

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0223.9
(324 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
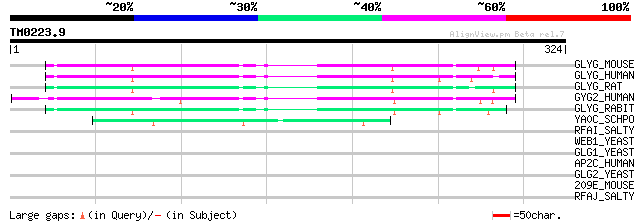
Score E
Sequences producing significant alignments: (bits) Value
GLYG_MOUSE (Q9R062) Glycogenin-1 (EC 2.4.1.186) 92 1e-18
GLYG_HUMAN (P46976) Glycogenin-1 (EC 2.4.1.186) 90 9e-18
GLYG_RAT (O08730) Glycogenin-1 (EC 2.4.1.186) 89 1e-17
GYG2_HUMAN (O15488) Glycogenin-2 (EC 2.4.1.186) (GN-2) (GN2) 87 7e-17
GLYG_RABIT (P13280) Glycogenin-1 (EC 2.4.1.186) 84 4e-16
YA0C_SCHPO (Q09680) Hypothetical protein C5H10.12c in chromosome I 49 1e-05
RFAI_SALTY (P19816) Lipopolysaccharide 1,3-galactosyltransferase... 35 0.33
WEB1_YEAST (P38968) WEB1 protein (Protein transport protein SEC31) 34 0.57
GLG1_YEAST (P36143) Glycogen synthesis initiator protein GLG1 34 0.57
AP2C_HUMAN (Q92754) Transcription factor AP-2 gamma (AP2-gamma) ... 32 2.2
GLG2_YEAST (P47011) Glycogen synthesis initiator protein GLG2 30 6.3
209E_MOUSE (Q91ZW7) CD209 antigen-like protein E (DC-SIGN relate... 30 6.3
RFAJ_SALTY (P19817) Lipopolysaccharide 1,2-glucosyltransferase (... 30 8.2
>GLYG_MOUSE (Q9R062) Glycogenin-1 (EC 2.4.1.186)
Length = 332
Score = 92.4 bits (228), Expect = 1e-18
Identities = 81/291 (27%), Positives = 121/291 (40%), Gaps = 53/291 (18%)
Query: 22 RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKIL--LSQGCIVRE 79
+AFVT L N Y KG + L L++ ++ +VV P V + RK+L + I+ +
Sbjct: 3 QAFVT-LTTNDAYAKGALVLGSSLKQHRTTRRMVVLTSPQVSDSMRKVLETVFDDVIMVD 61
Query: 80 IVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDNQF 139
++ + T I +KL W +YSK +++D D V NID LF+ +
Sbjct: 62 VLDSGDSAHLTLMKRPELGITLTKLHCWSLTQYSKCVFMDADTLVLSNIDDLFER--EEL 119
Query: 140 YAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHD 199
A D P W P FN+G FVY+P++ETY+
Sbjct: 120 SAAPD----PGW----------------------------PDCFNSGVFVYQPSIETYNQ 147
Query: 200 LLKTCEATTPTSFAEQDFLNMYF-----RDKYKPIPNVYNLVLAMLWRH-PENVQLDK-V 252
LL +Q LN YF D K +P VYNL ++ + P K
Sbjct: 148 LLHLASEQGSFDGGDQGLLNTYFSGWATTDITKHLPFVYNLSSISIYSYLPAFKAFGKNA 207
Query: 253 KVVHYCAFGSKPWRFTGEEE----NMDRKDIKM----LVKKWWDIYEDETL 295
KVVH+ +KPW +T + N D +D + + WWD + L
Sbjct: 208 KVVHFLG-RTKPWNYTYNPQTKSVNCDSQDPTVSHPEFLNLWWDTFTTNVL 257
>GLYG_HUMAN (P46976) Glycogenin-1 (EC 2.4.1.186)
Length = 349
Score = 89.7 bits (221), Expect = 9e-18
Identities = 78/294 (26%), Positives = 120/294 (40%), Gaps = 59/294 (20%)
Query: 22 RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKIL--LSQGCIVRE 79
+AFVT L N Y KG + L L++ ++ LVV P V + RK+L + I+ +
Sbjct: 3 QAFVT-LTTNDAYAKGALVLGSSLKQHRTTRRLVVLATPQVSDSMRKVLETVFDEVIMVD 61
Query: 80 IVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDNQF 139
++ + T + +KL W +YSK +++D D V NID LFD +
Sbjct: 62 VLDSGDSAHLTLMKRPELGVTLTKLHCWSLTQYSKCVFMDADTLVLANIDDLFDR--EEL 119
Query: 140 YAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHD 199
A D P W P FN+G FVY+P++ETY+
Sbjct: 120 SAAPD----PGW----------------------------PDCFNSGVFVYQPSVETYNQ 147
Query: 200 LLKTCEATTPTSFAEQDFLNMYF-----RDKYKPIPNVYNLVLAMLWRHPENVQL--DKV 252
LL +Q LN +F D K +P +YNL ++ + ++
Sbjct: 148 LLHLASEQGSFDGGDQGILNTFFSSWATTDIRKHLPFIYNLSSISIYSYLPAFKVFGASA 207
Query: 253 KVVHYCAFGSKPWRFT-----------GEEENMDRKDIKMLVKKWWDIYEDETL 295
KVVH+ KPW +T + NM + +L WW+I+ L
Sbjct: 208 KVVHFLG-RVKPWNYTYDPKTKSVKSEAHDPNMTHPEFLIL---WWNIFTTNVL 257
>GLYG_RAT (O08730) Glycogenin-1 (EC 2.4.1.186)
Length = 332
Score = 89.0 bits (219), Expect = 1e-17
Identities = 81/294 (27%), Positives = 118/294 (39%), Gaps = 59/294 (20%)
Query: 22 RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKIL--LSQGCIVRE 79
+AFVT L N Y KG + L L++ ++ VV P V + RK+L + I+ +
Sbjct: 3 QAFVT-LTTNDAYAKGALVLGSSLKQHRTTRRTVVLASPQVSDSMRKVLETVFDEVIMVD 61
Query: 80 IVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDNQF 139
++ + T I +KL W +YSK +++D D V NID LF+ +
Sbjct: 62 VLDSGDSAHLTLMKRPELGITLTKLHCWSLTQYSKCVFMDADTLVLSNIDDLFER--EEL 119
Query: 140 YAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHD 199
A D P W P FN+G FVY+P++ETY+
Sbjct: 120 SAAPD----PGW----------------------------PDCFNSGVFVYQPSIETYNQ 147
Query: 200 LLKTCEATTPTSFAEQDFLNMYF-----RDKYKPIPNVYNLVLAMLWRH-PENVQLDK-V 252
LL +Q LN YF D K +P VYNL ++ + P K
Sbjct: 148 LLHLASEQGSFDGGDQGLLNTYFSGWATTDITKHLPFVYNLSSLSIYSYLPAFKAFGKNA 207
Query: 253 KVVHYCAFGSKPWRFTGEEENMDRKDIKM-----------LVKKWWDIYEDETL 295
KVVH+ +KPW +T N K +K + WWD + L
Sbjct: 208 KVVHFLG-RTKPWNYT---YNPQTKSVKCESQDPIVSHPEFLNLWWDTFTTNVL 257
>GYG2_HUMAN (O15488) Glycogenin-2 (EC 2.4.1.186) (GN-2) (GN2)
Length = 501
Score = 86.7 bits (213), Expect = 7e-17
Identities = 84/316 (26%), Positives = 132/316 (41%), Gaps = 67/316 (21%)
Query: 2 APDLTTAATPITTTEQPLATRAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPD 61
+P + + +T T+Q AFVT LA N Y +G + L + LR+ + LVV I P
Sbjct: 22 SPTSASQSAGMTVTDQ-----AFVT-LATNDIYCQGALVLGQSLRRHRLTRKLVVLITPQ 75
Query: 62 VPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYV------INYSKLRIWEFVEYSKM 115
V R IL V E+ + + + H ++ + +KL W YSK
Sbjct: 76 VSSLLRVILSKVFDEVIEVNLI----DSADYIHLAFLKRPELGLTLTKLHCWTLTHYSKC 131
Query: 116 IYLDGDIQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNF 175
++LD D V N+D LFD +F A D P W
Sbjct: 132 VFLDADTLVLSNVDELFDR--GEFSAAPD----PGW------------------------ 161
Query: 176 GPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMYFR-----DKYKPIP 230
P FN+G FV++P+L T+ LL+ A+Q LN +FR D +K +P
Sbjct: 162 ----PDCFNSGVFVFQPSLHTHKLLLQHAMEHGSFDGADQGLLNSFFRNWSTTDIHKHLP 217
Query: 231 NVYNLVLAMLWRH-PENVQL-DKVKVVHYCAFGSKPWRFTGEEEN---MDRKDIK----- 280
+YNL ++ + P Q KVVH+ KPW + ++ +++ +
Sbjct: 218 FIYNLSSNTMYTYSPAFKQFGSSAKVVHFLG-SMKPWNYKYNPQSGSVLEQGSVSSSQHQ 276
Query: 281 -MLVKKWWDIYEDETL 295
+ WW +Y++ L
Sbjct: 277 AAFLHLWWTVYQNNVL 292
>GLYG_RABIT (P13280) Glycogenin-1 (EC 2.4.1.186)
Length = 332
Score = 84.3 bits (207), Expect = 4e-16
Identities = 74/286 (25%), Positives = 114/286 (38%), Gaps = 53/286 (18%)
Query: 22 RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKIL--LSQGCIVRE 79
+AFVT L N Y KG + L L++ ++ L V P V + RK L + I +
Sbjct: 3 QAFVT-LTTNDAYAKGALVLGSSLKQHRTSRRLAVLTTPQVSDTMRKALEIVFDEVITVD 61
Query: 80 IVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDNQF 139
I+ + T + +KL W +YSK +++D D V NID LF+ +
Sbjct: 62 ILDSGDSAHLTLMKRPELGVTLTKLHCWSLTQYSKCVFMDADTLVLANIDDLFER--EEL 119
Query: 140 YAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHD 199
A D P W P FN+G FVY+P++ETY+
Sbjct: 120 SAAPD----PGW----------------------------PDCFNSGVFVYQPSVETYNQ 147
Query: 200 LLKTCEATTPTSFAEQDFLNMYFR-----DKYKPIPNVYNLVLAMLWRHPENVQL--DKV 252
LL +Q LN +F D K +P +YNL ++ + +
Sbjct: 148 LLHVASEQGSFDGGDQGLLNTFFNSWATTDIRKHLPFIYNLSSISIYSYLPAFKAFGANA 207
Query: 253 KVVHYCAFGSKPWRFTGEEENMDRKD--------IKMLVKKWWDIY 290
KVVH+ +KPW +T + + + + WWDI+
Sbjct: 208 KVVHFLG-QTKPWNYTYDTKTKSVRSEGHDPTMTHPQFLNVWWDIF 252
>YA0C_SCHPO (Q09680) Hypothetical protein C5H10.12c in chromosome I
Length = 371
Score = 49.3 bits (116), Expect = 1e-05
Identities = 51/204 (25%), Positives = 78/204 (38%), Gaps = 32/204 (15%)
Query: 49 QSIYPLVVAILPDVPEEHRKILLSQGCIVREIVP------VYPPKNQTQFAHAYYVINYS 102
+S YP+ + L V E + G V I P VY + +Q A Y +S
Sbjct: 103 KSKYPIHILALRGVDEWKIERFRKDGASVIVIDPIASSDIVYDTSSFSQEISARYEQMFS 162
Query: 103 KLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMP---------------------DNQFYA 141
KLRI+E +++ K+ +D DI + +NID +FD P D+ Y
Sbjct: 163 KLRIFEQIQFDKICVIDSDILIMKNIDDIFDTPYMYQQINTLNYTRLPSYTKPDDDTVYH 222
Query: 142 VKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLL 201
+ F E ++ Y Y + P YFNAG + P+ ++ +L
Sbjct: 223 FNEDFKEYGASRSEFY--PYLLAAVSDRGEHHSIPPEDTPYFNAGLMLIRPSELHFNRIL 280
Query: 202 KTCE---ATTPTSFAEQDFLNMYF 222
K EQ LN+ F
Sbjct: 281 KIGRFPYMYENAKMMEQSLLNLAF 304
>RFAI_SALTY (P19816) Lipopolysaccharide 1,3-galactosyltransferase
(EC 2.4.1.44) (Lipopolysaccharide
3-alpha-galactosyltransferase)
Length = 337
Score = 34.7 bits (78), Expect = 0.33
Identities = 43/168 (25%), Positives = 70/168 (41%), Gaps = 28/168 (16%)
Query: 114 KMIYLDGDIQVFENIDHLFDM--PDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDW 171
+++YLD DI +I L D+ +N+ AV E W + S+ P V
Sbjct: 125 RVLYLDADIACKGSIQELIDLNFAENEIAAVV-AEGELEWWTKRSVSLA----TPGLVSG 179
Query: 172 PSNFGPRPPLYFNAGFFVYEPNLETYH-------DLLKTCEATTPTSFAEQDFLNMYFRD 224
YFNAGF + L T ++LK E + +QD LN++ +
Sbjct: 180 ----------YFNAGFILINIPLWTAENISKKAIEMLKDPEVVQRITHLDQDVLNIFLVN 229
Query: 225 KYKPIPNVYNLVLAMLWRHPENV--QLDKVKV-VHYCAFGSKPWRFTG 269
K + + +N ++ + ++V +D V VHY +KPW G
Sbjct: 230 KARFVDKKFNTQFSLNYELKDSVINPVDAETVFVHYIG-PTKPWHSWG 276
>WEB1_YEAST (P38968) WEB1 protein (Protein transport protein SEC31)
Length = 1273
Score = 33.9 bits (76), Expect = 0.57
Identities = 46/185 (24%), Positives = 70/185 (36%), Gaps = 46/185 (24%)
Query: 158 SIKYCQQCPDKVDW--PSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTC---EATTPTSF 212
S + Q P + +W + F P P F F + ++T +L T E T
Sbjct: 289 SAEQLSQFPARGNWCFKTKFAPEAPDLFACASFDNKIEVQTLQNLTNTLDEQETETKQQE 348
Query: 213 AEQDFLNMYFRDKYKPIPNVYNLVLAMLWRHPENVQLDKVKVVHYCAFGSKPWRFTGE-- 270
+E DF N R++ K P+V++L + P H+ AFG K + T +
Sbjct: 349 SETDFWNNVSREESKEKPSVFHLQAPTWYGEPS-------PAAHW-AFGGKLVQITPDGK 400
Query: 271 ------------------EENMDRKDIK-----MLVKKWWDIYEDETLDWNYINPSYTEK 307
E + KD K LVK D+ E+ DWN + EK
Sbjct: 401 GVSITNPKISGLESNTTLSEALKTKDFKPLINQRLVKVIDDVNEE---DWNLL-----EK 452
Query: 308 VLIDG 312
+ +DG
Sbjct: 453 LSMDG 457
>GLG1_YEAST (P36143) Glycogen synthesis initiator protein GLG1
Length = 618
Score = 33.9 bits (76), Expect = 0.57
Identities = 37/208 (17%), Positives = 73/208 (34%), Gaps = 51/208 (24%)
Query: 103 KLRIWEFVEYSKMIYLDGD-IQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKY 161
K R+WE ++ +++YLD D + + + LFD+ Q + + W
Sbjct: 106 KARLWELTQFEQVLYLDSDTLPLNKEFLKLFDIMSKQTTSQVGAIADIGW---------- 155
Query: 162 CQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMY 221
P FN+G + P+ +T L T ++Q LN +
Sbjct: 156 ------------------PDMFNSGVMMLIPDADTASVLQNYIFENTSIDGSDQGILNQF 197
Query: 222 FRD--------------KYKPIPNVYNLVLAMLWRHPE---NVQLDKVKVVHYCAFGSKP 264
F ++ + YN+ + L N +K++H+ KP
Sbjct: 198 FNQNCCTDELVKDSFSREWVQLSFTYNVTIPNLGYQSSPAMNYFKPSIKLIHFIG-KHKP 256
Query: 265 WRFTGEEENMDRKDIKMLVKKWWDIYED 292
W ++ + + +W ++YE+
Sbjct: 257 WSLWSQKNFIKNE----YHDQWNEVYEE 280
>AP2C_HUMAN (Q92754) Transcription factor AP-2 gamma (AP2-gamma)
(Activating enhancer-binding protein 2 gamma)
(Transcription factor ERF-1)
Length = 450
Score = 32.0 bits (71), Expect = 2.2
Identities = 16/42 (38%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Query: 151 WRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEP 192
W+ T ++KY + C D+ D SN PR P +AG +Y P
Sbjct: 3 WKITD--NVKYEEDCEDRHDGSSNGNPRVPHLSSAGQHLYSP 42
>GLG2_YEAST (P47011) Glycogen synthesis initiator protein GLG2
Length = 380
Score = 30.4 bits (67), Expect = 6.3
Identities = 33/136 (24%), Positives = 54/136 (39%), Gaps = 33/136 (24%)
Query: 103 KLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDN-QFYAVKDCFCEPSWRHTKQYSIKY 161
K R+WE V++ ++++LD D +P N +F+ + + E + ++ I
Sbjct: 104 KARLWELVQFDQVLFLDAD-----------TLPLNKEFFEILRLYPEQT-----RFQI-- 145
Query: 162 CQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMY 221
PD + WP FN G + P+L+ L T A+Q N +
Sbjct: 146 -AAVPD-IGWPD--------MFNTGVLLLIPDLDMATSLQDFLIKTVSIDGADQGIFNQF 195
Query: 222 FRDKYKPIPNVYNLVL 237
F PI N VL
Sbjct: 196 F----NPICNYSKEVL 207
>209E_MOUSE (Q91ZW7) CD209 antigen-like protein E (DC-SIGN related
protein 4) (DC-SIGNR4)
Length = 208
Score = 30.4 bits (67), Expect = 6.3
Identities = 21/77 (27%), Positives = 30/77 (38%), Gaps = 6/77 (7%)
Query: 148 EPSWRHTKQYSI-KYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTC-- 204
E W HTKQ + K Q +VD P +FN + + + +HD + C
Sbjct: 48 EIQWEHTKQEKMYKDLSQLKSEVDRLCRLCPWDWTFFNGNCYFFSKSQRDWHDSMTACKE 107
Query: 205 ---EATTPTSFAEQDFL 218
+ S EQ FL
Sbjct: 108 MGAQLVIIKSHEEQSFL 124
>RFAJ_SALTY (P19817) Lipopolysaccharide 1,2-glucosyltransferase (EC
2.4.1.58)
Length = 336
Score = 30.0 bits (66), Expect = 8.2
Identities = 25/99 (25%), Positives = 42/99 (42%), Gaps = 18/99 (18%)
Query: 182 YFNAGFFVYEPNLETYHD--------LLKTCEATTPTSFAEQDFLNMYFRDKYKPIPNVY 233
YFN+G V NL+ + + LL + + +QD LN+ +DK +P Y
Sbjct: 175 YFNSG--VVFVNLKLWKENALTKKAFLLLAGKEADSFKYPDQDVLNILLQDKVIFLPRPY 232
Query: 234 NLVLAM-------LWRHPENVQLDKVKVVHYCAFGSKPW 265
N + + + N+ D ++HY +KPW
Sbjct: 233 NTIYTIKSELKDKSHKKYSNIINDNTILIHYTG-ATKPW 270
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,304,982
Number of Sequences: 164201
Number of extensions: 1907058
Number of successful extensions: 3875
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 3854
Number of HSP's gapped (non-prelim): 20
length of query: 324
length of database: 59,974,054
effective HSP length: 110
effective length of query: 214
effective length of database: 41,911,944
effective search space: 8969156016
effective search space used: 8969156016
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0223.9


