

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0192.15
(1070 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
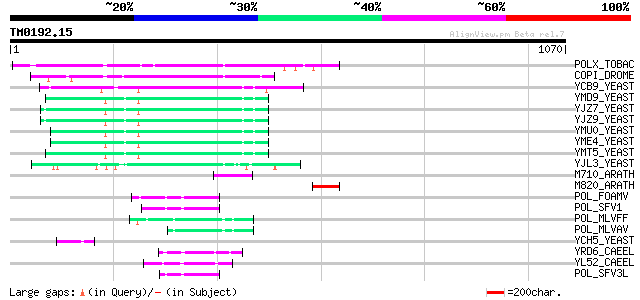
Score E
Sequences producing significant alignments: (bits) Value
POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from tran... 321 6e-87
COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contain... 253 2e-66
YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B) 123 3e-27
YMD9_YEAST (Q03434) Transposon Ty1 protein B 120 3e-26
YJZ7_YEAST (P47098) Transposon Ty1 protein B 119 3e-26
YJZ9_YEAST (P47100) Transposon Ty1 protein B 119 6e-26
YMU0_YEAST (Q04670) Transposon Ty1 protein B 118 1e-25
YME4_YEAST (Q04711) Transposon Ty1 protein B 118 1e-25
YMT5_YEAST (Q04214) Transposon Ty1 protein B 117 1e-25
YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein 86 4e-16
M710_ARATH (P92512) Hypothetical mitochondrial protein AtMg00710... 59 5e-08
M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820... 55 8e-07
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 54 2e-06
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 51 2e-05
POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC 3.4.2... 50 2e-05
POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.2... 50 2e-05
YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein 50 3e-05
YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II 49 6e-05
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 49 6e-05
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 49 7e-05
>POLX_TOBAC (P10978) Retrovirus-related Pol polyprotein from
transposon TNT 1-94 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1328
Score = 321 bits (823), Expect = 6e-87
Identities = 219/672 (32%), Positives = 337/672 (49%), Gaps = 62/672 (9%)
Query: 5 RESSRNFSKN-SQDKNPPCSIC*RLGHAEADCRYRDKPQCNYCKKFGHMEKYCYSKNRHQ 63
R +R SKN S+ + C C + GH + DC K G E + +
Sbjct: 214 RSGARGKSKNRSKSRVRNCYNCNQPGHFKRDCPNPRK---------GKGETSGQKNDDNT 264
Query: 64 ANLAEEQEQDQYLLYATQDSARETG--GSWYLDSGCSNHMAKDESIFKNIDDSVKVKVRL 121
A + + + + ++ +G W +D+ S+H +F V++
Sbjct: 265 AAMVQNNDNVVLFINEEEECMHLSGPESEWVVDTAASHHATPVRDLFCRYVAGDFGTVKM 324
Query: 122 GNGAVVESKGKGTVMVET*KG-TRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVCR 180
GN + + G G + ++T G T ++ DV VP+L+ NL+S + GY +F R
Sbjct: 325 GNTSYSKIAGIGDICIKTNVGCTLVLKDVRHVPDLRMNLISGIALDRDGYESYFANQKWR 384
Query: 181 IYDKHDKRVEIAQVKMQKSNISFPLNFKYVANIAMKAQVDDSW-LWHRRFGHFNTQVLKL 239
+ + + K + N + AQ + S LWH+R GH + + L++
Sbjct: 385 L-----TKGSLVIAKGVARGTLYRTNAEICQGELNAAQDEISVDLWHKRMGHMSEKGLQI 439
Query: 240 LYQKNMMRDLPSLKESNEACERCLLGMQRRVSFSTCKEWRAKDVLELIHTDVCGPMRTSS 299
L +K+++ + + C+ CL G Q RVSF T E R ++L+L+++DVCGPM S
Sbjct: 440 LAKKSLISYAKGT--TVKPCDYCLFGKQHRVSFQTSSE-RKLNILDLVYSDVCGPMEIES 496
Query: 300 HDNNRYFILFTDDFSRMTWVYFLKAKSEVFRVFKKFKALVEKQSGKHIKVLRSDRGKEYT 359
N+YF+ F DD SR WVY LK K +VF+VF+KF ALVE+++G+ +K LRSD G EYT
Sbjct: 497 MGGNKYFVTFIDDASRKLWVYILKTKDQVFQVFQKFHALVERETGRKLKRLRSDNGGEYT 556
Query: 360 SREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVY 419
SREF+++C GI + TV PQ NGV+ER RTI+E RSML+ + +F W EAV
Sbjct: 557 SREFEEYCSSHGIRHEKTVPGTPQHNGVAERMNRTIVEKVRSMLRMAKLPKSF-WGEAVQ 615
Query: 420 TAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGSICYTHVPDVKRHKLEDKNVRGI 479
TA Y++NR P+ + + P W K+ S HL+VFG + HVP +R KL+DK++ I
Sbjct: 616 TACYLINRSPSVPLAFEIPERVWTNKEVSYSHLKVFGCRAFAHVPKEQRTKLDDKSIPCI 675
Query: 480 FLGYSTKSKGYRVYNLQTKKLTISRDVEV-DGNASWNWDEEKVEKNILIP---------- 528
F+GY + GYR+++ KK+ SRDV + D + KN +IP
Sbjct: 676 FIGYGDEEFGYRLWDPVKKKVIRSRDVVFRESEVRTAADMSEKVKNGIIPNFVTIPSTSN 735
Query: 529 ----TQRPQEEVEEEAENPGEPTS--------------PPQQQEQQQDLSPESTPRRVRF 570
+ +EV E+ E PGE P Q +EQ Q L P R
Sbjct: 736 NPTSAESTTDEVSEQGEQPGEVIEQGEQLDEGVEEVEHPTQGEEQHQPLRRSERP---RV 792
Query: 571 LVDVYETCNLAIL----EPESFE---AASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGK 623
Y + ++ EPES + + ++ +KAM+EE++ ++KN T++LV+ P GK
Sbjct: 793 ESRRYPSTEYVLISDDREPESLKEVLSHPEKNQLMKAMQEEMESLQKNGTYKLVELPKGK 852
Query: 624 DIIGVKWVYKTK 635
+ KWV+K K
Sbjct: 853 RPLKCKWVFKLK 864
>COPI_DROME (P04146) Copia protein (Gag-int-pol protein) [Contains:
Copia VLP protein; Copia protease (EC 3.4.23.-)]
Length = 1409
Score = 253 bits (646), Expect = 2e-66
Identities = 169/492 (34%), Positives = 261/492 (52%), Gaps = 38/492 (7%)
Query: 40 KPQCNYCKKFGHMEKYCYSKNRHQANLAEEQEQD---------QYLLYATQDSARETGGS 90
K +C++C + GH++K C+ R N +E E+ +++ +++
Sbjct: 229 KVKCHHCGREGHIKKDCFHYKRILNNKNKENEKQVQTATSHGIAFMVKEVNNTSVMDNCG 288
Query: 91 WYLDSGCSNHMAKDESIFKNIDDSVKV----KVRLGN-GAVVESKGKGTVMVET*KGTRL 145
+ LDSG S+H+ DES++ DSV+V K+ + G + + +G V + L
Sbjct: 289 FVLDSGASDHLINDESLYT---DSVEVVPPLKIAVAKQGEFIYATKRGIVRLRNDHEITL 345
Query: 146 ISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVCRIYDKHDKRVEIAQVKMQKSNISFP- 204
DVL NL+S+ ++ E G ++ F+ I + + VK + P
Sbjct: 346 -EDVLFCKEAAGNLMSVKRLQEAGMSIEFDKSGVTI-----SKNGLMVVKNSGMLNNVPV 399
Query: 205 LNFK-YVANIAMKAQVDDSWLWHRRFGHFNTQVLKLLYQKNMMRD---LPSLKESNEACE 260
+NF+ Y N K ++ LWH RFGH + L + +KNM D L +L+ S E CE
Sbjct: 400 INFQAYSINAKHK---NNFRLWHERFGHISDGKLLEIKRKNMFSDQSLLNNLELSCEICE 456
Query: 261 RCLLGMQRRVSFSTCKE-WRAKDVLELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWV 319
CL G Q R+ F K+ K L ++H+DVCGP+ + D+ YF++F D F+
Sbjct: 457 PCLNGKQARLPFKQLKDKTHIKRPLFVVHSDVCGPITPVTLDDKNYFVIFVDQFTHYCVT 516
Query: 320 YFLKAKSEVFRVFKKFKALVEKQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVA 379
Y +K KS+VF +F+ F A E + L D G+EY S E +FC +GI LTV
Sbjct: 517 YLIKYKSDVFSMFQDFVAKSEAHFNLKVVYLYIDNGREYLSNEMRQFCVKKGISYHLTVP 576
Query: 380 YIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAV--QNKT 437
+ PQ NGVSER RTI E AR+M+ + +F W EAV TA Y++NR P+ A+ +KT
Sbjct: 577 HTPQLNGVSERMIRTITEKARTMVSGAKLDKSF-WGEAVLTATYLINRIPSRALVDSSKT 635
Query: 438 PIEAWCGKKPSTKHLRVFGSICYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYNLQT 497
P E W KKP KHLRVFG+ Y H+ + K+ K +DK+ + IF+GY + G+++++
Sbjct: 636 PYEMWHNKKPYLKHLRVFGATVYVHIKN-KQGKFDDKSFKSIFVGY--EPNGFKLWDAVN 692
Query: 498 KKLTISRDVEVD 509
+K ++RDV VD
Sbjct: 693 EKFIVARDVVVD 704
Score = 37.0 bits (84), Expect = 0.28
Identities = 16/54 (29%), Positives = 28/54 (51%), Gaps = 3/54 (5%)
Query: 585 PESFEAASKQE---VWVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWVYKTK 635
P SF+ ++ W +A+ E+ + NNTW + P K+I+ +WV+ K
Sbjct: 891 PNSFDEIQYRDDKSSWEEAINTELNAHKINNTWTITKRPENKNIVDSRWVFSVK 944
>YCB9_YEAST (P25384) Transposon Ty2 protein B (Ty1-17 protein B)
Length = 1770
Score = 123 bits (308), Expect = 3e-27
Identities = 122/538 (22%), Positives = 219/538 (40%), Gaps = 42/538 (7%)
Query: 57 YSKNRHQANLAEEQEQDQYLLYATQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVK 116
Y + ++ +L ++Q++ + T DS E +DSG S + + + + +
Sbjct: 422 YLSDDNELSLGQQQKESKPT--HTIDSNDELPDHLLIDSGASQTLVRSAHYLHHATPNSE 479
Query: 117 VKVRLGNGAVVESKGKGTVMVET*KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHF-- 174
+ + + G + GT+ L PN+ +LLS+ ++ + T F
Sbjct: 480 INIVDAQKQDIPINAIGNLHFNFQNGTKTSIKALHTPNIAYDLLSLSELANQNITACFTR 539
Query: 175 ------EGDVCRIYDKHDKRVEIAQVKMQKSNISFPLNFKYVANIAMKAQVDDSW---LW 225
+G V KH +++ + S+IS K N K++ + + L
Sbjct: 540 NTLERSDGTVLAPIVKHGDFYWLSKKYLIPSHIS-----KLTINNVNKSKSVNKYPYPLI 594
Query: 226 HRRFGHFNTQVLKLLYQKNMMR-----DLPSLKESNEACERCLLGMQ---RRVSFSTCKE 277
HR GH N + ++ +KN + D+ S C CL+G R V S K
Sbjct: 595 HRMLGHANFRSIQKSLKKNAVTYLKESDIEWSNASTYQCPDCLIGKSTKHRHVKGSRLKY 654
Query: 278 WRAKDVLELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSE--VFRVFKKF 335
+ + + +HTD+ GP+ YFI FTD+ +R WVY L + E + VF
Sbjct: 655 QESYEPFQYLHTDIFGPVHHLPKSAPSYFISFTDEKTRFQWVYPLHDRREESILNVFTSI 714
Query: 336 KALVEKQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTI 395
A ++ Q + V++ DRG EYT++ KF + GI T + +GV+ER RT+
Sbjct: 715 LAFIKNQFNARVLVIQMDRGSEYTNKTLHKFFTNRGITACYTTTADSRAHGVAERLNRTL 774
Query: 396 MEMARSMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVF 455
+ R++L G+ N W++ ++ + + ++ G +T + F
Sbjct: 775 LNDCRTLLHCSGLPNHLWFSAVEFSTIIRNSLVSPKNDKSARQHAGLAGLDITT--ILPF 832
Query: 456 GS--ICYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVY------NLQTKKLTISRDVE 507
G I H PD K H + + G L S S GY +Y + T I +D +
Sbjct: 833 GQPVIVNNHNPDSKIH---PRGIPGYALHPSRNSYGYIIYLPSLKKTVDTTNYVILQDKQ 889
Query: 508 VDGNASWNWDEEKVEKNILIPTQRPQEEVEEEAENPGEPTSPPQQQEQQQDLSPESTP 565
+ +N+D + ++ T Q +E+ + + Q ++ S P
Sbjct: 890 SKLD-QFNYDTLTFDDDLNRLTAHNQSFIEQNETEQSYDQNTESDHDYQSEIEINSDP 946
>YMD9_YEAST (Q03434) Transposon Ty1 protein B
Length = 1328
Score = 120 bits (300), Expect = 3e-26
Identities = 112/451 (24%), Positives = 183/451 (39%), Gaps = 29/451 (6%)
Query: 70 QEQDQYLLYATQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVKVKVRLGNGAVVES 129
QE + + T S E G LDSG S + + + + + V +
Sbjct: 10 QELTESTVNHTNHSDDELPGHLLLDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPI 69
Query: 130 KGKGTVMVET*KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVCRIYD------ 183
G + T+ VL PN+ +LLS+ ++ T F +V D
Sbjct: 70 NAIGDLQFHFQDNTKTSIKVLHTPNIAYDLLSLNELAAVDITACFTKNVLERSDGTVLAP 129
Query: 184 --KHDKRVEIAQVKMQKSNISFP-LNFKYVANIAMKAQVDDSWLWHRRFGHFNTQVLKLL 240
K+ +++ + SNIS P +N + + K HR H N Q ++
Sbjct: 130 IVKYGDFYWVSKKYLLPSNISVPTINNVHTSESTRKYPYP---FIHRMLAHANAQTIRYS 186
Query: 241 YQKNMMR-----DLPSLKESNEACERCLLGMQ---RRVSFSTCKEWRAKDVLELIHTDVC 292
+ N + D+ + C CL+G R + S K + + + +HTD+
Sbjct: 187 LKNNTITYFNESDVDRSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF 246
Query: 293 GPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSE--VFRVFKKFKALVEKQSGKHIKVL 350
GP+ YFI FTD+ ++ WVY L + E + VF A ++ Q + V+
Sbjct: 247 GPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVI 306
Query: 351 RSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHN 410
+ DRG EYT+R KF E GI T + +GV+ER RT+++ R+ L+ G+ N
Sbjct: 307 QMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPN 366
Query: 411 TFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGS--ICYTHVPDVKR 468
W++ ++ + + + ++ G ST L FG I H P+ K
Sbjct: 367 HLWFSAIEFSTIVRNSLASPKSKKSARQHAGLAGLDIST--LLPFGQPVIVNDHNPNSKI 424
Query: 469 HKLEDKNVRGIFLGYSTKSKGYRVYNLQTKK 499
H + + G L S S GY +Y KK
Sbjct: 425 H---PRGIPGYALHPSRNSYGYIIYLPSLKK 452
>YJZ7_YEAST (P47098) Transposon Ty1 protein B
Length = 1755
Score = 119 bits (299), Expect = 3e-26
Identities = 114/461 (24%), Positives = 187/461 (39%), Gaps = 31/461 (6%)
Query: 60 NRHQANLAEEQEQDQYLLYATQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVKVKV 119
N+H L QE + + T S E G LDSG S + + + + + V
Sbjct: 429 NKHDLTLG--QELTESTVNHTNHSDDELPGHLLLDSGASRTLIRSAHHIHSASSNPDINV 486
Query: 120 RLGNGAVVESKGKGTVMVET*KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVC 179
+ G + T+ VL PN+ +LLS+ ++ T F +V
Sbjct: 487 VDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHTPNIAYDLLSLNELAAVDITACFTKNVL 546
Query: 180 RIYD--------KHDKRVEIAQVKMQKSNISFP-LNFKYVANIAMKAQVDDSWLWHRRFG 230
D ++ +++ + SNIS P +N + + K HR
Sbjct: 547 ERSDGTVLAPIVQYGDFYWVSKRYLLPSNISVPTINNVHTSESTRKYPYP---FIHRMLA 603
Query: 231 HFNTQVLKLLYQKNMMR-----DLPSLKESNEACERCLLGMQ---RRVSFSTCKEWRAKD 282
H N Q ++ + N + D+ + C CL+G R + S K + +
Sbjct: 604 HANAQTIRYSLKNNTITYFNESDVDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYE 663
Query: 283 VLELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSE--VFRVFKKFKALVE 340
+ +HTD+ GP+ YFI FTD+ ++ WVY L + E + VF A ++
Sbjct: 664 PFQYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIK 723
Query: 341 KQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMAR 400
Q + V++ DRG EYT+R KF E GI T + +GV+ER RT+++ R
Sbjct: 724 NQFQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCR 783
Query: 401 SMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGS--I 458
+ L+ G+ N W++ ++ + + + ++ G ST L FG I
Sbjct: 784 TQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSKKSARQHAGLAGLDIST--LLPFGQPVI 841
Query: 459 CYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYNLQTKK 499
H P+ K H + + G L S S GY +Y KK
Sbjct: 842 VNDHNPNSKIH---PRGIPGYALHPSRNSYGYIIYLPSLKK 879
>YJZ9_YEAST (P47100) Transposon Ty1 protein B
Length = 1755
Score = 119 bits (297), Expect = 6e-26
Identities = 114/461 (24%), Positives = 187/461 (39%), Gaps = 31/461 (6%)
Query: 60 NRHQANLAEEQEQDQYLLYATQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVKVKV 119
N+H L Q+ + + T S E G LDSG S + + + + + V
Sbjct: 429 NKHDLILG--QKLTESTVNHTNHSDDELPGHLLLDSGASRTLIRSAHHIHSASSNPDINV 486
Query: 120 RLGNGAVVESKGKGTVMVET*KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVC 179
+ G + T+ VL PN+ +LLS+ ++ T F +V
Sbjct: 487 VDAQKRNIPINAIGDLQFHFQDNTKTSIKVLHTPNIAYDLLSLNELAAVDITACFTKNVL 546
Query: 180 RIYD--------KHDKRVEIAQVKMQKSNISFP-LNFKYVANIAMKAQVDDSWLWHRRFG 230
D K+ +++ + SNIS P +N + + K HR
Sbjct: 547 ERSDGTVLAPIVKYGDFYWVSKKYLLPSNISVPTINNVHTSESTRKYPYP---FIHRMLA 603
Query: 231 HFNTQVLKLLYQKNMMR-----DLPSLKESNEACERCLLGMQ---RRVSFSTCKEWRAKD 282
H N Q ++ + N + D+ + C CL+G R + S K + +
Sbjct: 604 HANAQTIRYSLKNNTITYFNESDVDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYE 663
Query: 283 VLELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSE--VFRVFKKFKALVE 340
+ +HTD+ GP+ YFI FTD+ ++ WVY L + E + VF A ++
Sbjct: 664 PFQYLHTDIFGPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIK 723
Query: 341 KQSGKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMAR 400
Q + V++ DRG EYT+R KF E GI T + +GV+ER RT+++ R
Sbjct: 724 NQFQASVLVIQMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCR 783
Query: 401 SMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGS--I 458
+ L+ G+ N W++ ++ + + + ++ G ST L FG I
Sbjct: 784 TQLQCSGLPNHLWFSAIEFSTIVRNSLASPKSKKSARQHAGLAGLDIST--LLPFGQPVI 841
Query: 459 CYTHVPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYNLQTKK 499
H P+ K H + + G L S S GY +Y KK
Sbjct: 842 VNDHNPNSKIH---PRGIPGYALHPSRNSYGYIIYLPSLKK 879
>YMU0_YEAST (Q04670) Transposon Ty1 protein B
Length = 1328
Score = 118 bits (295), Expect = 1e-25
Identities = 110/441 (24%), Positives = 179/441 (39%), Gaps = 29/441 (6%)
Query: 80 TQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVKVKVRLGNGAVVESKGKGTVMVET 139
T S E G LDSG S + + + + + V + G +
Sbjct: 20 TNHSDDELPGHLLLDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHF 79
Query: 140 *KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVCRIYD--------KHDKRVEI 191
T+ VL PN+ +LLS+ ++ T F +V D K+ +
Sbjct: 80 QDNTKTSIKVLHTPNIAYDLLSLNELAAVDITACFTKNVLERSDGTVLAPIVKYGDFYWV 139
Query: 192 AQVKMQKSNISFP-LNFKYVANIAMKAQVDDSWLWHRRFGHFNTQVLKLLYQKNMMR--- 247
++ + SNIS P +N + + K HR H N Q ++ + N +
Sbjct: 140 SKKYLLPSNISVPTINNVHTSESTRKYPYP---FIHRMLAHANAQTIRYSLKNNTITYFN 196
Query: 248 --DLPSLKESNEACERCLLGMQ---RRVSFSTCKEWRAKDVLELIHTDVCGPMRTSSHDN 302
D+ + C CL+G R + S K + + + +HTD+ GP+
Sbjct: 197 ESDVDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSA 256
Query: 303 NRYFILFTDDFSRMTWVYFLKAKSE--VFRVFKKFKALVEKQSGKHIKVLRSDRGKEYTS 360
YFI FTD+ ++ WVY L + E + VF A ++ Q + V++ DRG EYT+
Sbjct: 257 PSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTN 316
Query: 361 REFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYT 420
R KF E GI T + +GV+ER RT+++ R+ L+ G+ N W++ ++
Sbjct: 317 RTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFS 376
Query: 421 AVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGS--ICYTHVPDVKRHKLEDKNVRG 478
+ + + ++ G ST L FG I H P+ K H + + G
Sbjct: 377 TIVRNSLASPKSKKSARQHAGLAGLDIST--LLPFGQPVIVNDHNPNSKIH---PRGIPG 431
Query: 479 IFLGYSTKSKGYRVYNLQTKK 499
L S S GY +Y KK
Sbjct: 432 YALHPSRNSYGYIIYLPSLKK 452
>YME4_YEAST (Q04711) Transposon Ty1 protein B
Length = 1328
Score = 118 bits (295), Expect = 1e-25
Identities = 110/441 (24%), Positives = 179/441 (39%), Gaps = 29/441 (6%)
Query: 80 TQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVKVKVRLGNGAVVESKGKGTVMVET 139
T S E G LDSG S + + + + + V + G +
Sbjct: 20 TNHSDDELPGHLLLDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPINAIGDLQFHF 79
Query: 140 *KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVCRIYD--------KHDKRVEI 191
T+ VL PN+ +LLS+ ++ T F +V D K+ +
Sbjct: 80 QDNTKTSIKVLHTPNIAYDLLSLNELAAVDITACFTKNVLERSDGTVLAPIVKYGDFYWV 139
Query: 192 AQVKMQKSNISFP-LNFKYVANIAMKAQVDDSWLWHRRFGHFNTQVLKLLYQKNMMR--- 247
++ + SNIS P +N + + K HR H N Q ++ + N +
Sbjct: 140 SKKYLLPSNISVPTINNVHTSESTRKYPYP---FIHRMLAHANAQTIRYSLKNNTITYFN 196
Query: 248 --DLPSLKESNEACERCLLGMQ---RRVSFSTCKEWRAKDVLELIHTDVCGPMRTSSHDN 302
D+ + C CL+G R + S K + + + +HTD+ GP+
Sbjct: 197 ESDVDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIFGPVHNLPKSA 256
Query: 303 NRYFILFTDDFSRMTWVYFLKAKSE--VFRVFKKFKALVEKQSGKHIKVLRSDRGKEYTS 360
YFI FTD+ ++ WVY L + E + VF A ++ Q + V++ DRG EYT+
Sbjct: 257 PSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVIQMDRGSEYTN 316
Query: 361 REFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYT 420
R KF E GI T + +GV+ER RT+++ R+ L+ G+ N W++ ++
Sbjct: 317 RTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPNHLWFSAIEFS 376
Query: 421 AVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGS--ICYTHVPDVKRHKLEDKNVRG 478
+ + + ++ G ST L FG I H P+ K H + + G
Sbjct: 377 TIVRNSLASPKSKKSARQHAGLAGLDIST--LLPFGQPVIVNDHNPNSKIH---PRGIPG 431
Query: 479 IFLGYSTKSKGYRVYNLQTKK 499
L S S GY +Y KK
Sbjct: 432 YALHPSRNSYGYIIYLPSLKK 452
>YMT5_YEAST (Q04214) Transposon Ty1 protein B
Length = 1328
Score = 117 bits (294), Expect = 1e-25
Identities = 111/451 (24%), Positives = 183/451 (39%), Gaps = 29/451 (6%)
Query: 70 QEQDQYLLYATQDSARETGGSWYLDSGCSNHMAKDESIFKNIDDSVKVKVRLGNGAVVES 129
QE + + T S + G LDSG S + + + + + V +
Sbjct: 10 QELTESTVNHTNHSDDKLPGHLLLDSGASRTLIRSAHHIHSASSNPDINVVDAQKRNIPI 69
Query: 130 KGKGTVMVET*KGTRLISDVLLVPNLKENLLSIGQMIERGYTLHFEGDVCRIYD------ 183
G + T+ VL PN+ +LLS+ ++ T F +V D
Sbjct: 70 NAIGDLQFHFQDNTKTSIKVLHTPNIAYDLLSLNELAAVDITACFTKNVLERSDGTVLAP 129
Query: 184 --KHDKRVEIAQVKMQKSNISFP-LNFKYVANIAMKAQVDDSWLWHRRFGHFNTQVLKLL 240
K+ +++ + SNIS P +N + + K HR H N Q ++
Sbjct: 130 IVKYGDFYWVSKKYLLPSNISVPTINNVHTSESTRKYPYP---FIHRMLAHANAQTIRYS 186
Query: 241 YQKNMMR-----DLPSLKESNEACERCLLGMQ---RRVSFSTCKEWRAKDVLELIHTDVC 292
+ N + D+ + C CL+G R + S K + + + +HTD+
Sbjct: 187 LKNNTITYFNESDVDWSSAIDYQCPDCLIGKSTKHRHIKGSRLKYQNSYEPFQYLHTDIF 246
Query: 293 GPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSE--VFRVFKKFKALVEKQSGKHIKVL 350
GP+ YFI FTD+ ++ WVY L + E + VF A ++ Q + V+
Sbjct: 247 GPVHNLPKSAPSYFISFTDETTKFRWVYPLHDRREDSILDVFTTILAFIKNQFQASVLVI 306
Query: 351 RSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHN 410
+ DRG EYT+R KF E GI T + +GV+ER RT+++ R+ L+ G+ N
Sbjct: 307 QMDRGSEYTNRTLHKFLEKNGITPCYTTTADSRAHGVAERLNRTLLDDCRTQLQCSGLPN 366
Query: 411 TFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGS--ICYTHVPDVKR 468
W++ ++ + + + ++ G ST L FG I H P+ K
Sbjct: 367 HLWFSAIEFSTIVRNSLASPKSKKSARQHAGLAGLDIST--LLPFGQPVIVNDHNPNSKI 424
Query: 469 HKLEDKNVRGIFLGYSTKSKGYRVYNLQTKK 499
H + + G L S S GY +Y KK
Sbjct: 425 H---PRGIPGYALHPSRNSYGYIIYLPSLKK 452
>YJL3_YEAST (P47024) Transposon Ty4 207.7 kDa hypothetical protein
Length = 1803
Score = 86.3 bits (212), Expect = 4e-16
Identities = 129/591 (21%), Positives = 228/591 (37%), Gaps = 91/591 (15%)
Query: 43 CNYCKKFGHMEKYCYSKNRHQANLAEEQEQDQYLLYATQDS-----ARETGGSW------ 91
C YCK H C K L Q + +D+ + SW
Sbjct: 342 CMYCKSVFHCSINCKKKPNRNLGLTRPISQKPIIYKVHRDNNHLSPVQNEQKSWNKTQKR 401
Query: 92 -----------YLDSGCSNHMAKDESIFKNIDDSVKVK--VRLGNGAVVESKGKGTVMVE 138
+D+G ++ D+++ N +DS + +G + V KG G + ++
Sbjct: 402 SNKVYNSKKLVIIDTGSGVNITNDKTLLHNYEDSNRSTRFFGIGKNSSVSVKGYGYIKIK 461
Query: 139 T*KGTRLISDVLL--VPNLKENLLSIGQM-------IERGYTLHFEGDVCRIYDK----- 184
+L VP + ++S + + R YT + +I K
Sbjct: 462 NGHNNTDNKCLLTYYVPEEESTIISCYDLAKKTKMVLSRKYT-RLGNKIIKIKTKIVNGV 520
Query: 185 -HDKRVEIAQVKMQKSNIS---------FPLNFKYVANIAMKAQVDDSWLWHRRFGHFNT 234
H K E+ + S I+ F LN + + ++D+ H+R GH
Sbjct: 521 IHVKMNELIERPSDDSKINAIKPTSSPGFKLNKRSIT-------LEDA---HKRMGHTGI 570
Query: 235 QVLKLLYQKNMMRD-LPSLKESNEA-CERCLLGMQRRVSFSTCKEWRAKDVLELIHT--- 289
Q ++ + N + L +KE NE C+ C + + + T E +
Sbjct: 571 QQIENSIKHNHYEESLDLIKEPNEFWCQTCKISKATKRNHYTGSMNNHSTDHEPGSSWCM 630
Query: 290 DVCGPMRTSSHDNNRYFILFTDDFSR--MTWVYFLKAKSEVFRVFKKFKALVEKQSGKHI 347
D+ GP+ +S+ D RY ++ D+ +R MT +F K + +K VE Q + +
Sbjct: 631 DIFGPVSSSNADTKRYMLIMVDNNTRYCMTSTHFNKNAETILAQVRKNIQYVETQFDRKV 690
Query: 348 KVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKG 407
+ + SDRG E+T+ + +++ +GI LT NG +ER RTI+ A ++L++
Sbjct: 691 REINSDRGTEFTNDQIEEYFISKGIHHILTSTQDHAANGRAERYIRTIITDATTLLRQSN 750
Query: 408 MHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVF-----GSICYTH 462
+ F W AV +A I N + K P++A ++P T L F I + H
Sbjct: 751 LRVKF-WEYAVTSATNIRNYLEHKST-GKLPLKA-ISRQPVTVRLMSFLPFGEKGIIWNH 807
Query: 463 VPDVKRHKLEDKNVRGIFLGYSTKSKGYRVYNLQTKKLTISRDVEV-----DG------- 510
KL+ + I L S GY+ + K+ S + + DG
Sbjct: 808 ----NHKKLKPSGLPSIILCKDPNSYGYKFFIPSKNKIVTSDNYTIPNYTMDGRVRNTQN 863
Query: 511 -NASWNWDEEKVEKNILIPTQRPQEEVEEEAENPGEPTSPPQQQEQQQDLS 560
N S + + ++ I T E E E+ +P + + +++LS
Sbjct: 864 INKSHQFSSDNDDEEDQIETVTNLCEALENYEDDNKPITRLEDLFTEEELS 914
>M710_ARATH (P92512) Hypothetical mitochondrial protein AtMg00710
(ORF120)
Length = 120
Score = 59.3 bits (142), Expect = 5e-08
Identities = 32/75 (42%), Positives = 44/75 (58%), Gaps = 1/75 (1%)
Query: 393 RTIMEMARSMLKEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHL 452
RTI+E RSML E G+ TF A+A TAV+I+N+ P+ A+ P E W P+ +L
Sbjct: 3 RTIIEKVRSMLCECGLPKTFR-ADAANTAVHIINKYPSTAINFHVPDEVWFQSVPTYSYL 61
Query: 453 RVFGSICYTHVPDVK 467
R FG + Y H + K
Sbjct: 62 RRFGCVAYIHCDEGK 76
>M820_ARATH (P92520) Hypothetical mitochondrial protein AtMg00820
(ORF170)
Length = 170
Score = 55.5 bits (132), Expect = 8e-07
Identities = 25/53 (47%), Positives = 35/53 (65%)
Query: 584 EPESFEAASKQEVWVKAMEEEIKMIEKNNTWELVDCPHGKDIIGVKWVYKTKL 636
EP+S A K W +AM+EE+ + +N TW LV P ++I+G KWV+KTKL
Sbjct: 27 EPKSVIFALKDPGWCQAMQEELDALSRNKTWILVPPPVNQNILGCKWVFKTKL 79
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 54.3 bits (129), Expect = 2e-06
Identities = 44/170 (25%), Positives = 75/170 (43%), Gaps = 10/170 (5%)
Query: 235 QVLKLLYQKNMMRDLPSLKESNEACERCLL-GMQRRVSFSTCKEWRAKDVLELIHTDVCG 293
++ L + NM +D+ +K+ C++CL+ + S + R + + D G
Sbjct: 629 KIANLYWWPNMRKDV--VKQLGR-CQQCLITNASNKASGPILRPDRPQKPFDKFFIDYIG 685
Query: 294 PMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSEVFRVFKKFKALVEKQSGKHIKVLRSD 353
P+ S Y ++ D + TW+Y KA S V K+L S KV+ SD
Sbjct: 686 PLPPSQ--GYLYVLVVVDGMTGFTWLYPTKAPSTSATV----KSLNVLTSIAIPKVIHSD 739
Query: 354 RGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSML 403
+G +TS F ++ ++ GI + + Y PQ ERK I + +L
Sbjct: 740 QGAAFTSSTFAEWAKERGIHLEFSTPYHPQSGSKVERKNSDIKRLLTKLL 789
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 50.8 bits (120), Expect = 2e-05
Identities = 40/151 (26%), Positives = 67/151 (43%), Gaps = 7/151 (4%)
Query: 254 ESNEACERCLL-GMQRRVSFSTCKEWRAKDVLELIHTDVCGPMRTSSHDNNRYFILFTDD 312
+S C++CL+ S + + + + D GP+ S+ + ++ D
Sbjct: 853 KSIRQCKQCLVTNATNLTSPPILRPVKPLKPFDKFYIDYIGPLPPSN--GYLHVLVVVDS 910
Query: 313 FSRMTWVYFLKAKSEVFRVFKKFKALVEKQSGKHIKVLRSDRGKEYTSREFDKFCEDEGI 372
+ W+Y KA S V KAL S KVL SD+G +TS F + +++GI
Sbjct: 911 MTGFVWLYPTKAPSTSATV----KALNMLTSIAIPKVLHSDQGAAFTSSTFADWAKEKGI 966
Query: 373 ERQLTVAYIPQQNGVSERKKRTIMEMARSML 403
+ + + Y PQ +G ERK I + +L
Sbjct: 967 QLEFSTPYHPQSSGKVERKNSDIKRLLTKLL 997
>POL_MLVFF (P26809) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1204
Score = 50.4 bits (119), Expect = 2e-05
Identities = 61/246 (24%), Positives = 94/246 (37%), Gaps = 18/246 (7%)
Query: 231 HFNTQVLKLLYQKN-----MMRDLPSLKESNEACERCLLGMQRRVSFSTCKEW-RAKDVL 284
H + K L ++N M+ +LK+ E C+ C Q S S K+ R +
Sbjct: 857 HLSFSKTKALLERNYCPYYMLNRDRTLKDITETCQACA---QVNASKSAVKQGTRVRGHR 913
Query: 285 ELIHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSEVFRVF-KKFKALVEKQS 343
H ++ +Y ++F D FS WV K E +V KK + +
Sbjct: 914 PGTHWEIDFTEVKPGLYGYKYLLVFIDTFSG--WVEAFPTKKETAKVVTKKLLEEIFPRF 971
Query: 344 GKHIKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSML 403
G +VL +D G + S+ + G++ +L AY PQ +G ER RTI E +
Sbjct: 972 GMP-QVLGTDNGPAFVSKVSQTVADLLGVDWKLHCAYRPQSSGQVERMNRTIKETLTKLT 1030
Query: 404 KEKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGSICYTHV 463
G + W + A+Y P TP E G P + TH
Sbjct: 1031 LATGSRD---WVLLLPLALYRARNTP--GPHGLTPYEILYGAPPPLVNFPDPDMAKVTHN 1085
Query: 464 PDVKRH 469
P ++ H
Sbjct: 1086 PSLQAH 1091
>POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase/ribonuclease H (EC 2.7.7.49) (EC
3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 50.4 bits (119), Expect = 2e-05
Identities = 48/167 (28%), Positives = 68/167 (39%), Gaps = 9/167 (5%)
Query: 304 RYFILFTDDFSRMTWVYFLKAKSEVFRVF-KKFKALVEKQSGKHIKVLRSDRGKEYTSRE 362
+Y ++F D FS WV K E RV KK + + G +VL SD G +TS+
Sbjct: 928 KYLLVFVDTFSG--WVEAFPTKRETARVVSKKLLEEIFPRFGMP-QVLGSDNGPAFTSQV 984
Query: 363 FDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYTAV 422
+ GI+ +L AY PQ +G ER RTI E + G + W + A+
Sbjct: 985 SQSVADLLGIDWKLHCAYRPQSSGQVERMNRTIKETLTKLTLAAGTRD---WVLLLPLAL 1041
Query: 423 YILNRCPTNAVQNKTPIEAWCGKKPSTKHLRVFGSICYTHVPDVKRH 469
Y P TP E G P + T+ P ++ H
Sbjct: 1042 YRARNTP--GPHGLTPYEILYGAPPPLVNFHDPDMSELTNSPSLQAH 1086
>YCH5_YEAST (P25601) Transposon Ty5-1 16.0 kDa hypothetical protein
Length = 146
Score = 50.1 bits (118), Expect = 3e-05
Identities = 27/72 (37%), Positives = 38/72 (52%), Gaps = 4/72 (5%)
Query: 91 WYLDSGCSNHMAKDESIFKNIDDSVKVKVRLGNGAVVESKGKGTVMVET*KGTRLISDVL 150
W D+GC++HM D SIF + S + G G + G GTV + GT + DV
Sbjct: 79 WIFDTGCTSHMCHDRSIFSSFTRSSRKDFVRGVGGSIPIMGSGTVNI----GTVQLHDVS 134
Query: 151 LVPNLKENLLSI 162
VP+L NL+S+
Sbjct: 135 YVPDLPVNLISV 146
>YRD6_CAEEL (Q09575) Hypothetical protein K02A2.6 in chromosome II
Length = 1268
Score = 49.3 bits (116), Expect = 6e-05
Identities = 44/165 (26%), Positives = 70/165 (41%), Gaps = 18/165 (10%)
Query: 287 IHTDVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSEVFRVFKKFKALVEKQSGKH 346
IH D GP+ N Y ++ D ++ V ++ S V + L+E+ H
Sbjct: 852 IHIDFAGPL------NGCYLLVVVDAKTKYAEVKLTRSISAVTTI-----DLLEEIFSIH 900
Query: 347 --IKVLRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSMLK 404
+ + SD G + TS F + C+ GIE + + Y P+ NG +ER T + R + K
Sbjct: 901 GYPETIISDNGTQLTSHLFAQMCQSHGIEHKTSAVYYPRSNGAAERFVDT---LKRGIAK 957
Query: 405 EKGMHNTFWWAEAVYTAVYILNRCPTNAVQNKTPIEAWCGKKPST 449
KG + + Y P +A+ TP E G+K T
Sbjct: 958 IKGEGSVNQQILNKFLISY--RNTPHSALNGSTPAECHFGRKIRT 1000
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 49.3 bits (116), Expect = 6e-05
Identities = 51/174 (29%), Positives = 75/174 (42%), Gaps = 16/174 (9%)
Query: 259 CERCLLGMQRRVSFSTCKEWRAKDVLELIHTDVCGPMRTSSHDNNRYFILFTDDFSRM-T 317
C +CL S+ +R LE++ D+ S NRY + D F++ T
Sbjct: 1510 CAKCLCANDHSKLTSSLTPYRMTFPLEIVACDLMDV--GLSVQGNRYILTIIDLFTKYGT 1567
Query: 318 WVYFLKAKSEVFRVFKKFKALVEKQS---GKHIKVLRSDRGKEYTSREFDKFCEDEGIER 374
V K+E KA VE+ + G+ L +D+GKE+ + F +F IE
Sbjct: 1568 AVPIPDKKAETV-----LKAFVERWAIGEGRIPLKLLTDQGKEFVNGLFAQFTHMLKIEH 1622
Query: 375 QLTVAYIPQQNGVSERKKRTIMEMARSMLKEKGMHNTFWWAEAVYTAVYILNRC 428
T Y + NG ER +TIM ++K+K W + VY AVY N C
Sbjct: 1623 ITTKGYNSRANGAVERFNKTIMH----IMKKKTAVPMEWDDQVVY-AVYAYNNC 1671
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 48.9 bits (115), Expect = 7e-05
Identities = 34/114 (29%), Positives = 54/114 (46%), Gaps = 6/114 (5%)
Query: 290 DVCGPMRTSSHDNNRYFILFTDDFSRMTWVYFLKAKSEVFRVFKKFKALVEKQSGKHIKV 349
D GP+ S+ + ++ D + W+Y KA S V KAL S KV
Sbjct: 892 DYIGPLPPSN--GYLHVLVVVDSMTGFVWLYPTKAPSTSATV----KALNMLTSIAVPKV 945
Query: 350 LRSDRGKEYTSREFDKFCEDEGIERQLTVAYIPQQNGVSERKKRTIMEMARSML 403
+ SD+G +TS F + +++GI+ + + Y PQ +G ERK I + +L
Sbjct: 946 IHSDQGAAFTSATFADWAKNKGIQLEFSTPYHPQSSGKVERKNSDIKRLLTKLL 999
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.140 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 116,918,065
Number of Sequences: 164201
Number of extensions: 4803205
Number of successful extensions: 18392
Number of sequences better than 10.0: 132
Number of HSP's better than 10.0 without gapping: 80
Number of HSP's successfully gapped in prelim test: 52
Number of HSP's that attempted gapping in prelim test: 18055
Number of HSP's gapped (non-prelim): 261
length of query: 1070
length of database: 59,974,054
effective HSP length: 121
effective length of query: 949
effective length of database: 40,105,733
effective search space: 38060340617
effective search space used: 38060340617
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 71 (32.0 bits)
Lotus: description of TM0192.15


