

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0181.13
(507 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
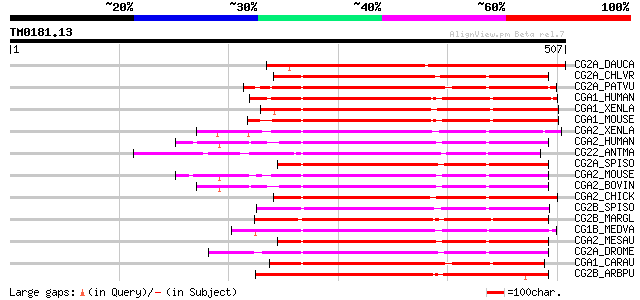
Score E
Sequences producing significant alignments: (bits) Value
CG2A_DAUCA (P25010) G2/mitotic-specific cyclin C13-1 (A-like cyc... 276 1e-73
CG2A_CHLVR (P51986) G2/mitotic-specific cyclin A (Fragment) 217 6e-56
CG2A_PATVU (P24861) G2/mitotic-specific cyclin A 211 3e-54
CGA1_HUMAN (P78396) Cyclin A1 211 4e-54
CGA1_XENLA (P18606) Cyclin A1 208 3e-53
CGA1_MOUSE (Q61456) Cyclin A1 207 7e-53
CGA2_XENLA (P47827) Cyclin A2 201 3e-51
CGA2_HUMAN (P20248) Cyclin A2 (Cyclin A) 197 5e-50
CG22_ANTMA (P34801) G2/mitotic-specific cyclin 2 197 5e-50
CG2A_SPISO (P04962) G2/mitotic-specific cyclin A 197 7e-50
CGA2_MOUSE (P51943) Cyclin A2 (Cyclin A) 196 1e-49
CGA2_BOVIN (P30274) Cyclin A2 (Cyclin A) (Fragment) 196 1e-49
CGA2_CHICK (P43449) Cyclin A2 (Cyclin A) 195 3e-49
CG2B_SPISO (P13952) G2/mitotic-specific cyclin B 193 8e-49
CG2B_MARGL (P15206) G2/mitotic-specific cyclin B 193 8e-49
CG1B_MEDVA (P46277) G2/mitotic-specific cyclin 1 (B-like cyclin)... 193 8e-49
CGA2_MESAU (P37881) Cyclin A2 (Cyclin A) 192 1e-48
CG2A_DROME (P14785) G2/mitotic-specific cyclin A 191 3e-48
CGA1_CARAU (Q92161) Cyclin A1 (Cyclin A) 191 4e-48
CG2B_ARBPU (P07818) G2/mitotic-specific cyclin B 189 1e-47
>CG2A_DAUCA (P25010) G2/mitotic-specific cyclin C13-1 (A-like
cyclin) (Fragment)
Length = 341
Score = 276 bits (705), Expect = 1e-73
Identities = 135/275 (49%), Positives = 196/275 (71%), Gaps = 3/275 (1%)
Query: 235 DPQLCASFACDIYKHLRASE--AKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRL 292
DPQ+C+++ D+Y++L+ E K+RP +++E+VQKD+ ++MR +L+DWLVEV+ EY+L
Sbjct: 67 DPQMCSAYVSDVYEYLKQMEMETKRRPMMNYIEQVQKDVTSNMRGVLVDWLVEVSLEYKL 126
Query: 293 VPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYF 352
+P+TLYL ++Y+DRYLS N ++RQKLQLLGV+S +IASKYEEI V +F ITDNTY
Sbjct: 127 LPETLYLAISYVDRYLSVNVLNRQKLQLLGVSSFLIASKYEEIKPKNVADFVDITDNTYS 186
Query: 353 KEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLME 412
++EV++ME+++L LKFEM +PT+K FL F+RA Q +VP L+ E L NY+AELSL++
Sbjct: 187 QQEVVKMEADLLKTLKFEMGSPTVKTFL-GFIRAVQENPDVPKLKFEFLANYLAELSLLD 245
Query: 413 YSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSP 472
Y L + PSL+AAS FLA+F + P++ PWS LQ + Y+ DL CV LH L
Sbjct: 246 YGCLEFVPSLIAASVTFLARFTIRPNVNPWSIALQKCSGYKSKDLKECVLLLHDLQMGRR 305
Query: 473 NSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEFFQN 507
+L A+++KY +HK+K V+ P IP F +
Sbjct: 306 GGSLSAVRDKYKKHKFKCVSTLSPAPEIPESIFND 340
>CG2A_CHLVR (P51986) G2/mitotic-specific cyclin A (Fragment)
Length = 420
Score = 217 bits (552), Expect = 6e-56
Identities = 118/252 (46%), Positives = 175/252 (68%), Gaps = 7/252 (2%)
Query: 242 FACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTV 301
+A DI+ +L+ SEAK RP +++M K Q DIN+SMRAILIDWLVEV+EEY+L+P TLYL+V
Sbjct: 165 YAQDIHNYLKKSEAKYRPKSNYMRK-QTDINSSMRAILIDWLVEVSEEYKLIPQTLYLSV 223
Query: 302 NYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMES 361
+YIDR+LS ++ R KLQL+G A M++A+K+EEI P+V EF YITD+TY ++VL+ME
Sbjct: 224 SYIDRFLSHMSVLRGKLQLVGAACMLVAAKFEEIYPPEVAEFVYITDDTYTAKQVLRMEH 283
Query: 362 EVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSM-LCYAP 420
+L L F+++ PT + FL R++ AA P QL+ L Y++EL+L+ + + YAP
Sbjct: 284 LILKTLAFDLSVPTCRDFLSRYLFAANA---KPESQLKYLAEYLSELTLINCDISVKYAP 340
Query: 421 SLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIK 480
S++AAS+I +A +L + PW+ TL+ Y+ Y DL C+ E+H L + + AI+
Sbjct: 341 SMIAASSICVANHML--NSIPWTPTLEFYSGYNIQDLRSCLNEIHLLHLAASTNPQQAIQ 398
Query: 481 EKYSQHKYKYVA 492
+KY K+ V+
Sbjct: 399 QKYKSPKFGCVS 410
>CG2A_PATVU (P24861) G2/mitotic-specific cyclin A
Length = 426
Score = 211 bits (538), Expect = 3e-54
Identities = 122/287 (42%), Positives = 182/287 (62%), Gaps = 15/287 (5%)
Query: 214 DILVEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINT 273
D+ +E+ EK ID + + + +A DIYKHLR +E++ R +M+K Q DI
Sbjct: 148 DVTIEDAEKKP----IDREAIILSV-PEYAEDIYKHLREAESRHRSKPGYMKK-QPDITN 201
Query: 274 SMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYE 333
SMR+IL+DW+VEV+EEY+L +TL+L +NYIDR+LS ++ R KLQL+G ASM IASKYE
Sbjct: 202 SMRSILVDWMVEVSEEYKLHRETLFLAINYIDRFLSQMSVLRGKLQLVGAASMFIASKYE 261
Query: 334 EICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEV 393
EI P+V EF YITD+TY +++VL+ME +L L F++ PTI F + + A E
Sbjct: 262 EIYPPEVSEFVYITDDTYEQKQVLRMEHLILKVLSFDVAQPTINWFTDTYAKMADTDETT 321
Query: 394 PSLQLESLTNYIAELSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLY 452
SL + Y++EL+L++ L Y PS +AA+++ LA L +PW S+L + Y
Sbjct: 322 KSLSM-----YLSELTLVDADPYLKYLPSTIAAASLCLANITL--GSEPWPSSLAKESKY 374
Query: 453 QPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPS 499
+ S+ C++E+++ + N+PN AI+EKY KY+ V+ PPS
Sbjct: 375 EISEFSECLQEMYQTYLNAPNHPQQAIREKYKSSKYQQVS-SISPPS 420
>CGA1_HUMAN (P78396) Cyclin A1
Length = 465
Score = 211 bits (537), Expect = 4e-54
Identities = 122/282 (43%), Positives = 180/282 (63%), Gaps = 13/282 (4%)
Query: 220 LEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAIL 279
L + E I ++ D ++ +A +IY++LR +E + RP +M+K Q DI MR IL
Sbjct: 192 LSQSEDISSLGTDVIN---VTEYAEEIYQYLREAEIRHRPKAHYMKK-QPDITEGMRTIL 247
Query: 280 IDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQ 339
+DWLVEV EEY+L +TLYL VN++DR+LS ++ R KLQL+G A+M++ASKYEEI P+
Sbjct: 248 VDWLVEVGEEYKLRAETLYLAVNFLDRFLSCMSVLRGKLQLVGTAAMLLASKYEEIYPPE 307
Query: 340 VEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLE 399
V+EF YITD+TY K ++L+ME +L L F++T PT FL +++R QGV ++ E
Sbjct: 308 VDEFVYITDDTYTKRQLLKMEHLLLKVLAFDLTVPTTNQFLLQYLR-RQGV----CVRTE 362
Query: 400 SLTNYIAELSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLC 458
+L Y+AELSL+E L Y PSL+AA+A LA + + W TL +T Y S++
Sbjct: 363 NLAKYVAELSLLEADPFLKYLPSLIAAAAFCLANYTVNKHF--WPETLAAFTGYSLSEIV 420
Query: 459 VCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSI 500
C+ ELH+ + + P+ AI+EKY KY V+ PP++
Sbjct: 421 PCLSELHKAYLDIPHRPQQAIREKYKASKYLCVSLME-PPAV 461
>CGA1_XENLA (P18606) Cyclin A1
Length = 418
Score = 208 bits (529), Expect = 3e-53
Identities = 117/275 (42%), Positives = 172/275 (62%), Gaps = 11/275 (4%)
Query: 230 DNDHMDPQLCA--SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVA 287
D+ DP A + +I+++LR +E K RP +M K Q DI ++MR IL+DWLVEV
Sbjct: 150 DDSVTDPDAVAVSEYIHEIHQYLREAELKHRPKAYYMRK-QPDITSAMRTILVDWLVEVG 208
Query: 288 EEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYIT 347
EEY+L +TLYL +NY+DR+LS ++ R KLQL+G A++++ASKYEEI P V+EF YIT
Sbjct: 209 EEYKLHTETLYLAMNYLDRFLSCMSVLRGKLQLVGTAAILLASKYEEIYPPDVDEFVYIT 268
Query: 348 DNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAE 407
D+TY K+++L+ME +L L F++T PT+ FL ++++ S+++E L Y+AE
Sbjct: 269 DDTYSKKQLLRMEHVLLKVLAFDLTVPTVNQFLLQYLQ-----RHAVSVKMEHLAMYMAE 323
Query: 408 LSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHR 466
L+L+E L Y PSL AA+A LA + L W TL+ +T Y SD+ C+ +LH+
Sbjct: 324 LTLLEVEPFLKYVPSLTAAAAYCLANYALNKVF--WPDTLEAFTGYALSDIAPCLSDLHQ 381
Query: 467 LFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIP 501
+P AI+EKY KY V+ P +P
Sbjct: 382 FCLGAPYQAQQAIREKYKTTKYMQVSLLEMPSILP 416
>CGA1_MOUSE (Q61456) Cyclin A1
Length = 421
Score = 207 bits (526), Expect = 7e-53
Identities = 120/285 (42%), Positives = 176/285 (61%), Gaps = 19/285 (6%)
Query: 218 EELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRA 277
E + G ++N+ +A +I+++L +E + RP +M K Q DI MRA
Sbjct: 153 EATDFGSDVINV----------TEYAEEIHRYLPEAEVRHRPKAHYMRK-QPDITEGMRA 201
Query: 278 ILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICA 337
IL+DWLVEV EEY+L +TLYL VN++DR+LS ++ R KLQL+G A++++ASKYEEI
Sbjct: 202 ILVDWLVEVGEEYKLRTETLYLAVNFLDRFLSCMSVLRGKLQLVGTAAILLASKYEEIYP 261
Query: 338 PQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQ 397
P V+EF YITD+TY K ++L+ME +L L F++T PT FL +++R QGV ++
Sbjct: 262 PDVDEFVYITDDTYTKRQLLRMEHLLLKVLAFDLTVPTTNQFLLQYLR-RQGV----CIR 316
Query: 398 LESLTNYIAELSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSD 456
E+L Y+AELSL+E L Y PSLVAA+A LA +I+ W TL +T Y ++
Sbjct: 317 TENLAKYVAELSLLEADPFLKYLPSLVAAAAYCLANYIVNRHF--WPETLAAFTGYSLNE 374
Query: 457 LCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIP 501
+ C+ ELH+ + P+ AI+EKY KY +V+ P +P
Sbjct: 375 IVPCLSELHKACLSIPHRPQQAIREKYKASKYLHVSLMEPPVVLP 419
>CGA2_XENLA (P47827) Cyclin A2
Length = 415
Score = 201 bits (512), Expect = 3e-51
Identities = 130/353 (36%), Positives = 198/353 (55%), Gaps = 35/353 (9%)
Query: 171 KSPAEIEYVDNSDVSAVD--SIERKTFSILNISDAKEQAGNICSRDIL------------ 216
K PA +VD D + ++ +KT N+ G+I +R L
Sbjct: 78 KQPAFTIHVDEPDCATNKRKAVHKKTVQDENLQQLNSVLGSIGTRKPLHPIQIAMETSFG 137
Query: 217 ----VEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDIN 272
V +++ +K+V +N A +A +I+ +LR E K +P +M+K Q DI
Sbjct: 138 SPMDVSIVDEEQKVVGCNN-------VADYAKEIHTYLREMEVKCKPKAGYMQK-QPDIT 189
Query: 273 TSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKY 332
+MRAIL+DWLVEV EEY+L +TLYL VNYIDR+LS ++ R KLQL+G A+M++ASK+
Sbjct: 190 GNMRAILVDWLVEVGEEYKLQNETLYLAVNYIDRFLSSMSVLRGKLQLVGTAAMLLASKF 249
Query: 333 EEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEE 392
EEI P+V EF YITD+TY K++VL+ME VL L F++ APTI +L ++ +
Sbjct: 250 EEIYPPEVAEFVYITDDTYTKKQVLKMEHLVLKVLSFDLAAPTILQYLNQYFQI-----H 304
Query: 393 VPSLQLESLTNYIAELSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTL 451
S ++ESL+ ++ ELSL++ L Y PS+VAA+A +A + + + WS L YT
Sbjct: 305 PVSPKVESLSMFLGELSLVDADPFLRYLPSVVAAAAFVIANCTI--NERTWSDPLVEYTS 362
Query: 452 YQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEF 504
Y L C+ +L++ + ++ + A++EKY K + PP S F
Sbjct: 363 YTLETLKPCILDLYQTYLSAASHQQQAVREKYKAPK-NHAVSLIIPPESMSTF 414
>CGA2_HUMAN (P20248) Cyclin A2 (Cyclin A)
Length = 432
Score = 197 bits (501), Expect = 5e-50
Identities = 128/348 (36%), Positives = 196/348 (55%), Gaps = 27/348 (7%)
Query: 152 KSDGCSVSMDESMSSCDSFKSPAEIEYVDNSDVSAVDSI-----ERKTFSILNIS-DAKE 205
K ++ +DE+ K PAE + ++ D A +S RK L+ D
Sbjct: 95 KQPAFTIHVDEAEKEAQ--KKPAESQKIEREDALAFNSAISLPGPRKPLVPLDYPMDGSF 152
Query: 206 QAGNICSRDILVEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFME 265
++ + I++E+ EK + + + H D I+ +LR E K +P +M+
Sbjct: 153 ESPHTMDMSIVLED-EKPVSVNEVPDYHED----------IHTYLREMEVKCKPKVGYMK 201
Query: 266 KVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVAS 325
K Q DI SMRAIL+DWLVEV EEY+L +TL+L VNYIDR+LS ++ R KLQL+G A+
Sbjct: 202 K-QPDITNSMRAILVDWLVEVGEEYKLQNETLHLAVNYIDRFLSSMSVLRGKLQLVGTAA 260
Query: 326 MMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVR 385
M++ASK+EEI P+V EF YITD+TY K++VL+ME VL L F++ APT+ FL ++
Sbjct: 261 MLLASKFEEIYPPEVAEFVYITDDTYTKKQVLRMEHLVLKVLTFDLAAPTVNQFLTQYFL 320
Query: 386 AAQGVEEVPSLQLESLTNYIAELSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSS 444
Q + ++ESL ++ ELSL++ L Y PS++A +A LA + + + + W
Sbjct: 321 HQQPA----NCKVESLAMFLGELSLIDADPYLKYLPSVIAGAAFHLALYTV--TGQSWPE 374
Query: 445 TLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
+L T Y L C+ +LH+ + +P +I+EKY KY V+
Sbjct: 375 SLIRKTGYTLESLKPCLMDLHQTYLKAPQHAQQSIREKYKNSKYHGVS 422
>CG22_ANTMA (P34801) G2/mitotic-specific cyclin 2
Length = 441
Score = 197 bits (501), Expect = 5e-50
Identities = 126/375 (33%), Positives = 214/375 (56%), Gaps = 20/375 (5%)
Query: 114 VATATASFTERNEDDAAAAASGARVSVPVSTSMDVSPCKSDGCSVSMDESMS-SCDSFKS 172
+A A A+ E N++ A A GA ++P+ ++ P + E + S D+ K
Sbjct: 69 IANAEAAAAENNKNSLAVNAKGADGALPIKRAVARVPVQKKTVKSKPQEIIEISPDTEKK 128
Query: 173 PAEIEYVDNSDVSAVDSIERKTFSILNISDAKEQAGNICSRDILVEELEKGEKIVNIDND 232
A + +++ S+++K ++ + A+ +A ++ + E+IV+ID
Sbjct: 129 KAPVL---EKEITGEKSLKKKAPTLTSTLTARSKAASVV-------RTKPKEQIVDIDAA 178
Query: 233 HMDPQLCA-SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYR 291
++ L + D+YK +++E RP D+M+ Q +IN MRAILIDWLV+V ++
Sbjct: 179 DVNNDLAVVEYVEDMYKFYKSAENDSRPH-DYMDS-QPEINEKMRAILIDWLVQVHYKFE 236
Query: 292 LVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTY 351
L P+TLYLT+N +DRYL+ SR++LQLLG++SM+IASKYEEI AP+V + I+D +Y
Sbjct: 237 LSPETLYLTINIVDRYLASKTTSRRELQLLGMSSMLIASKYEEIWAPEVNDLVCISDGSY 296
Query: 352 FKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLM 411
E+VL+ME ++L L++ +T PT FL RF++A+ +V +++ ++AEL +M
Sbjct: 297 SNEQVLRMEKKILGALEWYLTVPTPYVFLVRFIKASLPDSDVE----KNMVYFLAELGMM 352
Query: 412 EY-SMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCN 470
Y +++ Y PS++AA+A++ A+ L + W+ TL+ +T + L C K L
Sbjct: 353 NYATIIMYCPSMIAAAAVYAARCTL-NKMPIWNETLRMHTGFSEVQLMDCAKLLIDFHGG 411
Query: 471 SPNSNLPAIKEKYSQ 485
S + L I KYS+
Sbjct: 412 STDQKLQGIYRKYSR 426
>CG2A_SPISO (P04962) G2/mitotic-specific cyclin A
Length = 422
Score = 197 bits (500), Expect = 7e-50
Identities = 109/249 (43%), Positives = 160/249 (63%), Gaps = 9/249 (3%)
Query: 245 DIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYI 304
DIY +LR +E K R +M++ Q DI TSMR IL+DWLVEV+EE +L +TL+L VNYI
Sbjct: 166 DIYNYLRQAEMKNRAKPGYMKR-QTDITTSMRCILVDWLVEVSEEDKLHRETLFLGVNYI 224
Query: 305 DRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVL 364
DR+LS ++ R KLQL+G ASM +A+KYEEI P V+EF YITD+TY ++VL+ME +L
Sbjct: 225 DRFLSKISVLRGKLQLVGAASMFLAAKYEEIYPPDVKEFAYITDDTYTSQQVLRMEHLIL 284
Query: 365 NFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEY-SMLCYAPSLV 423
L F++ PT F F+++ + +L+SLT ++ EL+L++ + L Y PS+
Sbjct: 285 KVLTFDVAVPTTNWFCEDFLKSCDADD-----KLKSLTMFLTELTLIDMDAYLKYLPSIT 339
Query: 424 AASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKY 483
AA+A+ LA++ L I+PW L T Y+ C+K+LH+ + + A++EKY
Sbjct: 340 AAAALCLARYSL--GIEPWPQNLVKKTGYEIGHFVDCLKDLHKTSLGAESHQQQAVQEKY 397
Query: 484 SQHKYKYVA 492
Q KY V+
Sbjct: 398 KQDKYHQVS 406
>CGA2_MOUSE (P51943) Cyclin A2 (Cyclin A)
Length = 422
Score = 196 bits (498), Expect = 1e-49
Identities = 127/348 (36%), Positives = 199/348 (56%), Gaps = 29/348 (8%)
Query: 152 KSDGCSVSMDESMSSCDSFKSPAEIEYVDNSDVSAVDSI-----ERKTFSILNIS-DAKE 205
K ++ +DE+ ++ K PAE++ + D A ++ RK + L+ D
Sbjct: 87 KQPAFTIHVDEAE---ETQKRPAELKETECEDALAFNAAVSLPAARKPLTPLDYPMDGSF 143
Query: 206 QAGNICSRDILVEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFME 265
++ + I++E+ K VN++ + DI+ +LR E K +P +M+
Sbjct: 144 ESPHAMDMSIVLED-----KPVNVNE-------VPDYQEDIHTYLREMEVKCKPKVGYMK 191
Query: 266 KVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVAS 325
+ Q DI SMRAIL+DWLVEV EEY+L +TL+L VNYIDR+LS ++ R KLQL+G A+
Sbjct: 192 R-QPDITNSMRAILVDWLVEVGEEYKLQNETLHLAVNYIDRFLSSMSVLRGKLQLVGTAA 250
Query: 326 MMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVR 385
M++ASK+EEI P+V EF YITD+TY K++VL+ME VL L F++ APT+ FL ++
Sbjct: 251 MLLASKFEEIYPPEVAEFVYITDDTYSKKQVLRMEHLVLKVLAFDLAAPTVNQFLTQYFL 310
Query: 386 AAQGVEEVPSLQLESLTNYIAELSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSS 444
Q + ++ESL ++ ELSL++ L Y PSL+A +A LA + + + + W
Sbjct: 311 HLQPA----NCKVESLAMFLGELSLIDADPYLKYLPSLIAGAAFHLALYTV--TGQSWPE 364
Query: 445 TLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
+L T Y L C+ +LH+ + +P +I+EKY KY V+
Sbjct: 365 SLAQQTGYTLESLKPCLVDLHQTYLKAPQHAQQSIREKYKHSKYHSVS 412
>CGA2_BOVIN (P30274) Cyclin A2 (Cyclin A) (Fragment)
Length = 406
Score = 196 bits (498), Expect = 1e-49
Identities = 126/329 (38%), Positives = 191/329 (57%), Gaps = 25/329 (7%)
Query: 171 KSPAEIEYVDNSDVSAVDSI-----ERKTFSILNIS-DAKEQAGNICSRDILVEELEKGE 224
K P E + ++ DV A +S RK + L+ D ++ + +++E+ EK
Sbjct: 86 KRPTESKKSESEDVLAFNSAVTLPGPRKPLAPLDYPMDGSFESPHTMEMSVVLED-EKPV 144
Query: 225 KIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLV 284
+ + + H D I+ +LR E K +P +M+K Q DI SMRAIL+DWLV
Sbjct: 145 SVNEVPDYHED----------IHTYLREMEVKCKPKVGYMKK-QPDITNSMRAILVDWLV 193
Query: 285 EVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFC 344
EV EEY+L +TL+L VNYIDR+LS ++ R KLQL+G A+M++ASK+EEI P+V EF
Sbjct: 194 EVGEEYKLQNETLHLAVNYIDRFLSSMSVLRGKLQLVGTAAMLLASKFEEIYPPEVAEFV 253
Query: 345 YITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNY 404
YITD+TY K++VL+ME VL L F++ APTI FL ++ Q + ++ESL +
Sbjct: 254 YITDDTYTKKQVLRMEHLVLKVLAFDLAAPTINQFLTQYFLHQQPA----NCKVESLAMF 309
Query: 405 IAELSLMEYS-MLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKE 463
+ ELSL++ L Y PS++AA+A LA + + + + W +L T Y L C+ +
Sbjct: 310 LGELSLIDADPYLKYLPSVIAAAAFHLALYTV--TGQSWPESLVQKTGYTLETLKPCLLD 367
Query: 464 LHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
LH+ + +P +I+EKY KY V+
Sbjct: 368 LHQTYLRAPQHAQQSIREKYKNSKYHGVS 396
>CGA2_CHICK (P43449) Cyclin A2 (Cyclin A)
Length = 395
Score = 195 bits (495), Expect = 3e-49
Identities = 108/260 (41%), Positives = 165/260 (62%), Gaps = 9/260 (3%)
Query: 242 FACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTV 301
+ DI+ +LR E K +P +M+K Q DI +MRAIL+DWLVEV EEY+L +TL+L V
Sbjct: 142 YVSDIHTYLREMEVKCKPKIGYMKK-QPDITNNMRAILVDWLVEVGEEYKLQNETLHLAV 200
Query: 302 NYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMES 361
NYIDR+LS ++ R KLQL+G A+M++ASK+EEI P+V EF YITD+TY K++VL+ME
Sbjct: 201 NYIDRFLSSMSVLRGKLQLVGTAAMLLASKFEEIYPPEVAEFVYITDDTYNKKQVLRMEH 260
Query: 362 EVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYS-MLCYAP 420
+L L F++ APTI FL ++ + + + ++ESL+ Y+ EL+L++ L Y P
Sbjct: 261 LILKVLSFDLAAPTINQFLTQYF-----LHQQTNAKVESLSMYLGELTLIDADPYLKYLP 315
Query: 421 SLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIK 480
S++AA+A LA + + + + W +L T Y + C+ +LHR + + +I+
Sbjct: 316 SVIAAAAFHLASYTI--TGQTWPESLCKVTGYTLEHIKPCLMDLHRTYLKAAQHTQQSIR 373
Query: 481 EKYSQHKYKYVAKKYCPPSI 500
EKY KY V+ P ++
Sbjct: 374 EKYKSTKYHAVSLIDAPETL 393
>CG2B_SPISO (P13952) G2/mitotic-specific cyclin B
Length = 428
Score = 193 bits (491), Expect = 8e-49
Identities = 112/269 (41%), Positives = 159/269 (58%), Gaps = 10/269 (3%)
Query: 226 IVNID-NDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLV 284
+ NID ND +PQL + + DIY ++R E K +++E ++I MRAILIDWL
Sbjct: 152 VQNIDANDKENPQLVSEYVNDIYDYMRDLEGKYPIRHNYLEN--QEITGKMRAILIDWLC 209
Query: 285 EVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFC 344
+V + L+ +TLYLTV IDR L + + R KLQL+GV SM+IASKYEE+ AP+V +F
Sbjct: 210 QVHHRFHLLQETLYLTVAIIDRLLQESPVPRNKLQLVGVTSMLIASKYEEMYAPEVADFV 269
Query: 345 YITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNY 404
YITDN Y K+E+L+ME +L L F P FLRR +A Q +L Y
Sbjct: 270 YITDNAYTKKEILEMEQHILKKLNFSFGRPLCLHFLRRDSKAGQ-----VDANKHTLAKY 324
Query: 405 IAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKEL 464
+ EL++ EY M+ Y PS +AA+A+ L+ +L W+ TL HY+ Y DL +++L
Sbjct: 325 LMELTITEYDMVQYLPSKIAAAALCLSMKLL--DSTHWTETLTHYSSYCEKDLVSTMQKL 382
Query: 465 HRLFCNSPNSNLPAIKEKYSQHKYKYVAK 493
L + NS L A+ KYS K+ ++K
Sbjct: 383 ASLVIKAENSKLTAVHTKYSSSKFMKISK 411
>CG2B_MARGL (P15206) G2/mitotic-specific cyclin B
Length = 388
Score = 193 bits (491), Expect = 8e-49
Identities = 108/270 (40%), Positives = 168/270 (62%), Gaps = 9/270 (3%)
Query: 224 EKIVNID-NDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDW 282
E + +ID ND +PQLC+ F DIY+++R E + + TD+M ++I MR+ILIDW
Sbjct: 109 EGVEDIDKNDFDNPQLCSEFVNDIYQYMRKLEREFKVRTDYM--TIQEITERMRSILIDW 166
Query: 283 LVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEE 342
LV+V + L+ +TL+LT+ +DRYL +S+ KLQL+GV SM+IA+KYEE+ P++ +
Sbjct: 167 LVQVHLRFHLLQETLFLTIQILDRYLEVQPVSKNKLQLVGVTSMLIAAKYEEMYPPEIGD 226
Query: 343 FCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLT 402
F YITDN Y K ++ ME +L L F + P FLRR +A GV+ Q ++
Sbjct: 227 FVYITDNAYTKAQIRSMECNILRRLDFSLGKPLCIHFLRRNSKAG-GVDG----QKHTMA 281
Query: 403 NYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVK 462
Y+ EL+L EY+ + Y PS +AA+A+ L+ IL P ++ W +TL HY+ Y L V+
Sbjct: 282 KYLMELTLPEYAFVPYDPSEIAAAALCLSSKILEPDME-WGTTLVHYSAYSEDHLMPIVQ 340
Query: 463 ELHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
++ + N+P + A+++KYS K+ V+
Sbjct: 341 KMALVLKNAPTAKFQAVRKKYSSAKFMNVS 370
>CG1B_MEDVA (P46277) G2/mitotic-specific cyclin 1 (B-like cyclin)
(CycMs1)
Length = 428
Score = 193 bits (491), Expect = 8e-49
Identities = 114/302 (37%), Positives = 178/302 (58%), Gaps = 14/302 (4%)
Query: 203 AKEQAGNICSRDILVEELEKG----EKIVNIDN-DHMDPQLCASFACDIYKHLRASEAKK 257
A EQ + S +EE+E E +++ID D DP A + D+Y + R E+
Sbjct: 128 ALEQTEPMHSESDQMEEVEMEDIMEEPVMDIDTPDANDPLAVAEYIEDLYSYYRKVESTS 187
Query: 258 RPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQK 317
S ++M + Q DIN MRAIL+DWL+EV +++ L+ +TL+LTVN IDR+L ++ R+K
Sbjct: 188 CVSPNYMAQ-QFDINERMRAILVDWLIEVHDKFDLMHETLFLTVNLIDRFLEKQSVVRKK 246
Query: 318 LQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIK 377
LQL+G+ +M++A KYEE+ P V + I+D Y ++EVL+ME ++N LKF ++ PT
Sbjct: 247 LQLVGLVAMLLACKYEEVSVPVVGDLILISDRAYTRKEVLEMEKVMVNALKFNISVPTAY 306
Query: 378 CFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFP 437
F+RRF++AAQ +LE L ++ ELSL+EY+ML ++PS +AA+A++ A+ ++
Sbjct: 307 VFMRRFLKAAQA-----DRKLELLAFFLIELSLVEYAMLKFSPSQLAAAAVYTAQCTMY- 360
Query: 438 SIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCP 497
+K WS T + +T Y L C + + L KY K+ Y AK C
Sbjct: 361 GVKQWSKTCEWHTNYSEDQLLECSSLMVDFHKKAGTGKLTGAHRKYCTSKFSYTAK--CE 418
Query: 498 PS 499
P+
Sbjct: 419 PA 420
>CGA2_MESAU (P37881) Cyclin A2 (Cyclin A)
Length = 421
Score = 192 bits (489), Expect = 1e-48
Identities = 109/249 (43%), Positives = 158/249 (62%), Gaps = 8/249 (3%)
Query: 245 DIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYI 304
DI+ +LR E K +P +M+K Q DI SMRAIL+DWLVEV EEY+L +TL+L VNYI
Sbjct: 170 DIHTYLREMEIKCKPKVGYMKK-QPDITNSMRAILVDWLVEVGEEYKLQNETLHLAVNYI 228
Query: 305 DRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVL 364
DR+LS ++ R KLQL+G A+M++ASK+EEI P+V EF YITD+TY K++VL+ME VL
Sbjct: 229 DRFLSSMSVLRGKLQLVGTAAMLLASKFEEIYPPEVAEFVYITDDTYSKKQVLRMEHLVL 288
Query: 365 NFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYS-MLCYAPSLV 423
L F++ APT+ FL ++ Q + ++ESL ++ ELSL++ L Y PSL+
Sbjct: 289 KVLAFDLAAPTVNQFLNQYFLHQQPA----NCKVESLAMFLGELSLIDADPYLKYLPSLI 344
Query: 424 AASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKY 483
A +A LA + + + + W +L T Y L C+ +LH+ + + +I+EKY
Sbjct: 345 AGAAFHLALYTV--TGQSWPESLVQKTGYTLESLKPCLMDLHQTYLRAAQHTQQSIREKY 402
Query: 484 SQHKYKYVA 492
KY V+
Sbjct: 403 KHSKYHGVS 411
>CG2A_DROME (P14785) G2/mitotic-specific cyclin A
Length = 491
Score = 191 bits (486), Expect = 3e-48
Identities = 121/312 (38%), Positives = 180/312 (56%), Gaps = 16/312 (5%)
Query: 182 SDVSAVDSIERKTFSILNISDAKEQAGNICSRDILVEELEKGEKIVNIDNDHMDPQLCAS 241
+DV + S++R ++ SD S V+EL ND
Sbjct: 150 TDVQSPMSVDRSILGVIQSSDISVGTETGVSPTGRVKELPPR-------NDRQRFLEVVQ 202
Query: 242 FACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTV 301
+ DI ++ R SE K RP +M + QKDI+ +MR+ILIDWLVEV+EEY+L +TLYL+V
Sbjct: 203 YQMDILEYFRESEKKHRPKPLYMRR-QKDISHNMRSILIDWLVEVSEEYKLDTETLYLSV 261
Query: 302 NYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMES 361
Y+DR+LS A+ R KLQL+G A+M IA+KYEEI P+V EF ++TD++Y K +VL+ME
Sbjct: 262 FYLDRFLSQMAVVRSKLQLVGTAAMYIAAKYEEIYPPEVGEFVFLTDDSYTKAQVLRMEQ 321
Query: 362 EVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLME-YSMLCYAP 420
+L L F++ PT F+ + E +L+ +T YI+ELSLME + L Y P
Sbjct: 322 VILKILSFDLCTPTAYVFINTYAVLCDMPE-----KLKYMTLYISELSLMEGETYLQYLP 376
Query: 421 SLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIK 480
SL++++++ LA+ IL ++ W+ L+ T Y+ DL V L + N A++
Sbjct: 377 SLMSSASVALARHIL--GMEMWTPRLEEITTYKLEDLKTVVLHLCHTHKTAKELNTQAMR 434
Query: 481 EKYSQHKYKYVA 492
EKY++ YK VA
Sbjct: 435 EKYNRDTYKKVA 446
>CGA1_CARAU (Q92161) Cyclin A1 (Cyclin A)
Length = 391
Score = 191 bits (485), Expect = 4e-48
Identities = 109/253 (43%), Positives = 156/253 (61%), Gaps = 10/253 (3%)
Query: 238 LCA-SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDT 296
LC +A DI+++LR E K RP +M K Q DI MR IL+DWLVEV EEY+L +T
Sbjct: 132 LCVPEYAEDIHRYLRECEVKYRPKPGYMRK-QPDITNCMRVILVDWLVEVGEEYKLCSET 190
Query: 297 LYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEV 356
L+L VNY+DR+LS ++ R KLQL+G A++++A+KYEE+ P+V+EF YITD+TY K+++
Sbjct: 191 LFLAVNYLDRFLSCMSVLRGKLQLVGTAAVLLAAKYEEVYPPEVDEFVYITDDTYTKKQL 250
Query: 357 LQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYS-M 415
L+ME +L L F+MTAPT+ FL ++ +L L Y++ELSL+E
Sbjct: 251 LRMEQHLLRVLAFDMTAPTVHQFLMQYTLEGHICARTVNLAL-----YLSELSLLEVDPF 305
Query: 416 LCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSN 475
+ Y PS AA+A LA + L + W L +T Y + + C+ ELH+L +
Sbjct: 306 VQYLPSKTAAAAYCLANYTLNGVL--WPENLYAFTGYSLAVIIPCLMELHKLHLGAAGRP 363
Query: 476 LPAIKEKYSQHKY 488
AI+EKY KY
Sbjct: 364 QQAIQEKYKGSKY 376
>CG2B_ARBPU (P07818) G2/mitotic-specific cyclin B
Length = 409
Score = 189 bits (481), Expect = 1e-47
Identities = 105/272 (38%), Positives = 169/272 (61%), Gaps = 9/272 (3%)
Query: 225 KIVNIDNDHMD-PQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWL 283
++ +ID D D PQLC+ +A DIY +LR E + +++++ + I MR IL+DWL
Sbjct: 124 QVEDIDKDDGDNPQLCSEYAKDIYLYLRRLEVEMMVPANYLDRQETQITGRMRLILVDWL 183
Query: 284 VEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEF 343
V+V + L+ +TL+LTV IDR+L+ +++S+ KLQL+GV +M IASKYEE+ P++ +F
Sbjct: 184 VQVHLRFHLLQETLFLTVQLIDRFLAEHSVSKGKLQLVGVTAMFIASKYEEMYPPEINDF 243
Query: 344 CYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTN 403
YITDN Y K ++ QME +L LK+++ P FLRR +AA GV+ Q +L
Sbjct: 244 VYITDNAYTKAQIRQMEIAMLKGLKYKLGKPLCLHFLRRNSKAA-GVD----AQKHTLAK 298
Query: 404 YIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSI-KPWSSTLQHYTLYQPSDLCVCVK 462
Y+ E++L EYSM+ Y+PS +AA+AI+L+ +L P W + HY++Y L V+
Sbjct: 299 YLMEITLPEYSMVQYSPSEIAAAAIYLSMTLLDPETHSSWCPKMTHYSMYSEDHLRPIVQ 358
Query: 463 ELHRLFC--NSPNSNLPAIKEKYSQHKYKYVA 492
++ ++ +S + A+K KY K+ ++
Sbjct: 359 KIVQILLRDDSASQKYSAVKTKYGSSKFMKIS 390
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.315 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 55,290,509
Number of Sequences: 164201
Number of extensions: 2204924
Number of successful extensions: 13758
Number of sequences better than 10.0: 310
Number of HSP's better than 10.0 without gapping: 194
Number of HSP's successfully gapped in prelim test: 121
Number of HSP's that attempted gapping in prelim test: 10268
Number of HSP's gapped (non-prelim): 1480
length of query: 507
length of database: 59,974,054
effective HSP length: 115
effective length of query: 392
effective length of database: 41,090,939
effective search space: 16107648088
effective search space used: 16107648088
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0181.13


