

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
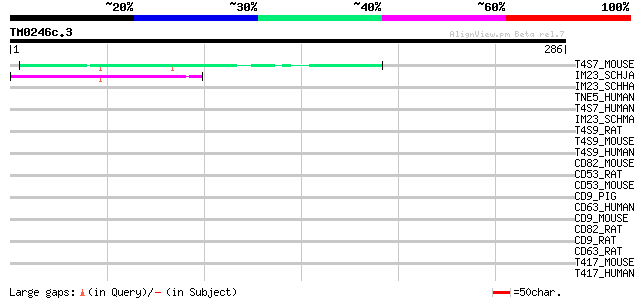
Query= TM0246c.3
(286 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
T4S7_MOUSE (Q9DCK3) Transmembrane 4 superfamily member 7 47 7e-05
IM23_SCHJA (P27591) 23 kDa integral membrane protein (SJ23) 45 3e-04
IM23_SCHHA (Q26499) 23 kDa integral membrane protein (Sh23) 43 0.001
TNE5_HUMAN (O75954) Tetraspan NET-5 42 0.002
T4S7_HUMAN (O14817) Transmembrane 4 superfamily member 7 (Novel ... 42 0.002
IM23_SCHMA (P19331) 23 kDa integral membrane protein (SM23) 42 0.002
T4S9_RAT (Q68VK5) Transmembrane 4 superfamily member 9 (Tetraspa... 41 0.003
T4S9_MOUSE (P62080) Transmembrane 4 superfamily member 9 (Tetras... 41 0.003
T4S9_HUMAN (P62079) Transmembrane 4 superfamily member 9 (Tetras... 41 0.003
CD82_MOUSE (P40237) CD82 antigen (Inducible membrane protein R2)... 40 0.009
CD53_RAT (P24485) Leukocyte surface antigen CD53 (Cell surface g... 39 0.011
CD53_MOUSE (Q61451) Leukocyte surface antigen CD53 (Cell surface... 38 0.025
CD9_PIG (Q8WMQ3) CD9 antigen 38 0.033
CD63_HUMAN (P08962) CD63 antigen (Melanoma-associated antigen ME... 38 0.033
CD9_MOUSE (P40240) CD9 antigen 37 0.043
CD82_RAT (O70352) CD82 antigen (Metastasis suppressor homolog) 37 0.043
CD9_RAT (P40241) CD9 antigen 37 0.056
CD63_RAT (P28648) CD63 antigen (AD1 antigen) 37 0.056
T417_MOUSE (Q9D7W4) Transmembrane 4 superfamily member 17 (Tetra... 37 0.073
T417_HUMAN (Q96FV3) Transmembrane 4 superfamily member 17 (Tetra... 37 0.073
>T4S7_MOUSE (Q9DCK3) Transmembrane 4 superfamily member 7
Length = 238
Score = 46.6 bits (109), Expect = 7e-05
Identities = 48/196 (24%), Positives = 76/196 (38%), Gaps = 29/196 (14%)
Query: 6 HLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-------LIIIGVSIMVVSL 58
+L+ N L +L +LG GIWL++ N L +P LI+ G +M +
Sbjct: 11 YLMFAFNLLFWLGGCGVLGVGIWLAATQGNFATLS-SSFPSLSAANLLIVTGTFVMAIGF 69
Query: 59 AGFAGACYRNTFLMRLYLVVMFLV--IAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQ 116
G GA N L+ + V++ LV + I + FAY ++ L +G Y Q
Sbjct: 70 VGCIGALKENKCLLLTFFVLLLLVFLLEATIAVLFFAYSDKIDSYAQQDLKKGLHLYGTQ 129
Query: 117 DYSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCC 176
V + W I + D + C G+ D F + T V CC
Sbjct: 130 G-------NVGLTNAWSIIQT---DFRCC---------GVSNYTDWFEVYNATRVPDSCC 170
Query: 177 KPPTDCGYIYQNETVW 192
+D +++ T W
Sbjct: 171 LEFSDSCGLHEPGTWW 186
>IM23_SCHJA (P27591) 23 kDa integral membrane protein (SJ23)
Length = 218
Score = 44.7 bits (104), Expect = 3e-04
Identities = 29/105 (27%), Positives = 53/105 (49%), Gaps = 7/105 (6%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-----LIIIGVSIMV 55
MR + +LN + L S+ ++G G ++ + + + W +I++GV I++
Sbjct: 8 MRCLKSCVFILNIICLLCSLVLIGAGAYVEVKFSQYEANLHKVWQAAPIAIIVVGVVILI 67
Query: 56 VSLAGFAGACYRNTFLMRLY-LVVMFLVIAVLIGFIIFAYVVTDK 99
VS G GA N ++ +Y ++ L+IA L+ I+ A V DK
Sbjct: 68 VSFLGCCGAIKENVCMLYMYAFFLIVLLIAELVAAIV-AVVYKDK 111
>IM23_SCHHA (Q26499) 23 kDa integral membrane protein (Sh23)
Length = 218
Score = 42.7 bits (99), Expect = 0.001
Identities = 25/104 (24%), Positives = 48/104 (46%), Gaps = 5/104 (4%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-----LIIIGVSIMV 55
MR + VLN + L S+ ++G G ++ + + W +I++GV I++
Sbjct: 8 MRCLKSCVFVLNIICLLCSLVLIGAGAYVEVKFSQYGDNLHKVWQAAPIAIIVVGVIILI 67
Query: 56 VSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDK 99
VS G GA N ++ +Y + +++ + I A V D+
Sbjct: 68 VSFLGCCGAIKENVCMLYMYAFFLIILLIAELAAAIVAVVYKDR 111
>TNE5_HUMAN (O75954) Tetraspan NET-5
Length = 239
Score = 42.0 bits (97), Expect = 0.002
Identities = 30/124 (24%), Positives = 57/124 (45%), Gaps = 10/124 (8%)
Query: 6 HLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-------LIIIGVSIMVVSL 58
+++ + N + +L +LG GIWLS N +P +I IG +MV
Sbjct: 11 YMMFLFNLIFWLCGCGLLGVGIWLSVSQGNFATFS-PSFPSLSAANLVIAIGTIVMVTGF 69
Query: 59 AGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDK--GSGRRVLNRGYMDYYLQ 116
G GA N L+ + +V+ +++ + +I +V DK + ++ L G + Y+ +
Sbjct: 70 LGCLGAIKENKCLLLSFFIVLLVILLAELILLILFFVYMDKVNENAKKDLKEGLLLYHTE 129
Query: 117 DYSG 120
+ G
Sbjct: 130 NNVG 133
>T4S7_HUMAN (O14817) Transmembrane 4 superfamily member 7 (Novel
antigen 2) (NAG-2) (Tetraspanin 4) (Tspan-4)
Length = 238
Score = 42.0 bits (97), Expect = 0.002
Identities = 46/201 (22%), Positives = 77/201 (37%), Gaps = 29/201 (14%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-------LIIIGVSI 53
++ +L+ N L +L +LG GIWL++ + L +P LII G +
Sbjct: 6 LQAVKYLMFAFNLLFWLGGCGVLGVGIWLAATQGSFATLS-SSFPSLSAANLLIITGAFV 64
Query: 54 MVVSLAGFAGACYRNTFLMRLYLVVMFLV--IAVLIGFIIFAYVVTDKGSGRRVLNRGYM 111
M + G GA N L+ + +++ LV + I + FAY ++ L +G
Sbjct: 65 MAIGFVGCLGAIKENKCLLLTFFLLLLLVFLLEATIAILFFAYTDKIDRYAQQDLKKGLH 124
Query: 112 DYYLQDYSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSV 171
Y Q V + W I + D + C G+ D F + T V
Sbjct: 125 LYGTQG-------NVGLTNAWSIIQT---DFRCC---------GVSNYTDWFEVYNATRV 165
Query: 172 QSGCCKPPTDCGYIYQNETVW 192
CC ++ ++ T W
Sbjct: 166 PDSCCLEFSESCGLHAPGTWW 186
>IM23_SCHMA (P19331) 23 kDa integral membrane protein (SM23)
Length = 218
Score = 42.0 bits (97), Expect = 0.002
Identities = 25/104 (24%), Positives = 48/104 (46%), Gaps = 5/104 (4%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-----LIIIGVSIMV 55
MR + VLN + L S+ ++G G ++ + + W +I++GV I++
Sbjct: 8 MRCLKSCVFVLNIICLLCSLVLIGAGAYVEVKFSQYGDNLHKVWQAAPIAIIVVGVIILI 67
Query: 56 VSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDK 99
VS G GA N ++ +Y + +++ + I A V D+
Sbjct: 68 VSFLGCCGAIKENVCMLYMYAFFLVVLLIAELAAAIVAVVYKDR 111
>T4S9_RAT (Q68VK5) Transmembrane 4 superfamily member 9 (Tetraspanin
5) (Tspan-5)
Length = 268
Score = 41.2 bits (95), Expect = 0.003
Identities = 26/95 (27%), Positives = 48/95 (50%), Gaps = 8/95 (8%)
Query: 12 NFLTFLLSIPILGGGIW-------LSSRANNTDCLKFLQ-WPLIIIGVSIMVVSLAGFAG 63
N + + L I LG G+W LS+ ++ TD F W +++G + ++ AG G
Sbjct: 23 NVIFWFLGITFLGIGLWAWNEKGVLSNISSITDLGGFDPVWLFLVVGGVMFILGFAGCIG 82
Query: 64 ACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTD 98
A NTFL++ + V + ++ + + + A+V D
Sbjct: 83 ALRENTFLLKFFSVFLGIIFFLELTAGVLAFVFKD 117
>T4S9_MOUSE (P62080) Transmembrane 4 superfamily member 9
(Tetraspanin 5) (Tspan-5) (Tetraspan NET-4)
Length = 268
Score = 41.2 bits (95), Expect = 0.003
Identities = 26/95 (27%), Positives = 48/95 (50%), Gaps = 8/95 (8%)
Query: 12 NFLTFLLSIPILGGGIW-------LSSRANNTDCLKFLQ-WPLIIIGVSIMVVSLAGFAG 63
N + + L I LG G+W LS+ ++ TD F W +++G + ++ AG G
Sbjct: 23 NVIFWFLGITFLGIGLWAWNEKGVLSNISSITDLGGFDPVWLFLVVGGVMFILGFAGCIG 82
Query: 64 ACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTD 98
A NTFL++ + V + ++ + + + A+V D
Sbjct: 83 ALRENTFLLKFFSVFLGIIFFLELTAGVLAFVFKD 117
>T4S9_HUMAN (P62079) Transmembrane 4 superfamily member 9
(Tetraspanin 5) (Tspan-5) (Tetraspan NET-4)
Length = 268
Score = 41.2 bits (95), Expect = 0.003
Identities = 26/95 (27%), Positives = 48/95 (50%), Gaps = 8/95 (8%)
Query: 12 NFLTFLLSIPILGGGIW-------LSSRANNTDCLKFLQ-WPLIIIGVSIMVVSLAGFAG 63
N + + L I LG G+W LS+ ++ TD F W +++G + ++ AG G
Sbjct: 23 NVIFWFLGITFLGIGLWAWNEKGVLSNISSITDLGGFDPVWLFLVVGGVMFILGFAGCIG 82
Query: 64 ACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTD 98
A NTFL++ + V + ++ + + + A+V D
Sbjct: 83 ALRENTFLLKFFSVFLGIIFFLELTAGVLAFVFKD 117
>CD82_MOUSE (P40237) CD82 antigen (Inducible membrane protein R2)
(C33 antigen) (IA4)
Length = 266
Score = 39.7 bits (91), Expect = 0.009
Identities = 25/107 (23%), Positives = 48/107 (44%), Gaps = 8/107 (7%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRAN--------NTDCLKFLQWPLIIIGVS 52
++ + + + + N L F+L ILG G+W+ + N ++ L+ + I +G
Sbjct: 6 VKVTKYFLFLFNLLFFILGAVILGFGVWILADKNSFISVLQTSSSSLQVGAYVFIGVGAI 65
Query: 53 IMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDK 99
+V+ G GA L+ LY V + L++ + + Y DK
Sbjct: 66 TIVMGFLGCIGAVNEVRCLLGLYFVFLLLILIAQVTVGVLFYFNADK 112
>CD53_RAT (P24485) Leukocyte surface antigen CD53 (Cell surface
glycoprotein CD53) (Leukocyte antigen MRC OX-44)
Length = 218
Score = 39.3 bits (90), Expect = 0.011
Identities = 29/103 (28%), Positives = 50/103 (48%), Gaps = 11/103 (10%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-------LIIIGVSI 53
++ +++ NFL ++ ILG GI L NT + F P L+I+G I
Sbjct: 5 LKLLKYVLFFFNFLFWVCGCCILGFGIHLL--VQNTYGILFRNLPFLTLGNVLVIVGSII 62
Query: 54 MVVSLAGFAGACYRNTFLMRLYLVVMFLVI--AVLIGFIIFAY 94
MVV+ G G+ N L+ + V++ L++ V + ++F Y
Sbjct: 63 MVVAFLGCMGSIKENKCLLMSFFVLLLLILLAEVTLAILLFVY 105
>CD53_MOUSE (Q61451) Leukocyte surface antigen CD53 (Cell surface
glycoprotein CD53)
Length = 218
Score = 38.1 bits (87), Expect = 0.025
Identities = 27/103 (26%), Positives = 50/103 (48%), Gaps = 11/103 (10%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWP-------LIIIGVSI 53
++ +++ + N L ++ ILG GI+ NT + F P L+I+G I
Sbjct: 5 LKLLKYVLFIFNLLFWVCGCCILGFGIYFL--VQNTYGVLFRNLPFLTLGNILVIVGSII 62
Query: 54 MVVSLAGFAGACYRNTFLMRLYLVVMFLVI--AVLIGFIIFAY 94
MVV+ G G+ N L+ + V++ +++ V I ++F Y
Sbjct: 63 MVVAFLGCMGSIKENKCLLMSFFVLLLIILLAEVTIAILLFVY 105
>CD9_PIG (Q8WMQ3) CD9 antigen
Length = 225
Score = 37.7 bits (86), Expect = 0.033
Identities = 29/123 (23%), Positives = 52/123 (41%), Gaps = 15/123 (12%)
Query: 6 HLIGVLNFLTFLLSIPILGGGIWLS---------SRANNTDCLKFLQWPLIIIGVSIMVV 56
+L+ NF+ +L I +L G+WL + NN + LI G +MVV
Sbjct: 11 YLLFGFNFIFWLAGIAVLAIGLWLRFDSQTKSIFEQENNNSSFYTGVYILIGAGALMMVV 70
Query: 57 SLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQ 116
G GA + ++ L+ + ++ A+ I I+ Y D+ + + D+Y
Sbjct: 71 GFLGCCGAVQESQCMLGLFFGFLLVIFAIEIAAAIWGYSHKDQ------VIKEVQDFYRD 124
Query: 117 DYS 119
Y+
Sbjct: 125 TYN 127
>CD63_HUMAN (P08962) CD63 antigen (Melanoma-associated antigen
ME491) (Lysosome-associated membrane glycoprotein 3)
(LAMP-3) (Ocular melanoma-associated antigen) (OMA81H)
(Granulophysin)
Length = 237
Score = 37.7 bits (86), Expect = 0.033
Identities = 18/54 (33%), Positives = 29/54 (53%)
Query: 46 LIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDK 99
+I +GV + +V+ G GAC N LM + + + L++ V + I YV DK
Sbjct: 56 IIAVGVFLFLVAFVGCCGACKENYCLMITFAIFLSLIMLVEVAAAIAGYVFRDK 109
>CD9_MOUSE (P40240) CD9 antigen
Length = 225
Score = 37.4 bits (85), Expect = 0.043
Identities = 29/118 (24%), Positives = 50/118 (41%), Gaps = 11/118 (9%)
Query: 6 HLIGVLNFLTFLLSIPILGGGIWLS---------SRANNTDCLKFLQWPLIIIGVSIMVV 56
+L+ NF+ +L I +L G+WL + NN + LI G +M+V
Sbjct: 11 YLLFGFNFIFWLAGIAVLAIGLWLRFDSQTKSIFEQENNHSSFYTGVYILIGAGALMMLV 70
Query: 57 SLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYY 114
G GA + ++ L+ + ++ A+ I ++ Y T K + L Y D Y
Sbjct: 71 GFLGCCGAVQESQCMLGLFFGFLLVIFAIEIAAAVWGY--THKDEVIKELQEFYKDTY 126
>CD82_RAT (O70352) CD82 antigen (Metastasis suppressor homolog)
Length = 266
Score = 37.4 bits (85), Expect = 0.043
Identities = 24/107 (22%), Positives = 47/107 (43%), Gaps = 8/107 (7%)
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIW--------LSSRANNTDCLKFLQWPLIIIGVS 52
++ + + + + N L F+L ILG G+W +S ++ L+ + I +G
Sbjct: 6 VKVTKYFLFLFNLLFFILGAVILGFGVWILADKSSFISVLQTSSSSLQVGAYVFIGVGAI 65
Query: 53 IMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDK 99
M++ G GA L+ LY V + L++ + + Y +K
Sbjct: 66 TMLMGFLGCIGAVNEVRCLLGLYFVFLLLILIAQVTVEVLFYFNANK 112
>CD9_RAT (P40241) CD9 antigen
Length = 225
Score = 37.0 bits (84), Expect = 0.056
Identities = 30/118 (25%), Positives = 51/118 (42%), Gaps = 11/118 (9%)
Query: 6 HLIGVLNFLTFLLSIPILGGGIWL-------SSRANNTDCLKFLQWPLIIIGVS--IMVV 56
+L+ NF+ +L I +L G+WL S T+ F I+IG +M+V
Sbjct: 11 YLLFGFNFIFWLAGIAVLAIGLWLRFDSQTKSIFEQETNHSSFYTGVYILIGAGALMMLV 70
Query: 57 SLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYY 114
G GA + ++ L+ + ++ A+ I ++ Y T K + L Y D Y
Sbjct: 71 GFLGCCGAVQESQCMLGLFFGFLLVIFAIEIAAAVWGY--THKDEVIKELQEFYKDTY 126
>CD63_RAT (P28648) CD63 antigen (AD1 antigen)
Length = 237
Score = 37.0 bits (84), Expect = 0.056
Identities = 33/155 (21%), Positives = 61/155 (39%), Gaps = 26/155 (16%)
Query: 46 LIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRV 105
+I +G + +V+ G GAC N LM + + + L++ V + I YV
Sbjct: 56 IIAVGAFLFLVAFVGCCGACKENYCLMITFAIFLSLIMLVEVAVAIAGYV---------- 105
Query: 106 LNRGYMDYYLQDYSGWLEERVASHSYWGKIASCV----RDSKVCAKMGRVDSNGIPEPAD 161
+ D ++S ++++ ++ K A+ + +++K C D IP A
Sbjct: 106 ----FRDQVKSEFSKSFQKQMQNYLTDNKTATILDKLQKENKCCGASNYTDWERIPGMAK 161
Query: 162 VFYLRKLTSVQSGCCKPPT-DCGYIYQNETVWNLG 195
V CC T CG ++ T+ G
Sbjct: 162 -------DRVPDSCCINITVGCGNDFKESTIHTQG 189
>T417_MOUSE (Q9D7W4) Transmembrane 4 superfamily member 17
(Tetraspan protein SB134) (F-box only protein 23)
Length = 270
Score = 36.6 bits (83), Expect = 0.073
Identities = 17/55 (30%), Positives = 30/55 (53%)
Query: 44 WPLIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTD 98
W +++G + V+ AG GA NTFL++ + V + L+ + + I A+V D
Sbjct: 65 WLFVVVGGVMSVLGFAGCIGALRENTFLLKFFSVFLGLIFFLELAAGILAFVFKD 119
>T417_HUMAN (Q96FV3) Transmembrane 4 superfamily member 17
(Tetraspan protein SB134) (F-box only protein 23)
Length = 263
Score = 36.6 bits (83), Expect = 0.073
Identities = 17/55 (30%), Positives = 30/55 (53%)
Query: 44 WPLIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTD 98
W +++G + V+ AG GA NTFL++ + V + L+ + + I A+V D
Sbjct: 65 WLFVVVGGVMSVLGFAGCIGALRENTFLLKFFSVFLGLIFFLELATGILAFVFKD 119
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,623,392
Number of Sequences: 164201
Number of extensions: 1432679
Number of successful extensions: 4644
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 29
Number of HSP's that attempted gapping in prelim test: 4602
Number of HSP's gapped (non-prelim): 59
length of query: 286
length of database: 59,974,054
effective HSP length: 109
effective length of query: 177
effective length of database: 42,076,145
effective search space: 7447477665
effective search space used: 7447477665
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0246c.3


