD 297
K E+L+ E E K+
Sbjct: 123 KPKKSVEELIKEILEKKE 140
Score = 31.6 bits (70), Expect = 2.5
Identities = 37/133 (27%), Positives = 60/133 (44%), Gaps = 17/133 (12%)
Query: 167 YGYRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREIL 226
+G +++E E+ E E A EE + +E ++ V EV E I+
Sbjct: 57 FGIKEEE----EKEEVVTEQAQAPAEVEE-----KKEEEKKEEVIVEEVVEEKKP-EVIV 106
Query: 227 REEEEDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYV-- 284
E EE EEE+ + K+ +EE +E + EKE EK +EK++ E + V
Sbjct: 107 EEIEEKKEEEEKKEEEKPKKSVEELIKEILE----KKEKEKEKKKVEKERKEEKVRVVEV 162
Query: 285 -SEDLMDEQSEAK 296
E+ +E+ E K
Sbjct: 163 KKEERKEEKKEEK 175
>CALD_RAT (Q62736) Non-muscle caldesmon (CDM) (L-caldesmon)
Length = 531
Score = 40.0 bits (92), Expect = 0.007
Identities = 66/301 (21%), Positives = 120/301 (38%), Gaps = 59/301 (19%)
Query: 2 ARNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEI 61
A EEK +S R+ + E ++ + E + E ++ ++ ++
Sbjct: 141 AEGEEKGESRSGRYEMEETEVVITSYQKNSYQDAEDKKKEEKEEEEEE---------EKL 191
Query: 62 QNEGLGEHRLRDLNDEINKLIRE--KSHWERR--IVELGGPN--YTRHSAKMTDLDGNIV 115
+ LGE++++D + +K +E K+ +R+ E+ N + H K T+ N
Sbjct: 192 KGGNLGENQIKDEKIKKDKEPKEEVKNFLDRKKGFTEVKAQNGEFMTHKLKQTE---NAF 248
Query: 116 DVPNPSGRGPGYRYFGAAKKLPGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDG 175
GR G + A ++ + L ELR+RR G + E+
Sbjct: 249 SPSRSGGRASGDKEAEGAPQVEAGKRL----EELRRRR--------------GETESEE- 289
Query: 176 ILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEE---- 231
E+L+ ++EAA E L + R+E RK ++ E REEEE
Sbjct: 290 -FEKLKQK-----QQEAALELEELKKKREERRKVLEEEEQRRKQEEADRKAREEEEKRRL 343
Query: 232 -DVVEEERMRGREMKER-----LEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVS 285
+ +E R E +++ L E+++ F P +IE ++ E LNK V
Sbjct: 344 KEEIERRRAEAAEKRQKMPEDGLSEDKKPFKCFTPKGSSLKIE------ERAEFLNKSVQ 397
Query: 286 E 286
+
Sbjct: 398 K 398
>YLX8_CAEEL (P46504) Hypothetical protein F23F12.8 in chromosome III
precursor
Length = 887
Score = 39.7 bits (91), Expect = 0.009
Identities = 39/159 (24%), Positives = 80/159 (49%), Gaps = 11/159 (6%)
Query: 133 AKKLPGVRELFDKPPELRKRRTRYDIYKRIDAAY-YGYRDDEDG--ILERLEPSAEAAMR 189
A ++ +REL + +L ++R + + ++AA Y +++E I ++ + +
Sbjct: 395 AMEISKIREL--ERLQLERQRKNERVRQELEAARKYKLQEEERQRKIQQQKVEMEQIRQQ 452
Query: 190 REAAEEWYR-LDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREMKERL 248
EA +E R L+ R + V+ E+ EILR++EED +++ + RE +E+
Sbjct: 453 EEARQEQLRVLEEERARELERVRQEELERQHQM--EILRQQEEDQKKKKLEKDREQREQQ 510
Query: 249 EENE-REFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSE 286
E E ++ + + K +KM+ EK K ++L K + +
Sbjct: 511 EAEELNRMIIEKEMKENK--QKMIEEKNKRKMLEKEMED 547
Score = 32.3 bits (72), Expect = 1.5
Identities = 29/112 (25%), Positives = 56/112 (49%), Gaps = 5/112 (4%)
Query: 178 ERLEPSAEAAMRREAAEEWYRLDRIRQ-EARKGVKSGEVAEVSAAVREILREEEEDVVEE 236
E+ E + +R+E E+ L+R R+ E + + E+ + E R E E
Sbjct: 315 EKFEKMEQERLRQEKEEKARELERRRKLEESETARQAELDRQATIYAEQERMAMERNREL 374
Query: 237 ERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDL 288
ER+R E K+R E R+ + + + +E+E++ LE+++ N+ V ++L
Sbjct: 375 ERIRLEE-KKRENERVRQEEIAMEISKIRELERLQLERQRK---NERVRQEL 422
>RA50_METKA (Q8TXI4) DNA double-strand break repair rad50 ATPase
Length = 876
Score = 39.7 bits (91), Expect = 0.009
Identities = 87/351 (24%), Positives = 147/351 (41%), Gaps = 62/351 (17%)
Query: 5 EEKAQSMLNRFITMKAEEKKKPKERRPF--LATECRDLS---EADKWRQQIMREIGRKVA 59
E K ++ R +K +K+ + R L E ++L E K R +RE R+
Sbjct: 181 EAKLETFRERVRDLKGSKKELKRVERELEELKREVKELEPEVEELKERLNELREAKREFE 240
Query: 60 EIQNE-GLGEHRLRDLN---DEINKLIREKSHWERRIVELGG-PNYTRH-SAKMTDLDGN 113
++ E L E+++ L D++ KL+ E ER + LG P+ R + +L
Sbjct: 241 RLEGELRLLENKIESLKGRRDDLRKLVEEGKEAERELQRLGDVPSKVRELENEEAELRRR 300
Query: 114 IVDVPN--PSGRGPGYRYFGAAKKLPGV-RELFDKPPELRKRRTRYDIYK-RIDAAYYGY 169
I ++ N R R A ++L GV REL + E R +K +I A
Sbjct: 301 IEELRNLLDDLRSLRNRLESAEEELEGVKRELEELKDEAGVDPERLVEFKDKIVEASERL 360
Query: 170 RD--DEDGILERLEPSA---------EAAMRREAAEEWYRLDRIRQEARK-GVKSGEVAE 217
RD E+ + +LE + E ++ E E RLD I+ E ++ VK E+ E
Sbjct: 361 RDLRREEELKRKLEKVSDELSELGDREETLQSEYEELQERLDEIQGELKEIRVKEKELLE 420
Query: 218 VSAAVRE-------------------ILREEEEDVV------EEERMRGREMKERLEENE 252
++RE +LR+ E+++ E+ R RE+K+RLE
Sbjct: 421 RIESLREAEGECPVCLRKLPRERAEKLLRDAEKELERLQGREEDLRKERRELKDRLESVR 480
Query: 253 REFVVHVPLPDEKEIEKMVLEKKKMELLNKYVS--EDLMDEQSEAKDMLNI 301
RE E E+M +++ E L + + E+L +E ++ L +
Sbjct: 481 REL--------EGTKERMWRLRERREELERELEEIEELKEELADLSRELGV 523
Score = 38.9 bits (89), Expect = 0.016
Identities = 77/326 (23%), Positives = 135/326 (40%), Gaps = 48/326 (14%)
Query: 6 EKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQNEG 65
+KA+ + + + + + +ER L ++L ++ +++ RE+ E++
Sbjct: 167 KKAREQAHELLRVAEAKLETFRERVRDLKGSKKELKRVERELEELKREVKELEPEVEELK 226
Query: 66 LGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPSGRGP 125
+ LR+ E +L E E +I L G R + +G +
Sbjct: 227 ERLNELREAKREFERLEGELRLLENKIESLKG---RRDDLRKLVEEGKEAE--------- 274
Query: 126 GYRYFGAAKKLPG-VRELFDKPPELRKR----RTRYDIYKRIDAAYYGYRDDEDGI---L 177
R +P VREL ++ ELR+R R D + + ++ +G+ L
Sbjct: 275 --RELQRLGDVPSKVRELENEEAELRRRIEELRNLLDDLRSLRNRLESAEEELEGVKREL 332
Query: 178 ERLEPSAEAAMRR---------EAAEEWYRLDRIRQEARKGVK-SGEVAEVSAAVREILR 227
E L+ A R EA+E L R + RK K S E++E+ E L+
Sbjct: 333 EELKDEAGVDPERLVEFKDKIVEASERLRDLRREEELKRKLEKVSDELSELGDR-EETLQ 391
Query: 228 EEEEDVVE----------EERMRGREMKERLE---ENEREFVVHVPLPDEKEIEKMVLE- 273
E E++ E E R++ +E+ ER+E E E E V + + EK++ +
Sbjct: 392 SEYEELQERLDEIQGELKEIRVKEKELLERIESLREAEGECPVCLRKLPRERAEKLLRDA 451
Query: 274 KKKMELLNKYVSEDLMDEQSEAKDML 299
+K++E L EDL E+ E KD L
Sbjct: 452 EKELERLQGR-EEDLRKERRELKDRL 476
>CALD_CHICK (P12957) Caldesmon (CDM)
Length = 771
Score = 39.7 bits (91), Expect = 0.009
Identities = 37/115 (32%), Positives = 54/115 (46%), Gaps = 12/115 (10%)
Query: 184 AEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEE-ERMRGR 242
A+A R+AAEE + + R+ A + K+ E E AA E+E EE ER +
Sbjct: 321 AKAEEERKAAEERAKAEEERKAAEERAKAEE--ERKAAEERAKAEKERKAAEERERAKAE 378
Query: 243 EMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAKD 297
E K EE R L EK EK +E+KK + + +L+ +Q E K+
Sbjct: 379 EEKRAAEEKAR-------LEAEKLKEKKKMEEKKAQ--EEKAQANLLRKQEEDKE 424
Score = 38.9 bits (89), Expect = 0.016
Identities = 32/114 (28%), Positives = 53/114 (46%), Gaps = 8/114 (7%)
Query: 184 AEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGRE 243
A+A R+AAEE R + +E + + + E A E + EEE EER + +
Sbjct: 306 AKAEEERKAAEERERA-KAEEERKAAEERAKAEEERKAAEERAKAEEERKAAEERAKAEK 364
Query: 244 MKERLEENEREFVVHVPLPDEKEI--EKMVLEKKKMELLNKYVSEDLMDEQSEA 295
++ EE ER +EK EK LE +K++ K + +E+++A
Sbjct: 365 ERKAAEERER-----AKAEEEKRAAEEKARLEAEKLKEKKKMEEKKAQEEKAQA 413
Score = 38.1 bits (87), Expect = 0.027
Identities = 31/119 (26%), Positives = 55/119 (46%), Gaps = 1/119 (0%)
Query: 179 RLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEER 238
+ E +AA R AEE + R +A K K+ E E + A E EE+ +E E+
Sbjct: 335 KAEEERKAAEERAKAEEERKAAEERAKAEKERKAAEERERAKAEEEKRAAEEKARLEAEK 394
Query: 239 MRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEAKD 297
++ ++ E + E + ++ L ++E ++ +E KK L K D+ + KD
Sbjct: 395 LKEKKKMEEKKAQEEKAQANL-LRKQEEDKEAKVEAKKESLPEKLQPTSKKDQVKDNKD 452
Score = 33.5 bits (75), Expect = 0.67
Identities = 26/85 (30%), Positives = 40/85 (46%), Gaps = 8/85 (9%)
Query: 170 RDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREE 229
++ E+ ER + AE R +A EE ++A + + E + +A RE + E
Sbjct: 243 KEKEEAEKEREKLEAEEKERLKAEEE--------KKAAEEKQKAEEEKKAAEERERAKAE 294
Query: 230 EEDVVEEERMRGREMKERLEENERE 254
EE EER R + +ER ERE
Sbjct: 295 EEKRAAEERERAKAEEERKAAEERE 319
Score = 32.0 bits (71), Expect = 1.9
Identities = 63/305 (20%), Positives = 124/305 (40%), Gaps = 48/305 (15%)
Query: 2 ARNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEI 61
A+ EE+ ++ R KAEE++K E R E + E ++ + + + + A +
Sbjct: 334 AKAEEERKAAEER---AKAEEERKAAEERAKAEKERKAAEERERAKAEEEKRAAEEKARL 390
Query: 62 QNEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGNIVDVPNPS 121
+ E L E + K+ +K+ E+ L AK+ ++ + P+
Sbjct: 391 EAEKLKEKK---------KMEEKKAQEEKAQANLLRKQEEDKEAKVEAKKESLPEKLQPT 441
Query: 122 GRGPGYRYFGAAKKLPG--VRELFDKP---PELRKRRTRYDI----YKRIDAAY-----Y 167
+ + +K P ++ ++D+ PE + + ++ K + A+
Sbjct: 442 SKKDQVKDNKDKEKAPKEEMKSVWDRKRGVPEQKAQNGERELTTPKLKSTENAFGRSNLK 501
Query: 168 GYRDDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILR 227
G + E G E+L+ ++EAA E L + R+E RK ++ E + +R
Sbjct: 502 GAANAEAGS-EKLKEK-----QQEAAVELDELKKRREERRKILEEEEQKKKQEEAERKIR 555
Query: 228 EEEE-----DVVEEERMRGREMKER-----LEENEREFVVHVPLPDEKEIEKMVLEKKKM 277
EEEE + +E R E +++ + E ++ F P +IE ++
Sbjct: 556 EEEEKKRMKEEIERRRAEAAEKRQKVPEDGVSEEKKPFKCFSPKGSSLKIE------ERA 609
Query: 278 ELLNK 282
E LNK
Sbjct: 610 EFLNK 614
Score = 31.2 bits (69), Expect = 3.3
Identities = 27/122 (22%), Positives = 45/122 (36%), Gaps = 2/122 (1%)
Query: 171 DDEDGILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEE 230
+ E I + + E A + E +R++ E K K+ E + + ++ E E
Sbjct: 232 EGEQSITDAADKEKEEAEKEREKLEAEEKERLKAEEEK--KAAEEKQKAEEEKKAAEERE 289
Query: 231 EDVVEEERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMD 290
EEE+ E + E ER+ +E K E+ K E K E
Sbjct: 290 RAKAEEEKRAAEERERAKAEEERKAAEERERAKAEEERKAAEERAKAEEERKAAEERAKA 349
Query: 291 EQ 292
E+
Sbjct: 350 EE 351
>SMC_METJA (Q59037) Chromosome partition protein smc homolog
Length = 1169
Score = 38.9 bits (89), Expect = 0.016
Identities = 66/290 (22%), Positives = 122/290 (41%), Gaps = 34/290 (11%)
Query: 3 RNEEKAQSMLNRFITMKAEEKKKPKERRPFLATECRDLSEADKWRQQIMREIGRKVAEIQ 62
R+ K + N +K E +K +E + ++L +K + + E+ K EI
Sbjct: 708 RSSAKKMEIENTLEIIKKNEMRK-REIAEKNTIKIKELELKNKDILEELEELNLKREEIL 766
Query: 63 NEGLGEHRLRDLNDEINKLIREKSHWERRIVELGGPNYTRHSAKMTDLDGN--IVDVPNP 120
N R+ ++ +IN+LI + E+ I EL + +M +++G I++
Sbjct: 767 N------RINEIESKINELIERR---EKIINELKEYESDENLKRMNEIEGELKILEKEKA 817
Query: 121 SGRGPGYRYFGAAKKL--PGVRELFDKPPELRKRRTRYDIYKRIDAAYYGYRDDEDGILE 178
+ + K++ P + EL K EL ++ I ++ + Y + ILE
Sbjct: 818 KLKNEIDKGLTLVKEILIPKIEELNKKVSELINKKV---ILEKNISFYKESIEKNLSILE 874
Query: 179 RLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILRE--EEEDVVEE 236
E E A+ L +++ K E+ + REILR+ + E+ + E
Sbjct: 875 EKRKRYE-----ELAKNLKELTEKKEQLEK-----EIETLERERREILRKVRDIENRINE 924
Query: 237 ERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSE 286
+ + + +LEE ER+ + + KE LEKK +E L Y+ E
Sbjct: 925 LMVEKAKYESKLEEEERKLYLCEKVDVSKE-----LEKKDIEELEIYIGE 969
Score = 35.8 bits (81), Expect = 0.13
Identities = 22/91 (24%), Positives = 49/91 (53%), Gaps = 1/91 (1%)
Query: 211 KSGEVAEVSAAVREILREEEEDVVEEERMRGREMKERLE-ENEREFVVHVPLPDEKEIEK 269
K ++A+ A+ LR+ +E++ ++ R +++E EN E + + + EK
Sbjct: 677 KLNKIADEIIAIESELRKIKEEIERLSKIVKRSSAKKMEIENTLEIIKKNEMRKREIAEK 736
Query: 270 MVLEKKKMELLNKYVSEDLMDEQSEAKDMLN 300
++ K++EL NK + E+L + + +++LN
Sbjct: 737 NTIKIKELELKNKDILEELEELNLKREEILN 767
>M4K4_MOUSE (P97820) Mitogen-activated protein kinase kinase kinase
kinase 4 (EC 2.7.1.37) (MAPK/ERK kinase kinase kinase 4)
(MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase
HGK) (Nck interacting kinase)
Length = 1233
Score = 38.9 bits (89), Expect = 0.016
Identities = 31/108 (28%), Positives = 53/108 (48%), Gaps = 16/108 (14%)
Query: 185 EAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREM 244
E R+ AE R+++ +++ R+ + E RE R++E E+R R +E
Sbjct: 384 EEYKRQLLAERQKRIEQQKEQRRR------LEEQQRREREARRQQER----EQRRREQEE 433
Query: 245 KERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQ 292
K RLEE ER +E+E + EK+++E +Y+ L +EQ
Sbjct: 434 KRRLEELERR------RKEEEERRRAEEEKRRVEREQEYIRRQLEEEQ 475
Score = 31.2 bits (69), Expect = 3.3
Identities = 32/128 (25%), Positives = 61/128 (47%), Gaps = 21/128 (16%)
Query: 176 ILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVE 235
+ ER + + +R EE R +R + ++ + E + E+ R +E E
Sbjct: 391 LAERQKRIEQQKEQRRRLEEQQRREREARRQQEREQRRREQEEKRRLEELERRRKE---E 447
Query: 236 EERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQSEA 295
EER R E K R+E E+E++ +++E+ E++ +E+L + L+ EQ+
Sbjct: 448 EERRRAEEEKRRVER-EQEYI-------RRQLEE---EQRHLEIL----QQQLLQEQAM- 491
Query: 296 KDMLNIHR 303
+L+ HR
Sbjct: 492 --LLHDHR 497
>M4K4_HUMAN (O95819) Mitogen-activated protein kinase kinase kinase
kinase 4 (EC 2.7.1.37) (MAPK/ERK kinase kinase kinase 4)
(MEK kinase kinase 4) (MEKKK 4) (HPK/GCK-like kinase
HGK) (Nck interacting kinase)
Length = 1239
Score = 38.9 bits (89), Expect = 0.016
Identities = 31/108 (28%), Positives = 53/108 (48%), Gaps = 16/108 (14%)
Query: 185 EAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVEEERMRGREM 244
E R+ AE R+++ +++ R+ + E RE R++E E+R R +E
Sbjct: 385 EEYKRQLLAERQKRIEQQKEQRRR------LEEQQRREREARRQQER----EQRRREQEE 434
Query: 245 KERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQ 292
K RLEE ER +E+E + EK+++E +Y+ L +EQ
Sbjct: 435 KRRLEELERR------RKEEEERRRAEEEKRRVEREQEYIRRQLEEEQ 476
Score = 30.4 bits (67), Expect = 5.6
Identities = 29/118 (24%), Positives = 56/118 (46%), Gaps = 18/118 (15%)
Query: 176 ILERLEPSAEAAMRREAAEEWYRLDRIRQEARKGVKSGEVAEVSAAVREILREEEEDVVE 235
+ ER + + +R EE R +R + ++ + E + E+ R +E E
Sbjct: 392 LAERQKRIEQQKEQRRRLEEQQRREREARRQQEREQRRREQEEKRRLEELERRRKE---E 448
Query: 236 EERMRGREMKERLEENEREFVVHVPLPDEKEIEKMVLEKKKMELLNKYVSEDLMDEQS 293
EER R E K R+E E+E++ +++E+ E++ +E+L + L+ EQ+
Sbjct: 449 EERRRAEEEKRRVER-EQEYI-------RRQLEE---EQRHLEVL----QQQLLQEQA 491
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.134 0.374
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,867,520
Number of Sequences: 164201
Number of extensions: 1746688
Number of successful extensions: 9150
Number of sequences better than 10.0: 414
Number of HSP's better than 10.0 without gapping: 69
Number of HSP's successfully gapped in prelim test: 359
Number of HSP's that attempted gapping in prelim test: 7541
Number of HSP's gapped (non-prelim): 1121
length of query: 303
length of database: 59,974,054
effective HSP length: 110
effective length of query: 193
effective length of database: 41,911,944
effective search space: 8089005192
effective search space used: 8089005192
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0282.6


