

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.5
(471 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
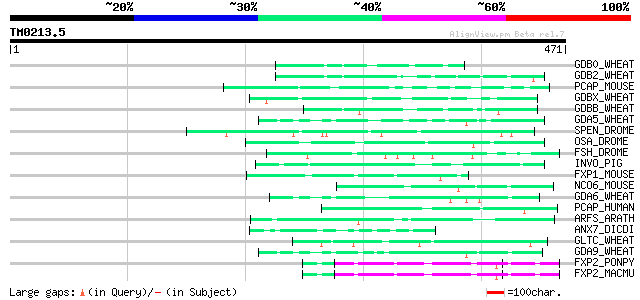
Score E
Sequences producing significant alignments: (bits) Value
GDB0_WHEAT (P08079) Gamma-gliadin precursor (Fragment) 57 1e-07
GDB2_WHEAT (P08453) Gamma-gliadin precursor 55 5e-07
PCAP_MOUSE (Q924H2) Positive cofactor 2 glutamine/Q-rich-associa... 52 3e-06
GDBX_WHEAT (P21292) Gamma-gliadin precursor 52 4e-06
GDBB_WHEAT (P06659) Gamma-gliadin B precursor 52 4e-06
GDA5_WHEAT (P04725) Alpha/beta-gliadin A-V precursor (Prolamin) 52 4e-06
SPEN_DROME (Q8SX83) Split ends protein 50 1e-05
OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein) 49 2e-05
FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chor... 49 2e-05
INVO_PIG (P18175) Involucrin 48 5e-05
FXP1_MOUSE (P58462) Forkhead box protein P1 (Forkhead-related tr... 48 5e-05
NCO6_MOUSE (Q9JL19) Nuclear receptor coactivator 6 (Amplified in... 48 6e-05
GDA6_WHEAT (P04726) Alpha/beta-gliadin clone PW1215 precursor (P... 47 8e-05
PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associa... 47 1e-04
ARFS_ARATH (Q8RYC8) Auxin response factor 19 (Auxin-responsive p... 47 1e-04
ANX7_DICDI (P24639) Annexin A7 (Annexin VII) (Synexin) 47 1e-04
GLTC_WHEAT (P16315) Glutenin, low molecular weight subunit PTDUC... 47 1e-04
GDA9_WHEAT (P18573) Alpha/beta-gliadin MM1 precursor (Prolamin) 47 1e-04
FXP2_PONPY (Q8MJ98) Forkhead box protein P2 47 1e-04
FXP2_MACMU (Q8MJ97) Forkhead box protein P2 47 1e-04
>GDB0_WHEAT (P08079) Gamma-gliadin precursor (Fragment)
Length = 251
Score = 56.6 bits (135), Expect = 1e-07
Identities = 55/161 (34%), Positives = 62/161 (38%), Gaps = 18/161 (11%)
Query: 226 VQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSF 285
+Q+D Q QQ V P Q Q P FPHQ PQQ P QQ F
Sbjct: 21 MQVDPSSQVQWPQQQPVPQPHQPFSQQPQQTFPQPQQTFPHQ--PQQQFPQPQQPQQ-QF 77
Query: 286 QQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQL 345
Q P P Q YPQQ Q F Q P FP PQQ + P Q
Sbjct: 78 LQPQQPFPQQPQQPYPQQ--------PQQPFPQTQQPQQLFPQSQQPQQQFSQP----QQ 125
Query: 346 SFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPMQQ 386
F Q P +FP QQ IQ S QQ + P + +QQ
Sbjct: 126 QFPQPQQPQQSFPQQQPPFIQPS---LQQQVNPCKNFLLQQ 163
Score = 38.5 bits (88), Expect = 0.038
Identities = 52/196 (26%), Positives = 69/196 (34%), Gaps = 33/196 (16%)
Query: 206 QNQVPQLNAPMQE-LSPQQNGVQL---------DAPMQQVSSQQNQVDALDFPMQQLSFQ 255
Q Q PQ P Q+ L PQQ Q P Q Q P QQ S
Sbjct: 64 QQQFPQPQQPQQQFLQPQQPFPQQPQQPYPQQPQQPFPQTQQPQQLFPQSQQPQQQFSQP 123
Query: 256 QNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQ--------------VYP 301
Q P P Q FPQQ P +Q S QQ +P NF Q ++P
Sbjct: 124 QQQFPQPQQPQQSFPQQQ---PPFIQP-SLQQQVNPCKNFLLQQCKPVSLVSSLWSMIWP 179
Query: 302 QQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQ 361
Q +C ++Q QQ P Q + +QQ +Q + P+ Q
Sbjct: 180 QSDCQV---MRQQCCQQLAQIPQQL--QCAAIHTIIHSIIMQQEQQEQQQGMHILLPLYQ 234
Query: 362 LQEIQQSGLASQQHMV 377
Q++ Q L Q ++
Sbjct: 235 QQQVGQGTLVQGQGII 250
Score = 31.2 bits (69), Expect = 6.0
Identities = 35/127 (27%), Positives = 44/127 (34%), Gaps = 25/127 (19%)
Query: 323 PLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVP--PL 380
P+ PHQ + QQ Q +F Q P FP Q Q+ Q QQ + P P
Sbjct: 36 PVPQPHQPFSQQ--------PQQTFPQ---PQQTFPHQPQQQFPQPQQPQQQFLQPQQPF 84
Query: 381 DAPMQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVP--PL 438
QQ +Q Q + P P QQ + QQ P P P
Sbjct: 85 PQQPQQPYPQQPQQPFPQTQQPQQLFPQSQQPQQQFSQ----------PQQQFPQPQQPQ 134
Query: 439 EVMPMQQ 445
+ P QQ
Sbjct: 135 QSFPQQQ 141
>GDB2_WHEAT (P08453) Gamma-gliadin precursor
Length = 327
Score = 54.7 bits (130), Expect = 5e-07
Identities = 75/234 (32%), Positives = 90/234 (38%), Gaps = 29/234 (12%)
Query: 226 VQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSF 285
+Q+D Q QQ V L P+ Q Q P FPHQ PQQ P QQ F
Sbjct: 21 IQVDPSGQVQWLQQQLVPQLQQPLSQQPQQTFPQPQQTFPHQ--PQQQVPQPQQPQQ-PF 77
Query: 286 QQNGDPPLNFPHQVYPQ-QNCATPLNVQ-QLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQ 343
Q P P Q +PQ Q P Q Q F Q P FP Q PQQ
Sbjct: 78 LQPQQPFPQQPQQPFPQTQQPQQPFPQQPQQPFPQTQQPQQPFPQQ--PQQ--------- 126
Query: 344 QLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQ 403
F Q P FP QLQ+ QQ QQ + P P Q ++Q L QQ
Sbjct: 127 --PFPQTQQPQQPFP--QLQQPQQPFPQPQQQLPQP-QQPQQSFPQQQRPFIQPSL--QQ 179
Query: 404 LMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLEVMPM---QQLQERPQEHQ 454
+ P + +QQ + S L S P +VM QQL + PQ+ Q
Sbjct: 180 QLNPCKNILLQQSKPASLV---SSLWSIIWPQSDCQVMRQQCCQQLAQIPQQLQ 230
Score = 36.6 bits (83), Expect = 0.14
Identities = 51/194 (26%), Positives = 66/194 (33%), Gaps = 33/194 (17%)
Query: 204 PQQNQ--VPQLNAPMQEL--SPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGD 259
PQQ Q PQ P Q PQQ Q P Q Q P QQL Q
Sbjct: 103 PQQPQQPFPQTQQPQQPFPQQPQQPFPQTQQPQQPFPQLQQPQQPFPQPQQQLPQPQQ-- 160
Query: 260 PPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQ--------------VYPQQNC 305
P Q FPQQ Q S QQ +P N Q ++PQ +C
Sbjct: 161 -----PQQSFPQQQ----RPFIQPSLQQQLNPCKNILLQQSKPASLVSSLWSIIWPQSDC 211
Query: 306 ATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEI 365
++Q QQ P + + QQ QQ G + P+ Q +++
Sbjct: 212 QV---MRQQCCQQLAQIPQQLQCAAIHSVVHSIIMQQQQQQQQQQG-IDIFLPLSQHEQV 267
Query: 366 QQSGLASQQHMVPP 379
Q L Q ++ P
Sbjct: 268 GQGSLVQGQGIIQP 281
>PCAP_MOUSE (Q924H2) Positive cofactor 2 glutamine/Q-rich-associated
protein (PC2 glutamine/Q-rich-associated protein)
(mPcqap)
Length = 792
Score = 52.4 bits (124), Expect = 3e-06
Identities = 76/278 (27%), Positives = 103/278 (36%), Gaps = 47/278 (16%)
Query: 182 STPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQ-QVSSQQN 240
ST + + + + + QQ Q Q A +Q+ QQ Q Q Q ++ Q
Sbjct: 145 STTTPQTQLQLQQVALQQQQQRQQQQQFQQQQAALQQQQQQQQQQQQQQQFQAQQNAMQQ 204
Query: 241 QVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVY 300
Q A+ QQL QQ + HQ QQ +QQ Q+ L Q
Sbjct: 205 QFQAV-VQQQQLQQQQQQQHLIKLHHQSQQQQ-------IQQQQLQRMAQLQLQQQQQQQ 256
Query: 301 PQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQ 360
QQ +QQ S QQ P P Q PQQ + QL Q+ PP
Sbjct: 257 QQQALQAQPPMQQPSMQQPQPP----PSQALPQQ-------LSQLHHPQHHQPP------ 299
Query: 361 QLQEIQQSGLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERK 420
+ QQS +A Q PP P Q Q +Q L P L A QQL+ +
Sbjct: 300 --PQAQQSPIAQNQ---PPQIPPQSQSQPLVS-------RAQALPGPMLYAAQQQLKFVR 347
Query: 421 HTLGDSVLASQQHPVPPLEVMPMQQLQERPQEHQAVNS 458
+ + QQ V P +QQ+Q + Q AV +
Sbjct: 348 -----APMVVQQPQVQP----QVQQVQPQVQPQAAVQA 376
Score = 39.7 bits (91), Expect = 0.017
Identities = 69/307 (22%), Positives = 102/307 (32%), Gaps = 37/307 (12%)
Query: 152 QWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTIDDTDLVPQQNQVPQ 211
Q Q ++F + Q A +L +++ +H + Q ++ Q
Sbjct: 187 QQQQQQQQFQAQQNAMQQQFQAVVQQQQLQQQQQQQHLIKLHHQSQQQQIQQQQLQRMAQ 246
Query: 212 LNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQ 271
L Q+ QQ +Q PMQQ S QQ Q QQLS Q+
Sbjct: 247 LQLQQQQQQQQQQALQAQPPMQQPSMQQPQPPPSQALPQQLS-------------QLHHP 293
Query: 272 QNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPL-------NVQQLSFQQNGDPPL 324
Q+ P QQ QN P + Q P + A L QQL F + P +
Sbjct: 294 QHHQPPPQAQQSPIAQNQPPQIPPQSQSQPLVSRAQALPGPMLYAAQQQLKFVR--APMV 351
Query: 325 NFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFP-MQQLQEIQQSGLASQQHMVPPLDA- 382
QV PQ P Q + Q + P +Q + E G + PP
Sbjct: 352 VQQPQVQPQVQQVQPQVQPQAAVQAAQSAQMVAPGVQMIAEALAQGGMHVRARFPPTSTM 411
Query: 383 -------------PMQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLA 429
P Q+ + T+ S QQ+ P P Q + + +S ++
Sbjct: 412 SAGPSSSISLGGQPTTQVSQSSLTMLSSPSPGQQVQTPQSMPPPPQPSPQPGSQPNSNVS 471
Query: 430 SQQHPVP 436
S P P
Sbjct: 472 SGPAPSP 478
>GDBX_WHEAT (P21292) Gamma-gliadin precursor
Length = 302
Score = 51.6 bits (122), Expect = 4e-06
Identities = 70/251 (27%), Positives = 86/251 (33%), Gaps = 39/251 (15%)
Query: 204 PQQNQVPQLNAPM----QELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGD 259
PQQ PQ P Q PQ + P Q Q P QQ Q
Sbjct: 32 PQQQPFPQPQQPFCQQPQRTIPQPHQTFHHQPQQTFPQPQQTYPHQ--PQQQFPQTQQPQ 89
Query: 260 PPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQN 319
P P Q FPQQ Q F Q P FP PQQ P Q F Q
Sbjct: 90 QPFPQPQQTFPQQPQLPFPQQPQQPFPQPQQPQQPFPQSQQPQQ----PFPQPQQQFPQP 145
Query: 320 GDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQ-QSGLASQQHMVP 378
P +FP Q P Q QQ +P NF +QQ + S L S ++P
Sbjct: 146 QQPQQSFPQQQQP---------AIQSFLQQQMNPCKNFLLQQCNHVSLVSSLVS--IILP 194
Query: 379 PLDAPMQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQ-ERKHTLGDSVLASQQHPVPP 437
D + Q Q QQL P QQLQ H++ S++ Q+
Sbjct: 195 RSDCQVMQQQ-----------CCQQLAQIP-----QQLQCAAIHSVAHSIIMQQEQQQGV 238
Query: 438 LEVMPMQQLQE 448
+ P+ QL +
Sbjct: 239 PILRPLFQLAQ 249
>GDBB_WHEAT (P06659) Gamma-gliadin B precursor
Length = 291
Score = 51.6 bits (122), Expect = 4e-06
Identities = 59/203 (29%), Positives = 79/203 (38%), Gaps = 24/203 (11%)
Query: 250 QQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNF--PHQVYPQQNCAT 307
QQ F Q P P Q+FPQ P QQ F Q P F P Q +PQQ
Sbjct: 33 QQQPFLQPHQPFSQQPQQIFPQPQQTFPHQPQQ-QFPQPQQPQQQFLQPRQPFPQQPQQP 91
Query: 308 PLNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQ 367
Q F Q P FP PQQ F Q P +FP QQ IQQ
Sbjct: 92 YPQQPQQPFPQTQQPQQPFPQSKQPQQ-----------PFPQPQQPQQSFPQQQPSLIQQ 140
Query: 368 SGLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQLMVPPLDAPM--QQLQERKHTLGD 425
S QQ + P + +QQ + S L S +++PP D + QQ ++ +
Sbjct: 141 S---LQQQLNPCKNFLLQQCKPVSLV---SSLWS--IILPPSDCQVMRQQCCQQLAQIPQ 192
Query: 426 SVLASQQHPVPPLEVMPMQQLQE 448
+ + H V +M +Q ++
Sbjct: 193 QLQCAAIHSVVHSIIMQQEQQEQ 215
Score = 34.3 bits (77), Expect = 0.71
Identities = 51/191 (26%), Positives = 71/191 (36%), Gaps = 26/191 (13%)
Query: 206 QNQVPQLNAPMQE-LSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNF 264
Q Q PQ P Q+ L P+Q Q QQ QQ Q Q F Q+ P F
Sbjct: 64 QQQFPQPQQPQQQFLQPRQPFPQQP---QQPYPQQPQQPFPQTQQPQQPFPQSKQPQQPF 120
Query: 265 PHQVFPQQNC--ATPLNVQQLSFQQNGDPPLNFPHQ--------------VYPQQNCATP 308
P PQQ+ P +QQ S QQ +P NF Q + P +C
Sbjct: 121 PQPQQPQQSFPQQQPSLIQQ-SLQQQLNPCKNFLLQQCKPVSLVSSLWSIILPPSDCQV- 178
Query: 309 LNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQS 368
++Q QQ P Q + + +QQ +Q + P+ Q Q++ Q
Sbjct: 179 --MRQQCCQQLAQIPQQL--QCAAIHSVVHSIIMQQEQQEQLQGVQILVPLSQQQQVGQG 234
Query: 369 GLASQQHMVPP 379
L Q ++ P
Sbjct: 235 ILVQGQGIIQP 245
>GDA5_WHEAT (P04725) Alpha/beta-gliadin A-V precursor (Prolamin)
Length = 319
Score = 51.6 bits (122), Expect = 4e-06
Identities = 73/255 (28%), Positives = 98/255 (37%), Gaps = 55/255 (21%)
Query: 212 LNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQ 271
+ P+ +L PQ Q P +QV Q Q FP QQ F P Q +PQ
Sbjct: 21 VRVPVPQLQPQNPSQQ--QPQEQVPLVQQQ----QFPGQQQQFP---------PQQPYPQ 65
Query: 272 QNCATPLNVQQLSFQQNGDP-PLNFPHQV-YPQQNCATPLNVQQLSFQQNGDPPLNFPHQ 329
P QQ Q P P FP Q+ YPQ SF P P
Sbjct: 66 PQ---PFPSQQPYLQLQPFPQPQPFPPQLPYPQPQ----------SFPPQQPYPQQQPQY 112
Query: 330 VYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPMQQ--- 386
+ PQQ P++ QQ QQ QQ Q+I Q L QQ ++P D +QQ
Sbjct: 113 LQPQQ----PISQQQAQQQQQQQQQ----QQQQQQILQQIL--QQQLIPCRDVVLQQHNI 162
Query: 387 -------LQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLE 439
LQ+ + L L QQL L P Q + H + +++ QQ +
Sbjct: 163 AHASSQVLQQSTYQLLQQ-LCCQQL----LQIPEQSQCQAIHNVAHAIIMHQQQQQQQEQ 217
Query: 440 VMPMQQLQERPQEHQ 454
+QQ Q++ Q+ Q
Sbjct: 218 KQQLQQQQQQQQQLQ 232
Score = 34.3 bits (77), Expect = 0.71
Identities = 54/193 (27%), Positives = 67/193 (33%), Gaps = 21/193 (10%)
Query: 204 PQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALD---FPMQQLSFQQNGDP 260
P Q PQ P Q +S QQ Q QQ QQ L P + + QQ+
Sbjct: 104 PYPQQQPQYLQPQQPISQQQAQQQQQQQQQQQQQQQILQQILQQQLIPCRDVVLQQH--- 160
Query: 261 PLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQ--QNCATPLNVQQLSFQQ 318
N H +T +QQL QQ L P Q Q N A + + Q QQ
Sbjct: 161 --NIAHASSQVLQQSTYQLLQQLCCQQ----LLQIPEQSQCQAIHNVAHAIIMHQQQQQQ 214
Query: 319 NGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNF--PMQQLQEIQQSGLASQQHM 376
Q+ QQ L QQ QQ ++F P QQ Q S SQ +
Sbjct: 215 QEQ-----KQQLQQQQQQQQQLQQQQQQQQQQPSSQVSFQQPQQQYPSSQVSFQPSQLNP 269
Query: 377 VPPLDAPMQQLQE 389
QQL +
Sbjct: 270 QAQGSVQPQQLPQ 282
Score = 31.2 bits (69), Expect = 6.0
Identities = 29/118 (24%), Positives = 43/118 (35%), Gaps = 19/118 (16%)
Query: 203 VPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPL 262
+P+Q+Q ++ + Q Q QQ+ QQ Q L QQ Q +
Sbjct: 189 IPEQSQCQAIHNVAHAIIMHQQQQQQQEQKQQLQQQQQQQQQLQQQQQQQQQQPSSQVSF 248
Query: 263 NFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQ-QNCATPLNVQQLSFQQN 319
P Q +P Q+SFQ P Q+ PQ Q P + Q + +N
Sbjct: 249 QQPQQQYPS---------SQVSFQ---------PSQLNPQAQGSVQPQQLPQFAEIRN 288
>SPEN_DROME (Q8SX83) Split ends protein
Length = 5560
Score = 50.4 bits (119), Expect = 1e-05
Identities = 84/342 (24%), Positives = 124/342 (35%), Gaps = 52/342 (15%)
Query: 151 GQWQTISREFLPSKTPTQIASHAQKYYLRLNS----------TPKRRKRASIHDLTIDDT 200
GQ + R + +P Q Q+ + + S +P R +S + T
Sbjct: 3669 GQLSPVGRPMVSQPSPQQQVQQTQQQHALITSPQSSNISPLASPTTRVLSSSNSPTTSKV 3728
Query: 201 DLV-PQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQ--------NQVDALDFPMQQ 251
+ P+ QVPQ +P Q + P+Q+++ Q ++V + P Q
Sbjct: 3729 NSYQPRNQQVPQQPSPKSVAEVQTTPQLMTIPLQKMTPIQVPHHPTIISKVVTVQ-PQQA 3787
Query: 252 LSFQQNGDPPLNF--PHQ---VFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCA 306
Q PPL PH+ + QN P + +++ Q+ F HQ Q+
Sbjct: 3788 TQSQVASSPPLGSLPPHKNVHLNAHQNQQQPQVIAKMTAHQHQQHMQQFMHQQMIQRQ-- 3845
Query: 307 TPLNVQQL--SFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQE 364
+ QQL QQ P + HQ + Q N Q L+ Q H + +Q Q
Sbjct: 3846 QHMQQQQLHGQSQQITSAPQHQMHQQHQAQQQQQHHNQQHLN--QQLHAQQHPTQKQHQA 3903
Query: 365 IQQSGLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQL-------- 416
QQ QQH QQ Q +Q L SQQ + A QQL
Sbjct: 3904 QQQFNQQIQQHQSQQQHQVQQQNQAQQQHLSQQQHQSQQQLNQQHQAQQQQLQQIQKLQQ 3963
Query: 417 -----QERKHTLG-------DSVLASQQHPVP-PLEVMPMQQ 445
Q++K G S+ ASQQH P +P QQ
Sbjct: 3964 MHGPQQQQKSPQGVGHLGGSTSIFASQQHNSQLPARGVPQQQ 4005
>OSA_DROME (Q8IN94) Trithorax group protein OSA (Eyelid protein)
Length = 2716
Score = 49.3 bits (116), Expect = 2e-05
Identities = 68/265 (25%), Positives = 88/265 (32%), Gaps = 30/265 (11%)
Query: 201 DLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVS-SQQNQVDALDFPMQQLSFQQNGD 259
D + P NA Q Q+G Q P S + QN A +
Sbjct: 1486 DFIKDSQPYPGYNARPQIYGAWQSGTQQYRPQYPSSPAPQNWGGAPP---------RGAA 1536
Query: 260 PPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQN 319
PP PH QQ Q Q G PP PQQ QQ +QQ
Sbjct: 1537 PPPGAPHGPPIQQPAGVAQWDQHRYPPQQGPPPP-------PQQQQQPQQQQQQPPYQQV 1589
Query: 320 GDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGL-ASQQHMVP 378
PP P Q PQ P Q G P L P Q+ + G+ A QQ
Sbjct: 1590 AGPPGQQPPQAPPQWAQMNPGQTAQSGIAPPGSP-LRPPSGPGQQNRMPGMPAQQQQSQQ 1648
Query: 379 PLDAPMQQLQERQH---------TLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLA 429
P Q+ H +G G+ +PP P Q + ++ +
Sbjct: 1649 QGGVPQPPPQQASHGGVPSPGLPQVGPGGMVKPPYAMPP--PPSQGVGQQVGQGPPGGMM 1706
Query: 430 SQQHPVPPLEVMPMQQLQERPQEHQ 454
SQ+ P P + M Q LQ++P HQ
Sbjct: 1707 SQKPPPMPGQAMQQQPLQQQPPSHQ 1731
Score = 36.2 bits (82), Expect = 0.19
Identities = 67/276 (24%), Positives = 89/276 (31%), Gaps = 37/276 (13%)
Query: 204 PQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLN 263
PQQ Q P Q+ P G + P QQ + L N PP
Sbjct: 173 PQQQQPHPQQQPPQQPGP---GGSPNRPPQQRYIPGQPPQGPTPTLNSLLQSSNPPPPPQ 229
Query: 264 --FPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGD 321
+ + PQQ A+ QQ G PP H P Q+ +P QQ +
Sbjct: 230 HRYANTYDPQQAAASAAAAAAAQQQQAGGPPPP-GHGPPPPQHQPSPYGGQQGGW---AP 285
Query: 322 PPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLN------FP----MQQLQEIQQSGLA 371
PP + Q+ P Q TP S Q+ +PP + +P QQ Q+ QQ A
Sbjct: 286 PPRPYSPQLGPSQQYRTPPPTNT-SRGQSPYPPAHGQNSGSYPSSPQQQQQQQQQQQQQA 344
Query: 372 SQQHMVPPLDAP-----MQQLQERQHTLGDSGLTSQQLMVPP------------LDAPMQ 414
QQ P P Q Q+ Q+ PP
Sbjct: 345 GQQPGGPVPGGPPPGTGQQPPQQNTPPTSQYSPYPQRYPTPPGLPAGGSNHRTAYSTHQY 404
Query: 415 QLQERKHTLGDSVLASQQHPVPPLEVMPMQQLQERP 450
R G S HP+PP + LQ++P
Sbjct: 405 PEPNRPWPGGSSPSPGSGHPLPPASPHHVPPLQQQP 440
>FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chorion
membrane protein)
Length = 2038
Score = 49.3 bits (116), Expect = 2e-05
Identities = 72/287 (25%), Positives = 93/287 (32%), Gaps = 68/287 (23%)
Query: 219 LSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQ----LSFQQNGDPPLNFPHQVFPQQNC 274
L ++A + ++S Q+ D P++Q L F QQ+
Sbjct: 1323 LGGSHGDAMVNASLASLASGLKQIPQFDDPVEQSLASLEFSAGSTGKSGLTDNFLMQQHL 1382
Query: 275 ATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQ---------------QN 319
P QQ QQ P F HQ QQ QQ N
Sbjct: 1383 MQPAGPQQQQQQQQQQP---FGHQQQQQQQQQQQQQQQQQHMDYVTELLSKGAENVGGMN 1439
Query: 320 GDPPLNFP-------HQVYPQQNCTTPLN--VQQLSFQQNGHPPLNF---------PMQQ 361
G+ LNF Q +PQQ N F G LN P Q
Sbjct: 1440 GNHLLNFNLDMAAAYQQKHPQQQQQQAHNNGFNVADFGMAGFDGLNMTAASFLDLEPSLQ 1499
Query: 362 LQEIQQSGLASQQHMVPPLDAPMQQLQERQ--HTLGDSGLTSQQLMVPPLDAPMQQLQER 419
Q++QQ L Q H QQ Q +Q H LT QQL QQ Q++
Sbjct: 1500 QQQMQQMQLQQQHHQQQQQQTHQQQQQHQQQHHQQQQQQLTQQQLQ--------QQQQQQ 1551
Query: 420 KHTLGDSVLASQQHPVPPLEVMPMQQLQERPQEHQAVNSRHFVWKPV 466
+ QQH +QQ Q + Q HQA N + KP+
Sbjct: 1552 QQ---------QQH---------LQQQQHQQQHHQAANKLLIIPKPI 1580
>INVO_PIG (P18175) Involucrin
Length = 347
Score = 48.1 bits (113), Expect = 5e-05
Identities = 61/247 (24%), Positives = 97/247 (38%), Gaps = 30/247 (12%)
Query: 209 VPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQV 268
VP+ Q PQQ + ++ QQ Q++QV L QQ Q++ + L+ Q
Sbjct: 70 VPEQECEPQPQEPQQQELHVE---QQQQQQESQVQELHVDQQQQQ-QESQEQELHVDQQQ 125
Query: 269 FPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNG-DPPLNFP 327
Q++ L+V Q Q++ L+ H Q++ L+V QQ + L+
Sbjct: 126 QQQESQEQELHVDQQQQQESQVQELHVGHHQQQQESQEQELHVDHHQQQQESQEQELHVD 185
Query: 328 HQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPMQQL 387
Q Q++ L+V Q QQ QE Q+ L H QQ
Sbjct: 186 QQQQQQESQEQELHVDQ--------------QQQQQESQEQELHVDHH---------QQQ 222
Query: 388 QERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLEVMPMQQLQ 447
QE Q QQ + + Q Q+++ + + QQ L+V +QQ Q
Sbjct: 223 QESQVQELHVDHQQQQQESQEQELHVDQHQQQQESQEQELHVDQQQ--QELQVQEVQQQQ 280
Query: 448 ERPQEHQ 454
++ QE Q
Sbjct: 281 QQQQEQQ 287
>FXP1_MOUSE (P58462) Forkhead box protein P1 (Forkhead-related
transcription factor 1)
Length = 705
Score = 48.1 bits (113), Expect = 5e-05
Identities = 58/194 (29%), Positives = 75/194 (37%), Gaps = 18/194 (9%)
Query: 202 LVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPP 261
L+ QQ Q Q Q+ QQ Q QQ QQ QV L P + + P
Sbjct: 68 LLLQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQVSGLKSPKRN-----DKQPA 122
Query: 262 LNFPHQV-FPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNG 320
L P V TP +QQ+ QQ P QV QQ A L Q F +
Sbjct: 123 LQVPVSVAMMTPQVITPQQMQQILQQQVLSPQ---QLQVLLQQQQALMLQQQLQEFYKKQ 179
Query: 321 DPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQE-----IQQSGLASQQH 375
L Q+ QQ+ QQ++ QQ MQQLQ+ +Q+ GL + Q
Sbjct: 180 QEQLQL--QLLQQQHAGKQPKEQQVATQQLAFQQQLLQMQQLQQQHLLSLQRQGLLTIQP 237
Query: 376 MVPPLDAPMQQLQE 389
P L P+Q L +
Sbjct: 238 GQPAL--PLQPLAQ 249
>NCO6_MOUSE (Q9JL19) Nuclear receptor coactivator 6 (Amplified in
breast cancer-3 protein) (Cancer-amplified
transcriptional coactivator ASC-2) (Activating signal
cointegrator-2) (ASC-2) (Peroxisome
proliferator-activated receptor-interacting protein) (PP
Length = 2067
Score = 47.8 bits (112), Expect = 6e-05
Identities = 57/189 (30%), Positives = 69/189 (36%), Gaps = 14/189 (7%)
Query: 278 LNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCT 337
L++QQ S PP + Q P P N QL QQ Q QQ
Sbjct: 224 LHIQQQSHPSGSLPPAHHSMQPVPVNRQMNPANFPQLQQQQQQQQQQQQQQQQQQQQQ-- 281
Query: 338 TPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPP----LDAPMQQLQERQHT 393
QQ QQ PL QQ Q I+ A Q VPP L + Q Q +
Sbjct: 282 ---QQQQQQQQQLQTRPLQQHQQQPQGIRPQFTAPTQVPVPPGWNQLPSGALQPPPAQGS 338
Query: 394 LGDSGLTSQQLMVPPLDAPMQ-QLQERKHTLGDSVLASQQHPVPPLEVMPMQQLQERPQE 452
LG + T+Q PL +PMQ QLQ R + + + HP PP Q Q
Sbjct: 339 LG-TMTTNQGWKKAPLPSPMQAQLQARPSL---ATVQTPSHPPPPYPFGSQQASQAHTNF 394
Query: 453 HQAVNSRHF 461
Q N F
Sbjct: 395 PQMSNPGQF 403
Score = 37.4 bits (85), Expect = 0.084
Identities = 67/289 (23%), Positives = 95/289 (32%), Gaps = 50/289 (17%)
Query: 210 PQLNAPMQELSPQQNGVQ-----LDAPMQQ-----------VSSQQNQVDALDFPMQQLS 253
PQ++ P Q +PQ G+Q + P+QQ SS Q A + Q
Sbjct: 395 PQMSNPGQFTAPQMKGLQGGPSRVPTPLQQPHLTNKSPASSPSSFQQGSPASSPTVNQTQ 454
Query: 254 FQQNGDPPLNFPHQVFPQQNCATPLN---VQQLSFQQN-----GDPPLNFPHQVYPQQNC 305
Q PP N P QQ ++P VQQ + N PP P ++P
Sbjct: 455 QQMGPRPPQNNPLSQGFQQPVSSPGRNPMVQQGNVPPNFMVMQQQPPNQGPQSLHPGLG- 513
Query: 306 ATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEI 365
P + +P NF P TP N L Q N Q +Q
Sbjct: 514 GMPKRLPPGFSAGQANP--NFMQGQVPSTTAATPGNSGALQLQAN---------QNVQHA 562
Query: 366 QQSGLASQQHMVPPLDAPMQQLQE---------RQHTLGDSG---LTSQQLMVPPLDAPM 413
G Q+ + P +Q G SG +T + P P
Sbjct: 563 GGQGAGPPQNQMQVSHGPPNMMQPSLMGIHGNINNQQAGSSGVPQVTLGNMQGQPQQGPP 622
Query: 414 QQLQERKHTL--GDSVLASQQHPVPPLEVMPMQQLQERPQEHQAVNSRH 460
QL + +A QQ + P M + + Q PQ VN+++
Sbjct: 623 SQLMGMHQQIVPSQGQMAQQQGTLNPQNPMILSRAQLMPQGQMMVNAQN 671
Score = 37.4 bits (85), Expect = 0.084
Identities = 57/271 (21%), Positives = 86/271 (31%), Gaps = 32/271 (11%)
Query: 203 VPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPL 262
VP Q+ N P + QQ Q QQ QQ Q +Q QQ+ P
Sbjct: 246 VPVNRQMNPANFPQLQQQQQQQQQQQQQQQQQ-QQQQQQQQQQQQQLQTRPLQQHQQQPQ 304
Query: 263 NFPHQVFPQQNCATPLNVQQLSFQQNGDPPLN------FPHQVYPQQNCATPLNVQ---- 312
Q P QL PP +Q + + +P+ Q
Sbjct: 305 GIRPQFTAPTQVPVPPGWNQLPSGALQPPPAQGSLGTMTTNQGWKKAPLPSPMQAQLQAR 364
Query: 313 --QLSFQQNGDPPLNFP---------HQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQ 361
+ Q PP +P H +PQ + Q+ Q G + P+QQ
Sbjct: 365 PSLATVQTPSHPPPPYPFGSQQASQAHTNFPQMSNPGQFTAPQMKGLQGGPSRVPTPLQQ 424
Query: 362 LQEIQQSGLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQ-LQERK 420
+S +S A + + Q +G PP + P+ Q Q+
Sbjct: 425 PHLTNKSPASSPSSFQQGSPASSPTVNQTQQQMGPR---------PPQNNPLSQGFQQPV 475
Query: 421 HTLGDSVLASQQHPVPPLEVMPMQQLQERPQ 451
+ G + + Q + P VM Q + PQ
Sbjct: 476 SSPGRNPMVQQGNVPPNFMVMQQQPPNQGPQ 506
Score = 32.3 bits (72), Expect = 2.7
Identities = 66/267 (24%), Positives = 93/267 (34%), Gaps = 55/267 (20%)
Query: 200 TDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGD 259
TD V N L+ M ++S QQ + P S Q N F +SF
Sbjct: 806 TDPVTTANNDVNLSQMMPDVSMQQASM---VPPHVQSMQGNSASGSHFSGHGVSF----- 857
Query: 260 PPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQN 319
N P P N Q+S QN P+N +PL V L +
Sbjct: 858 ---NAPFGGAP--------NGSQMSCGQNPGFPVN------KDVTLTSPLLVNLLQSDIS 900
Query: 320 -GDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQ---LQEIQQSGLASQQH 375
G +N N P + ++N H LN P + L+E+ Q L +Q
Sbjct: 901 AGHFGVNNKQN---NTNANKPKKKKPPRKKKNCHQDLNTPDNRPTGLEEVDQQSLPGEQG 957
Query: 376 M--------VP-----PLDAPMQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHT 422
+ +P P P Q +++R L Q + PP P QQ Q +
Sbjct: 958 INLDTTGPKLPDFSNRPPGYPTQPVEQRP--LPQMPPQLMQHVAPPPQPPQQQPQPQ--- 1012
Query: 423 LGDSVLASQQHPVPPLEVMPMQQLQER 449
L QQ P PP + QQ Q++
Sbjct: 1013 -----LPQQQQPPPPSQPQSQQQQQQQ 1034
>GDA6_WHEAT (P04726) Alpha/beta-gliadin clone PW1215 precursor
(Prolamin)
Length = 296
Score = 47.4 bits (111), Expect = 8e-05
Identities = 69/242 (28%), Positives = 84/242 (34%), Gaps = 51/242 (21%)
Query: 221 PQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATPLNV 280
PQ P QV Q Q FP QQ F P Q +PQ P
Sbjct: 28 PQPQNPSQPQPQGQVPLVQQQ----QFPGQQQQFP---------PQQPYPQPQ---PFPS 71
Query: 281 QQLSFQQNGDP-PLNFPHQV-YPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCTT 338
QQ Q P P FP Q+ YPQ PP P Q YPQ
Sbjct: 72 QQPYLQLQPFPQPQPFPPQLPYPQ-------------------PPPFSPQQPYPQPQPQY 112
Query: 339 PLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLAS--QQHMVPPLDAPMQQ---LQERQHT 393
P Q +S QQ QQ Q+ QQ L QQ ++P D +QQ R
Sbjct: 113 PQPQQPISQQQAQQQQQQQQQQQQQQQQQQILQQILQQQLIPCRDVVLQQHNIAHARSQV 172
Query: 394 LGDS------GLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLEVMPMQQLQ 447
L S L QQL P + Q + H + +L QQ P + +QQ Q
Sbjct: 173 LQQSTYQPLQQLCCQQLWQIPEQSRCQAIHNVVHAI---ILHQQQRQQQPSSQVSLQQPQ 229
Query: 448 ER 449
++
Sbjct: 230 QQ 231
Score = 34.7 bits (78), Expect = 0.54
Identities = 58/190 (30%), Positives = 67/190 (34%), Gaps = 26/190 (13%)
Query: 204 PQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQL------SFQQN 257
P PQ P Q +S QQ Q QQ QQ Q QQL QQ+
Sbjct: 104 PYPQPQPQYPQPQQPISQQQAQQQQQQQQQQQQQQQQQQILQQILQQQLIPCRDVVLQQH 163
Query: 258 GDPPLNFPH---QVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQL 314
N H QV QQ+ PL QQL QQ P Q A L+ QQ
Sbjct: 164 -----NIAHARSQVL-QQSTYQPL--QQLCCQQLWQIPEQSRCQAIHNVVHAIILHQQQR 215
Query: 315 SFQQNGDPPLNFPHQVYPQ-QNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQ 373
Q + L P Q YP Q P QQ Q P P Q +EI+ L +
Sbjct: 216 QQQPSSQVSLQQPQQQYPSGQGFFQP--SQQNPQAQGSVQPQQLP--QFEEIRNLALQTL 271
Query: 374 QHM----VPP 379
M +PP
Sbjct: 272 PRMCNVYIPP 281
>PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associated
protein (PC2 glutamine/Q-rich-associated protein)
(TPA-inducible gene-1) (TIG-1) (Activator-recruited
cofactor 105 kDa component) (ARC105) (CTG repeat protein
7a)
Length = 788
Score = 47.0 bits (110), Expect = 1e-04
Identities = 53/205 (25%), Positives = 66/205 (31%), Gaps = 14/205 (6%)
Query: 265 PHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGDPPL 324
P Q A QQ FQQ L Q QQ + Q FQ
Sbjct: 149 PQTQLQLQQVALQQQQQQQQFQQQQQAALQQQQQQQQQQQFQAQQSAMQQQFQAVVQQQQ 208
Query: 325 NFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPM 384
Q QQ+ + Q QQ QQLQ I Q L QQ
Sbjct: 209 QLQQQQQQQQHLIKLHHQNQQQIQQQ--------QQQLQRIAQLQLQQQQQQQQQQQQQQ 260
Query: 385 QQLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPV----PPLEV 440
QQ + Q + + QQ PP A QQLQ+ HT Q P P ++
Sbjct: 261 QQALQAQPPIQQPPM--QQPQPPPSQALPQQLQQMHHTQHHQPPPQPQQPPVAQNQPSQL 318
Query: 441 MPMQQLQERPQEHQAVNSRHFVWKP 465
P Q Q + QA+ + +P
Sbjct: 319 PPQSQTQPLVSQAQALPGQMLYTQP 343
Score = 41.6 bits (96), Expect = 0.004
Identities = 59/229 (25%), Positives = 77/229 (32%), Gaps = 33/229 (14%)
Query: 205 QQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNF 264
QQ Q A +Q+ QQ Q A Q S+ Q Q A+ QQL QQ L
Sbjct: 166 QQQFQQQQQAALQQQQQQQQQQQFQA---QQSAMQQQFQAVVQQQQQLQQQQQQQQHLIK 222
Query: 265 PHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGDPPL 324
H QQ + QQ Q+ L Q QQ QQ PP+
Sbjct: 223 LHHQNQQQ-----IQQQQQQLQRIAQLQLQQQQQQQQQQQ-------QQQQQALQAQPPI 270
Query: 325 NFP--HQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDA 382
P Q P + P +QQ+ Q+ PP P Q + Q +Q +PP
Sbjct: 271 QQPPMQQPQPPPSQALPQQLQQMHHTQHHQPP---PQPQQPPVAQ----NQPSQLPPQSQ 323
Query: 383 PMQQLQERQHTLGDSGLTSQQL---------MVPPLDAPMQQLQERKHT 422
+ + Q G T L PP+ +QQ Q T
Sbjct: 324 TQPLVSQAQALPGQMLYTQPPLKFVRAPMVVQQPPVQPQVQQQQTAVQT 372
Score = 40.4 bits (93), Expect = 0.010
Identities = 71/266 (26%), Positives = 86/266 (31%), Gaps = 49/266 (18%)
Query: 202 LVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPP 261
+ P V P +L QQ V L QQ QQ Q AL QQ QQ
Sbjct: 136 MAPHSMAVVSTATPQTQLQLQQ--VALQQQQQQQQFQQQQQAALQQQQQQQQQQQ----- 188
Query: 262 LNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGD 321
QQ A QQL QQ L H QQ + QQ Q+
Sbjct: 189 FQAQQSAMQQQFQAVVQQQQQLQQQQQQQQHLIKLHHQNQQQ-----IQQQQQQLQRIAQ 243
Query: 322 PPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLD 381
L Q QQ QQ PP+ P QQ PP
Sbjct: 244 LQLQQQQQQQQQQQ-------QQQQQALQAQPPIQQP------------PMQQPQPPPSQ 284
Query: 382 APMQQLQERQHTLGDSG---------LTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQ 432
A QQLQ+ HT +Q +PP + Q L + L +L +Q
Sbjct: 285 ALPQQLQQMHHTQHHQPPPQPQQPPVAQNQPSQLPP-QSQTQPLVSQAQALPGQMLYTQ- 342
Query: 433 HPVPPLEV----MPMQQLQERPQEHQ 454
PPL+ M +QQ +PQ Q
Sbjct: 343 ---PPLKFVRAPMVVQQPPVQPQVQQ 365
>ARFS_ARATH (Q8RYC8) Auxin response factor 19 (Auxin-responsive
protein IAA22)
Length = 1086
Score = 47.0 bits (110), Expect = 1e-04
Identities = 65/263 (24%), Positives = 94/263 (35%), Gaps = 34/263 (12%)
Query: 205 QQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNF 264
QQ Q Q Q+L QQ QL QQV Q + Q+S Q P
Sbjct: 498 QQQQQQQHQHQQQQLQQQQ---QLQMSQQQVQQQGIYNNGTIAVANQVSCQSPNQPTGFS 554
Query: 265 PHQVFPQQNCATPLNVQQLSFQQNGDPPLN----FPHQVYPQQNCATPLNVQQLSFQQNG 320
Q+ Q T + + G+ L+ F ++ QQ + Q L QQ
Sbjct: 555 QSQLQQQSMLPTGAKMTHQNINSMGNKGLSQMTSFAQEMQFQQQLEMHNSSQLLRNQQEQ 614
Query: 321 DPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPL 380
+ QQN + N QQL QQ P QLQ +Q+ QQ +PP+
Sbjct: 615 SSLHSL------QQNLSQ--NPQQLQMQQQSSKPSPSQQLQLQLLQKLQQQQQQQSIPPV 666
Query: 381 DAPMQ-QLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLE 439
+ +Q QL Q T QLQ+ + LA + P
Sbjct: 667 SSSLQPQLSALQQT------------------QSHQLQQLLSSQNQQPLAHGNNSFPAST 708
Query: 440 VMPMQQLQERPQEHQAVNSRHFV 462
M Q+Q PQ+ +++++ V
Sbjct: 709 FMQPPQIQVSPQQQGQMSNKNLV 731
>ANX7_DICDI (P24639) Annexin A7 (Annexin VII) (Synexin)
Length = 462
Score = 47.0 bits (110), Expect = 1e-04
Identities = 54/158 (34%), Positives = 59/158 (37%), Gaps = 35/158 (22%)
Query: 204 PQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLN 263
P Q PQ N+P P Q G AP Q QQ +P QQ Q G PP
Sbjct: 5 PNQGYPPQSNSPQ----PGQYG----APQQGYPPQQGYPPQQGYPPQQGYPPQQGYPP-- 54
Query: 264 FPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGDPP 323
Q +P Q P Q G PP Q YP Q P QQ Q G PP
Sbjct: 55 --QQGYPPQQGYPP---------QQGYPP----QQGYPPQQGYPP---QQGYPPQQGYPP 96
Query: 324 LNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQ 361
Q YP Q P QQ Q G+PP +P QQ
Sbjct: 97 ----QQGYPPQQGYPP---QQGYPPQQGYPPQGYPPQQ 127
>GLTC_WHEAT (P16315) Glutenin, low molecular weight subunit PTDUCD1
precursor
Length = 295
Score = 46.6 bits (109), Expect = 1e-04
Identities = 65/230 (28%), Positives = 92/230 (39%), Gaps = 21/230 (9%)
Query: 241 QVDALDFPMQQLSFQQNGDPPLN--FPHQV-FPQQNCATPLNVQQLSFQQNGD--PPLNF 295
Q++ P + +Q+ PP + FP Q FPQQ P + QQ SF Q P L F
Sbjct: 20 QMETSCIPGLERPWQEQPLPPQHTLFPQQQPFPQQQ-QPPFSQQQPSFLQQQPILPQLPF 78
Query: 296 PHQVYPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQ-NCTTPLNVQQLS----FQQN 350
Q P +P + QQL PP QV QQ P +QQL+ F Q
Sbjct: 79 SQQQQPVLPQQSPFSQQQLVL-----PPQQQYQQVLQQQIPIVQPSVLQQLNPCKVFLQQ 133
Query: 351 GHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQLMVPPLD 410
P+ P Q+L Q +S M + Q+ E+ +T ++
Sbjct: 134 QCNPVAMP-QRLARSQMLQQSSCHVMQQQCCQQLPQIPEQSRYDVIRAITYSIILQEQQQ 192
Query: 411 APMQQLQERKHTLGDSVLASQQHPVPPLEV----MPMQQLQERPQEHQAV 456
+Q Q++ LG V SQQ L P QQL ++PQ+ Q +
Sbjct: 193 GFVQAQQQQPQQLGQGVSQSQQQSQQQLGQCSFQQPQQQLGQQPQQQQVL 242
>GDA9_WHEAT (P18573) Alpha/beta-gliadin MM1 precursor (Prolamin)
Length = 307
Score = 46.6 bits (109), Expect = 1e-04
Identities = 70/253 (27%), Positives = 93/253 (36%), Gaps = 42/253 (16%)
Query: 212 LNAPMQELSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQ 271
+ P+ +L PQ Q P +QV Q Q FP QQ F P Q +PQ
Sbjct: 21 VRVPVPQLQPQNPSQQ--QPQEQVPLVQQQ----QFPGQQQPFP---------PQQPYPQ 65
Query: 272 QNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQNGDPPLNFPHQVY 331
P QQ Q P P YPQ P QL + Q P P Q Y
Sbjct: 66 PQ---PFPSQQPYLQLQPFPQPQLP---YPQPQLPYPQ--PQLPYPQ---PQPFRPQQPY 114
Query: 332 PQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQ-SGLASQQHMVPPLDAPMQQ---- 386
PQ Q +S QQ QQ Q+ QQ QQ ++P D +QQ
Sbjct: 115 PQSQPQYSQPQQPISQQQQQQQQQQQQKQQQQQQQQILQQILQQQLIPCRDVVLQQHSIA 174
Query: 387 ------LQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLEV 440
LQ+ + L L QQL P + Q + H + +L QQ +
Sbjct: 175 YGSSQVLQQSTYQLVQQ-LCCQQLWQIPEQSRCQAIHNVVHAI---ILHQQQQQQQQQQQ 230
Query: 441 MPMQQLQ-ERPQE 452
P+ Q+ ++PQ+
Sbjct: 231 QPLSQVSFQQPQQ 243
Score = 33.5 bits (75), Expect = 1.2
Identities = 48/174 (27%), Positives = 61/174 (34%), Gaps = 21/174 (12%)
Query: 204 PQQNQVPQLNAPMQELSPQQNGVQLDAPMQQVSSQQNQ----------VDALDFPMQQLS 253
P PQ + P Q +S QQ Q +Q QQ Q + D +QQ S
Sbjct: 113 PYPQSQPQYSQPQQPISQQQQQQQQQQQQKQQQQQQQQILQQILQQQLIPCRDVVLQQHS 172
Query: 254 FQQNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQ 313
QV Q +T VQQL QQ P Q A L+ QQ
Sbjct: 173 IAYGSS-------QVLQQ---STYQLVQQLCCQQLWQIPEQSRCQAIHNVVHAIILHQQQ 222
Query: 314 LSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQ 367
QQ PL+ PQQ + Q S QQN + QQL + ++
Sbjct: 223 QQQQQQQQQPLSQVSFQQPQQQYPSGQGSFQPS-QQNPQAQGSVQPQQLPQFEE 275
Score = 31.2 bits (69), Expect = 6.0
Identities = 36/131 (27%), Positives = 47/131 (35%), Gaps = 19/131 (14%)
Query: 327 PHQVYPQQNCTTPLNVQQLSF--QQNGHPPLN-FPMQQLQEIQQSGLASQQHMVPPLDAP 383
P Q PQ+ PL VQQ F QQ PP +P Q QQ L Q P L P
Sbjct: 33 PSQQQPQEQ--VPL-VQQQQFPGQQQPFPPQQPYPQPQPFPSQQPYLQLQPFPQPQLPYP 89
Query: 384 MQQLQERQHTLGDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLEVMPM 443
QL Q L P P ++ + + Q P+ +
Sbjct: 90 QPQLPYPQPQL-------------PYPQPQPFRPQQPYPQSQPQYSQPQQPISQQQQQQQ 136
Query: 444 QQLQERPQEHQ 454
QQ Q++ Q+ Q
Sbjct: 137 QQQQQKQQQQQ 147
>FXP2_PONPY (Q8MJ98) Forkhead box protein P2
Length = 714
Score = 46.6 bits (109), Expect = 1e-04
Identities = 55/173 (31%), Positives = 68/173 (38%), Gaps = 22/173 (12%)
Query: 249 MQQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATP 308
MQQ+ QQ P Q QQ A L QQL + Y +Q
Sbjct: 104 MQQILQQQVLSPQ---QLQALLQQQQAVMLQQQQLQ-------------EFYKKQQEQLH 147
Query: 309 LNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQS 368
L + Q QQ Q QQ QQ QQ HP QQ Q+ QQ
Sbjct: 148 LQLLQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQHPGKQAKEQQQQQQQQQ 207
Query: 369 GLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQLM-VPPLDA--PMQQLQE 418
LA+QQ + MQQLQ++QH L L Q L+ +PP A P+Q L +
Sbjct: 208 QLAAQQLVFQQQLLQMQQLQQQQHLL---SLQRQGLISIPPGQAALPVQSLPQ 257
Score = 44.7 bits (104), Expect = 5e-04
Identities = 59/193 (30%), Positives = 79/193 (40%), Gaps = 22/193 (11%)
Query: 276 TPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLS-FQQNGDPPLNFPHQVYPQQ 334
TP +QQ+ QQ P Q QQ A L QQL F + L+ Q+ QQ
Sbjct: 100 TPQQMQQILQQQVLSPQ---QLQALLQQQQAVMLQQQQLQEFYKKQQEQLHL--QLLQQQ 154
Query: 335 NCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPMQQLQERQHTL 394
QQ QQ QQ Q+ QQ QQ P A QQ Q++Q
Sbjct: 155 QQQQQQQQQQQQQQQQQQ------QQQQQQQQQQQQQQQQQQHPGKQAKEQQQQQQQQ-- 206
Query: 395 GDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLE-VMPMQQLQERPQEH 453
L +QQL+ MQQLQ+++H L S+ +PP + +P+Q L PQ
Sbjct: 207 --QQLAAQQLVFQQQLLQMQQLQQQQHLL--SLQRQGLISIPPGQAALPVQSL---PQAG 259
Query: 454 QAVNSRHFVWKPV 466
+ +WK V
Sbjct: 260 LSPAEIQQLWKEV 272
Score = 32.7 bits (73), Expect = 2.1
Identities = 46/151 (30%), Positives = 54/151 (35%), Gaps = 23/151 (15%)
Query: 205 QQNQVPQLNAPMQE-----LSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGD 259
QQ Q+ + QE L QQ Q QQ QQ Q QQ QQ
Sbjct: 131 QQQQLQEFYKKQQEQLHLQLLQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ 190
Query: 260 PPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQN 319
P + QQ L QQL FQQ L Q+ QQ+ LS Q+
Sbjct: 191 HPGKQAKEQQQQQQQQQQLAAQQLVFQQQ----LLQMQQLQQQQHL--------LSLQRQ 238
Query: 320 G---DPP--LNFPHQVYPQQNCTTPLNVQQL 345
G PP P Q PQ +P +QQL
Sbjct: 239 GLISIPPGQAALPVQSLPQAG-LSPAEIQQL 268
>FXP2_MACMU (Q8MJ97) Forkhead box protein P2
Length = 714
Score = 46.6 bits (109), Expect = 1e-04
Identities = 55/173 (31%), Positives = 68/173 (38%), Gaps = 22/173 (12%)
Query: 249 MQQLSFQQNGDPPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATP 308
MQQ+ QQ P Q QQ A L QQL + Y +Q
Sbjct: 104 MQQILQQQVLSPQ---QLQALLQQQQAVMLQQQQLQ-------------EFYKKQQEQLH 147
Query: 309 LNVQQLSFQQNGDPPLNFPHQVYPQQNCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQS 368
L + Q QQ Q QQ QQ QQ HP QQ Q+ QQ
Sbjct: 148 LQLLQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQHPGKQAKEQQQQQQQQQ 207
Query: 369 GLASQQHMVPPLDAPMQQLQERQHTLGDSGLTSQQLM-VPPLDA--PMQQLQE 418
LA+QQ + MQQLQ++QH L L Q L+ +PP A P+Q L +
Sbjct: 208 QLAAQQLVFQQQLLQMQQLQQQQHLL---SLQRQGLISIPPGQAALPVQSLPQ 257
Score = 44.7 bits (104), Expect = 5e-04
Identities = 59/193 (30%), Positives = 79/193 (40%), Gaps = 22/193 (11%)
Query: 276 TPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLS-FQQNGDPPLNFPHQVYPQQ 334
TP +QQ+ QQ P Q QQ A L QQL F + L+ Q+ QQ
Sbjct: 100 TPQQMQQILQQQVLSPQ---QLQALLQQQQAVMLQQQQLQEFYKKQQEQLHL--QLLQQQ 154
Query: 335 NCTTPLNVQQLSFQQNGHPPLNFPMQQLQEIQQSGLASQQHMVPPLDAPMQQLQERQHTL 394
QQ QQ QQ Q+ QQ QQ P A QQ Q++Q
Sbjct: 155 QQQQQQQQQQQQQQQQQQ------QQQQQQQQQQQQQQQQQQHPGKQAKEQQQQQQQQ-- 206
Query: 395 GDSGLTSQQLMVPPLDAPMQQLQERKHTLGDSVLASQQHPVPPLE-VMPMQQLQERPQEH 453
L +QQL+ MQQLQ+++H L S+ +PP + +P+Q L PQ
Sbjct: 207 --QQLAAQQLVFQQQLLQMQQLQQQQHLL--SLQRQGLISIPPGQAALPVQSL---PQAG 259
Query: 454 QAVNSRHFVWKPV 466
+ +WK V
Sbjct: 260 LSPAEIQQLWKEV 272
Score = 32.7 bits (73), Expect = 2.1
Identities = 46/151 (30%), Positives = 54/151 (35%), Gaps = 23/151 (15%)
Query: 205 QQNQVPQLNAPMQE-----LSPQQNGVQLDAPMQQVSSQQNQVDALDFPMQQLSFQQNGD 259
QQ Q+ + QE L QQ Q QQ QQ Q QQ QQ
Sbjct: 131 QQQQLQEFYKKQQEQLHLQLLQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQQ 190
Query: 260 PPLNFPHQVFPQQNCATPLNVQQLSFQQNGDPPLNFPHQVYPQQNCATPLNVQQLSFQQN 319
P + QQ L QQL FQQ L Q+ QQ+ LS Q+
Sbjct: 191 HPGKQAKEQQQQQQQQQQLAAQQLVFQQQ----LLQMQQLQQQQHL--------LSLQRQ 238
Query: 320 G---DPP--LNFPHQVYPQQNCTTPLNVQQL 345
G PP P Q PQ +P +QQL
Sbjct: 239 GLISIPPGQAALPVQSLPQAG-LSPAEIQQL 268
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.132 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,474,949
Number of Sequences: 164201
Number of extensions: 2828727
Number of successful extensions: 7294
Number of sequences better than 10.0: 209
Number of HSP's better than 10.0 without gapping: 33
Number of HSP's successfully gapped in prelim test: 179
Number of HSP's that attempted gapping in prelim test: 6441
Number of HSP's gapped (non-prelim): 590
length of query: 471
length of database: 59,974,054
effective HSP length: 114
effective length of query: 357
effective length of database: 41,255,140
effective search space: 14728084980
effective search space used: 14728084980
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0213.5


