

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.4
(606 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
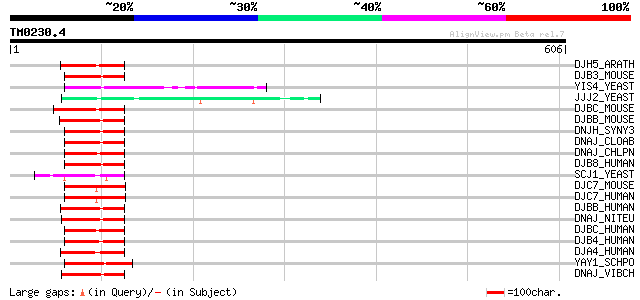
Score E
Sequences producing significant alignments: (bits) Value
DJH5_ARATH (Q9FH28) J domain protein At5g49060 63 3e-09
DJB3_MOUSE (O35723) DnaJ homolog subfamily B member 3 (DnaJ prot... 60 2e-08
YIS4_YEAST (P40564) Hypothetical 48.6 kDa protein in BET1-PAN1 i... 60 2e-08
JJJ2_YEAST (P46997) J protein type 2 60 2e-08
DJBC_MOUSE (Q9QYI4) DnaJ homolog subfamily B member 12 (mDJ10) 59 3e-08
DJBB_MOUSE (Q99KV1) DnaJ homolog subfamily B member 11 precursor 59 3e-08
DNJH_SYNY3 (P50027) DnAJ-like protein slr0093 59 5e-08
DNAJ_CLOAB (P30725) Chaperone protein dnaJ 59 5e-08
DNAJ_CHLPN (Q9Z9E9) Chaperone protein dnaJ 59 5e-08
DJB8_HUMAN (Q8NHS0) DnaJ homolog subfamily B member 8 59 5e-08
SCJ1_YEAST (P25303) DnaJ-related protein SCJ1 58 8e-08
DJC7_MOUSE (Q9QYI3) DnaJ homolog subfamily C member 7 (Tetratric... 57 1e-07
DJC7_HUMAN (Q99615) DnaJ homolog subfamily C member 7 (Tetratric... 57 1e-07
DJBB_HUMAN (Q9UBS4) DnaJ homolog subfamily B member 11 precursor... 57 1e-07
DNAJ_NITEU (O06431) Chaperone protein dnaJ 57 1e-07
DJBC_HUMAN (Q9NXW2) DnaJ homolog subfamily B member 12 57 1e-07
DJB4_HUMAN (Q9UDY4) DnaJ homolog subfamily B member 4 (Heat shoc... 57 1e-07
DJA4_HUMAN (Q8WW22) DnaJ homolog subfamily A member 4 57 1e-07
YAY1_SCHPO (Q10209) Hypothetical J-domain protein C4H3.01 in chr... 57 2e-07
DNAJ_VIBCH (O34242) Chaperone protein dnaJ 57 2e-07
>DJH5_ARATH (Q9FH28) J domain protein At5g49060
Length = 350
Score = 62.8 bits (151), Expect = 3e-09
Identities = 32/70 (45%), Positives = 45/70 (63%), Gaps = 3/70 (4%)
Query: 56 ILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVL 115
I+ D+Y +LG+E + + +RK Y+KLSL +HPDKN A SEEAFK V +AF L
Sbjct: 94 IIRNNDYYAILGLEKNCSVDEIRKAYRKLSLKVHPDKNK---APGSEEAFKKVSKAFTCL 150
Query: 116 SDSALKKGYD 125
SD ++ +D
Sbjct: 151 SDGNSRRQFD 160
>DJB3_MOUSE (O35723) DnaJ homolog subfamily B member 3 (DnaJ protein
homolog 3) (Heat shock J3 protein) (HSJ-3) (MSJ-1)
Length = 242
Score = 60.1 bits (144), Expect = 2e-08
Identities = 32/65 (49%), Positives = 42/65 (64%), Gaps = 1/65 (1%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y VLGV A++ A+RK Y+KL+L HPDKN H A E FK V +A+ VLSD
Sbjct: 3 DYYEVLGVPRQASAEAIRKAYRKLALKWHPDKNPEHKEEA-ERRFKQVAQAYEVLSDVRK 61
Query: 121 KKGYD 125
++ YD
Sbjct: 62 REVYD 66
>YIS4_YEAST (P40564) Hypothetical 48.6 kDa protein in BET1-PAN1
intergenic region
Length = 432
Score = 59.7 bits (143), Expect = 2e-08
Identities = 58/226 (25%), Positives = 102/226 (44%), Gaps = 24/226 (10%)
Query: 60 TDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
T++Y +LGV A+S ++K Y+K S+ HPDKN A E F+ + EA++VL D
Sbjct: 5 TEYYDLLGVSTTASSIEIKKAYRKKSIQEHPDKNPNDPTAT--ERFQAISEAYQVLGDDD 62
Query: 120 LKKGYDAELRKKEAPT--FWTACSACRLLHQFE--RRYVGHSLVCPNCNKSFEAVEAVLS 175
L+ YD RK+ P F A ++ + Y+G ++ N K+ E
Sbjct: 63 LRAKYDKYGRKEAIPQGGFEDAAEQFSVIFGGDAFASYIGELMLLKNLQKTEEL------ 116
Query: 176 DGSSDEGEKVGVRRNEKREVVLGVAAPNGVKGKVGG-EKGLKGKVGDGDGVGDVGGGGSK 234
+ DE EK EK V +P GK G + +G+ + D +
Sbjct: 117 -NAEDEAEK------EKENVETMEESP--ADGKTNGTTNAVDAALGNTNEKDDKNKARTT 167
Query: 235 KRMRSVGDVFERSKPKGIITGEETMTLAEFQSQVKRKVQQEKVKDK 280
+V D ++++ G ++ L +F+ + ++V+++K D+
Sbjct: 168 SGNLTVHDGNKKNEQVGAEAKKKKTKLEQFEEE--QEVEKQKRVDQ 211
>JJJ2_YEAST (P46997) J protein type 2
Length = 482
Score = 59.7 bits (143), Expect = 2e-08
Identities = 76/308 (24%), Positives = 119/308 (37%), Gaps = 45/308 (14%)
Query: 57 LSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLS 116
L T +Y++LG+ A S+ V K Y KL+ LLHPDK + SEE FK V A +L+
Sbjct: 9 LDRTTYYSILGLTSNATSSEVHKSYLKLARLLHPDKTKSD---KSEELFKAVVHAHSILT 65
Query: 117 DSALKKGYDAELRKKEAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAVLSD 176
D K YD +L+ K T+ + H F+ + P +S
Sbjct: 66 DEDQKLRYDRDLKIKGLHTY----QPKKNCHIFKTKAKESQGASPTLGQSEAYHRQNKPY 121
Query: 177 GSSDEGEKVGVRRNEKREVVLGVAAPNGVKG-----KVGGEKGLKGKVGDGDGVGDVGGG 231
G VG + + + + +K +K K D D + G
Sbjct: 122 EQQPYGFGVGKKMTSSSKSKVPIFKSFNLKSYQRNHYYSSKKERKHGSPDIDSLFHETNG 181
Query: 232 GSKKRMRSVGDVFERSKPKGI--ITGEETMTLAEF------------------QSQVKRK 271
SK RM G + S+ + I I G+ T + + K++
Sbjct: 182 ASKVRMTDAGKMDTNSQFQEIWEILGKNAYTHKSYSEDPNSCLGSALSDHEEEEEAGKQQ 241
Query: 272 VQQEKVKDKEEEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVKSKRGSEVGEER 331
QQ++ + +++ +G T K SS E + K P +E+ V + E GEE
Sbjct: 242 QQQQQQQQQQQHYGMTSKSSSPDE----------EKKNNKEPKRESRVSPE---ENGEEE 288
Query: 332 SLKKNVKL 339
+ K KL
Sbjct: 289 TGHKQFKL 296
>DJBC_MOUSE (Q9QYI4) DnaJ homolog subfamily B member 12 (mDJ10)
Length = 376
Score = 59.3 bits (142), Expect = 3e-08
Identities = 32/79 (40%), Positives = 48/79 (60%), Gaps = 4/79 (5%)
Query: 48 SEMVTSLT-ILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFK 106
SE V ++ + D+Y +LGV A+ ++K Y+KL+L HPDKN+ A + EAFK
Sbjct: 97 SEQVAAVKRVKQCKDYYEILGVSRSASDEDLKKAYRKLALKFHPDKNH---APGATEAFK 153
Query: 107 LVGEAFRVLSDSALKKGYD 125
+G A+ VLS+ +K YD
Sbjct: 154 AIGTAYAVLSNPEKRKQYD 172
>DJBB_MOUSE (Q99KV1) DnaJ homolog subfamily B member 11 precursor
Length = 358
Score = 59.3 bits (142), Expect = 3e-08
Identities = 30/71 (42%), Positives = 46/71 (64%), Gaps = 2/71 (2%)
Query: 55 TILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRV 114
T+++ D+Y +LGV A+ ++K Y+KL+L LHPD+N A +E F+ +G A+ V
Sbjct: 19 TVIAGRDFYKILGVPRSASIKDIKKAYRKLALQLHPDRNPDDPQA--QEKFQDLGAAYEV 76
Query: 115 LSDSALKKGYD 125
LSDS +K YD
Sbjct: 77 LSDSEKRKQYD 87
>DNJH_SYNY3 (P50027) DnAJ-like protein slr0093
Length = 332
Score = 58.5 bits (140), Expect = 5e-08
Identities = 30/65 (46%), Positives = 42/65 (64%), Gaps = 2/65 (3%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y +LGV A+ ++KQ++KL+L HPDKN AA EE FK + EA+ VLSD
Sbjct: 8 DYYQILGVTKTASEAEIKKQFRKLALKYHPDKNPGDKAA--EEKFKEISEAYEVLSDPEK 65
Query: 121 KKGYD 125
++ YD
Sbjct: 66 RQKYD 70
>DNAJ_CLOAB (P30725) Chaperone protein dnaJ
Length = 374
Score = 58.5 bits (140), Expect = 5e-08
Identities = 29/65 (44%), Positives = 43/65 (65%), Gaps = 2/65 (3%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y VLG+E A+ + ++K ++KL++ HPDKN + A EE FK + EA++VLSD
Sbjct: 5 DYYEVLGLEKGASDDEIKKAFRKLAIKYHPDKNRGNKEA--EEKFKEINEAYQVLSDPDK 62
Query: 121 KKGYD 125
K YD
Sbjct: 63 KANYD 67
>DNAJ_CHLPN (Q9Z9E9) Chaperone protein dnaJ
Length = 392
Score = 58.5 bits (140), Expect = 5e-08
Identities = 28/65 (43%), Positives = 43/65 (66%), Gaps = 2/65 (3%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y++LG+ A++ ++K Y+KL++ HPDKN AA+E+ FK V EA+ VLSD
Sbjct: 2 DYYSILGISKTASAEEIKKAYRKLAVKYHPDKNPGD--AAAEKRFKEVSEAYEVLSDPQK 59
Query: 121 KKGYD 125
+ YD
Sbjct: 60 RDSYD 64
>DJB8_HUMAN (Q8NHS0) DnaJ homolog subfamily B member 8
Length = 232
Score = 58.5 bits (140), Expect = 5e-08
Identities = 31/65 (47%), Positives = 43/65 (65%), Gaps = 1/65 (1%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
++Y VLGV+ A+ ++K Y+KL+L HPDKN + A E+ FKLV EA+ VLSDS
Sbjct: 3 NYYEVLGVQASASPEDIKKAYRKLALRWHPDKNPDNKEEA-EKKFKLVSEAYEVLSDSKK 61
Query: 121 KKGYD 125
+ YD
Sbjct: 62 RSLYD 66
>SCJ1_YEAST (P25303) DnaJ-related protein SCJ1
Length = 404
Score = 57.8 bits (138), Expect = 8e-08
Identities = 41/104 (39%), Positives = 57/104 (54%), Gaps = 13/104 (12%)
Query: 28 KSALKYAKRAQRLFPELEGISEMVTSLTILS---ATDWYTVLGVEPFANSNAVRKQYKKL 84
KS AKR++ + P+L ++ SL +L A D+Y +L ++ A ++ Y++L
Sbjct: 16 KSVHWLAKRSRTMIPKL--YIHLILSLLLLPLILAQDYYAILEIDKDATEKEIKSAYRQL 73
Query: 85 SLLLHPDKNNTHVAAASEEA---FKLVGEAFRVLSDSALKKGYD 125
S HPDKN A SEEA F VGEA+ VLSD KK YD
Sbjct: 74 SKKYHPDKN-----AGSEEAHQKFIEVGEAYDVLSDPEKKKIYD 112
>DJC7_MOUSE (Q9QYI3) DnaJ homolog subfamily C member 7
(Tetratricopeptide repeat protein 2) (TPR repeat protein
2) (MDj11)
Length = 494
Score = 57.4 bits (137), Expect = 1e-07
Identities = 28/69 (40%), Positives = 44/69 (63%), Gaps = 3/69 (4%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKN---NTHVAAASEEAFKLVGEAFRVLSD 117
D+Y +LGV+ A+ + ++K Y+K +L+ HPD++ + V E+ FK VGEAF +LSD
Sbjct: 381 DYYKILGVDKNASEDEIKKAYRKRALMHHPDRHSGASAEVQKEEEKKFKEVGEAFTILSD 440
Query: 118 SALKKGYDA 126
K YD+
Sbjct: 441 PKKKTRYDS 449
>DJC7_HUMAN (Q99615) DnaJ homolog subfamily C member 7
(Tetratricopeptide repeat protein 2) (TPR repeat protein
2)
Length = 494
Score = 57.4 bits (137), Expect = 1e-07
Identities = 28/69 (40%), Positives = 44/69 (63%), Gaps = 3/69 (4%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKN---NTHVAAASEEAFKLVGEAFRVLSD 117
D+Y +LGV+ A+ + ++K Y+K +L+ HPD++ + V E+ FK VGEAF +LSD
Sbjct: 381 DYYKILGVDKNASEDEIKKAYRKRALMHHPDRHSGASAEVQKEEEKKFKEVGEAFTILSD 440
Query: 118 SALKKGYDA 126
K YD+
Sbjct: 441 PKKKTRYDS 449
>DJBB_HUMAN (Q9UBS4) DnaJ homolog subfamily B member 11 precursor
(ER-associated dnaJ protein 3) (ErJ3) (ER-associated
Hsp40 co-chaperone) (hDj9) (PWP1-interacting protein 4)
(UNQ537/PRO1080)
Length = 358
Score = 57.4 bits (137), Expect = 1e-07
Identities = 29/70 (41%), Positives = 45/70 (63%), Gaps = 2/70 (2%)
Query: 56 ILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVL 115
+++ D+Y +LGV A+ ++K Y+KL+L LHPD+N A +E F+ +G A+ VL
Sbjct: 20 VIAGRDFYKILGVPRSASIKDIKKAYRKLALQLHPDRNPDDPQA--QEKFQDLGAAYEVL 77
Query: 116 SDSALKKGYD 125
SDS +K YD
Sbjct: 78 SDSEKRKQYD 87
>DNAJ_NITEU (O06431) Chaperone protein dnaJ
Length = 369
Score = 57.0 bits (136), Expect = 1e-07
Identities = 29/69 (42%), Positives = 43/69 (62%), Gaps = 2/69 (2%)
Query: 57 LSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLS 116
+S +D+Y VLGV A+ N ++K Y+KL++ HPD+N A EE FK + EA+ +LS
Sbjct: 1 MSQSDYYEVLGVGRDADENELKKAYRKLAMKYHPDRNAGDTKA--EERFKNIKEAYEILS 58
Query: 117 DSALKKGYD 125
D + YD
Sbjct: 59 DPNKRAAYD 67
>DJBC_HUMAN (Q9NXW2) DnaJ homolog subfamily B member 12
Length = 375
Score = 57.0 bits (136), Expect = 1e-07
Identities = 28/65 (43%), Positives = 42/65 (64%), Gaps = 3/65 (4%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y +LGV A+ ++K Y++L+L HPDKN+ A + EAFK +G A+ VLS+
Sbjct: 110 DYYEILGVSRGASDEDLKKAYRRLALKFHPDKNH---APGATEAFKAIGTAYAVLSNPEK 166
Query: 121 KKGYD 125
+K YD
Sbjct: 167 RKQYD 171
>DJB4_HUMAN (Q9UDY4) DnaJ homolog subfamily B member 4 (Heat shock
40 kDa protein 1 homolog) (Heat shock protein 40
homolog) (HSP40 homolog)
Length = 337
Score = 57.0 bits (136), Expect = 1e-07
Identities = 29/65 (44%), Positives = 41/65 (62%), Gaps = 3/65 (4%)
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y +LG+E A+ ++K Y+K +L HPDKN + A EE FK V EA+ VLSD
Sbjct: 4 DYYCILGIEKGASDEDIKKAYRKQALKFHPDKNKSPQA---EEKFKEVAEAYEVLSDPKK 60
Query: 121 KKGYD 125
++ YD
Sbjct: 61 REIYD 65
>DJA4_HUMAN (Q8WW22) DnaJ homolog subfamily A member 4
Length = 397
Score = 57.0 bits (136), Expect = 1e-07
Identities = 28/70 (40%), Positives = 42/70 (60%), Gaps = 5/70 (7%)
Query: 56 ILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVL 115
++ T +Y +LGV+P A+ ++K Y+KL+L HPDKN E FKL+ +A+ VL
Sbjct: 1 MVKETQYYDILGVKPSASPEEIKKAYRKLALKYHPDKN-----PDEGEKFKLISQAYEVL 55
Query: 116 SDSALKKGYD 125
SD + YD
Sbjct: 56 SDPKKRDVYD 65
>YAY1_SCHPO (Q10209) Hypothetical J-domain protein C4H3.01 in
chromosome I
Length = 392
Score = 56.6 bits (135), Expect = 2e-07
Identities = 28/75 (37%), Positives = 46/75 (61%), Gaps = 1/75 (1%)
Query: 60 TDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
T++Y +LG+ A + ++K Y+KL++ HPDKN ASE+ F+ + EA++VL D
Sbjct: 7 TEYYDLLGISTDATAVDIKKAYRKLAVKYHPDKNPDDPQGASEK-FQKISEAYQVLGDEK 65
Query: 120 LKKGYDAELRKKEAP 134
L+ YD ++K P
Sbjct: 66 LRSQYDQFGKEKAVP 80
>DNAJ_VIBCH (O34242) Chaperone protein dnaJ
Length = 381
Score = 56.6 bits (135), Expect = 2e-07
Identities = 29/69 (42%), Positives = 43/69 (62%), Gaps = 2/69 (2%)
Query: 57 LSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLS 116
+S D+Y VLGV A+ ++K YK+L++ HPD+N+ AA E FK V EA+ +L+
Sbjct: 1 MSKRDFYEVLGVGRDASERDIKKAYKRLAMKYHPDRNSGDAGAA--EKFKEVKEAYEILT 58
Query: 117 DSALKKGYD 125
D+ K YD
Sbjct: 59 DAQKKAAYD 67
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.134 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 73,935,025
Number of Sequences: 164201
Number of extensions: 3377701
Number of successful extensions: 11769
Number of sequences better than 10.0: 336
Number of HSP's better than 10.0 without gapping: 181
Number of HSP's successfully gapped in prelim test: 158
Number of HSP's that attempted gapping in prelim test: 10734
Number of HSP's gapped (non-prelim): 843
length of query: 606
length of database: 59,974,054
effective HSP length: 116
effective length of query: 490
effective length of database: 40,926,738
effective search space: 20054101620
effective search space used: 20054101620
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 69 (31.2 bits)
Lotus: description of TM0230.4


