

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0249.1
(68 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
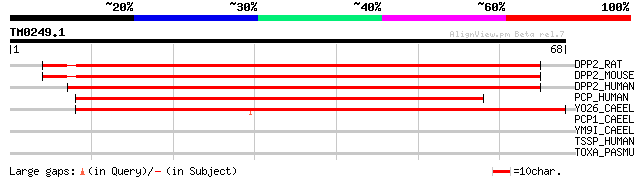
Sequences producing significant alignments: (bits) Value
DPP2_RAT (Q9EPB1) Dipeptidyl-peptidase II precursor (EC 3.4.14.2... 70 1e-12
DPP2_MOUSE (Q9ET22) Dipeptidyl-peptidase II precursor (EC 3.4.14... 69 3e-12
DPP2_HUMAN (Q9UHL4) Dipeptidyl-peptidase II precursor (EC 3.4.14... 68 5e-12
PCP_HUMAN (P42785) Lysosomal Pro-X carboxypeptidase precursor (E... 65 4e-11
YO26_CAEEL (P34676) Putative serine protease Z688.6 precursor (E... 59 2e-09
PCP1_CAEEL (P34610) Putative serine protease pcp-1 precursor (EC... 40 0.001
YM9I_CAEEL (P90893) Putative serine protease F56F10.1 precursor ... 28 5.7
TSSP_HUMAN (Q9NQE7) Thymus-specific serine protease precursor (E... 28 5.7
TOXA_PASMU (P17452) Dermonecrotic toxin (DNT) (PMT) (Mitogenic t... 27 9.8
>DPP2_RAT (Q9EPB1) Dipeptidyl-peptidase II precursor (EC 3.4.14.2)
(DPP II) (Dipeptidyl aminopeptidase II) (Quiescent cell
proline dipeptidase)
Length = 500
Score = 69.7 bits (169), Expect = 1e-12
Identities = 33/61 (54%), Positives = 45/61 (73%), Gaps = 1/61 (1%)
Query: 5 DLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLKDV 64
DLK +ASNIIF NG DPW+GGG+ +N+S +++A+ + GAHH DLR S EDP + +V
Sbjct: 413 DLK-AASNIIFSNGDLDPWAGGGIQRNLSTSIIAVTIQGGAHHLDLRASNSEDPPSVVEV 471
Query: 65 R 65
R
Sbjct: 472 R 472
>DPP2_MOUSE (Q9ET22) Dipeptidyl-peptidase II precursor (EC 3.4.14.2)
(DPP II) (Dipeptidyl aminopeptidase II) (Quiescent cell
proline dipeptidase) (Dipeptidyl peptidase 7)
Length = 506
Score = 68.6 bits (166), Expect = 3e-12
Identities = 33/61 (54%), Positives = 44/61 (72%), Gaps = 1/61 (1%)
Query: 5 DLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLKDV 64
DLK +ASNIIF NG DPW+GGG+ N+S +++A+ + GAHH DLR S EDP + +V
Sbjct: 413 DLK-AASNIIFSNGDLDPWAGGGIQSNLSTSVIAVTIQGGAHHLDLRASNSEDPPSVVEV 471
Query: 65 R 65
R
Sbjct: 472 R 472
>DPP2_HUMAN (Q9UHL4) Dipeptidyl-peptidase II precursor (EC 3.4.14.2)
(DPP II) (Dipeptidyl aminopeptidase II) (Quiescent cell
proline dipeptidase) (Dipeptidyl peptidase 7)
Length = 492
Score = 67.8 bits (164), Expect = 5e-12
Identities = 30/58 (51%), Positives = 42/58 (71%)
Query: 8 RSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLKDVR 65
R+ASNIIF NG DPW+GGG+ +N+S +++A+ + GAHH DLR S EDP + + R
Sbjct: 405 RAASNIIFSNGNLDPWAGGGIRRNLSASVIAVTIQGGAHHLDLRASHPEDPASVVEAR 462
>PCP_HUMAN (P42785) Lysosomal Pro-X carboxypeptidase precursor (EC
3.4.16.2) (Prolylcarboxypeptidase) (PRCP) (Proline
carboxypeptidase) (Angiotensinase C) (Lysosomal
carboxypeptidase C)
Length = 496
Score = 64.7 bits (156), Expect = 4e-11
Identities = 30/50 (60%), Positives = 35/50 (70%)
Query: 9 SASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDP 58
S +NI+F NG DPWSGGGV K+I+ TLVA+ EGAHH DLR DP
Sbjct: 418 SHTNIVFSNGELDPWSGGGVTKDITDTLVAVTISEGAHHLDLRTKNALDP 467
>YO26_CAEEL (P34676) Putative serine protease Z688.6 precursor (EC
3.4.-.-)
Length = 507
Score = 59.3 bits (142), Expect = 2e-09
Identities = 28/62 (45%), Positives = 41/62 (65%), Gaps = 2/62 (3%)
Query: 9 SASNIIFYNGLRDPWSGGGV--LKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLKDVRI 66
SASNI+F NG DPWSGGG + ++++++ K+GAHH DLR + +D + +K VR
Sbjct: 427 SASNIVFSNGYLDPWSGGGYDHSDKVQGSVISVILKQGAHHYDLRGAHPQDTEEVKKVRA 486
Query: 67 KE 68
E
Sbjct: 487 ME 488
>PCP1_CAEEL (P34610) Putative serine protease pcp-1 precursor (EC
3.4.-.-)
Length = 565
Score = 40.0 bits (92), Expect = 0.001
Identities = 21/61 (34%), Positives = 30/61 (48%), Gaps = 3/61 (4%)
Query: 10 ASNIIFYNGLRDPWSGGGV---LKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLKDVRI 66
+SN+I G DPWSGGG N ++ + + AHH DLR DP + + R
Sbjct: 440 SSNLILTQGHLDPWSGGGYKVDQNNAARGIYVLEIPGSAHHLDLRQPNTCDPNTVTNARF 499
Query: 67 K 67
+
Sbjct: 500 Q 500
>YM9I_CAEEL (P90893) Putative serine protease F56F10.1 precursor (EC
3.4.-.-)
Length = 540
Score = 27.7 bits (60), Expect = 5.7
Identities = 16/51 (31%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Query: 9 SASNIIFYNGLRDPWSGGGVLKNI-SKTLVAIVAKEGAHHRDLRYSTKEDP 58
+A+N++ NG DPW G I S++L+ + AH D+ S +P
Sbjct: 441 NATNVVLPNGSLDPWHALGTYGTIKSQSLLPYLINGTAHCGDMYPSYDGEP 491
>TSSP_HUMAN (Q9NQE7) Thymus-specific serine protease precursor (EC
3.4.-.-)
Length = 514
Score = 27.7 bits (60), Expect = 5.7
Identities = 13/53 (24%), Positives = 25/53 (46%)
Query: 10 ASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWLK 62
A+ ++F NG DPW V + + + ++ + G+H D+ D L+
Sbjct: 436 ANKVLFVNGDTDPWHVLSVTQALGSSESTLLIRTGSHCLDMAPERPSDSPSLR 488
>TOXA_PASMU (P17452) Dermonecrotic toxin (DNT) (PMT) (Mitogenic
toxin)
Length = 1285
Score = 26.9 bits (58), Expect = 9.8
Identities = 14/47 (29%), Positives = 23/47 (48%), Gaps = 3/47 (6%)
Query: 14 IFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKW 60
IF D W +L +++ + V KEGA++ D Y + +P W
Sbjct: 117 IFVERYFDDWD---LLNSLASNGIYSVGKEGAYYPDHDYGPEYNPVW 160
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,953,463
Number of Sequences: 164201
Number of extensions: 245834
Number of successful extensions: 419
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 410
Number of HSP's gapped (non-prelim): 9
length of query: 68
length of database: 59,974,054
effective HSP length: 44
effective length of query: 24
effective length of database: 52,749,210
effective search space: 1265981040
effective search space used: 1265981040
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0249.1


