

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.8
(488 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
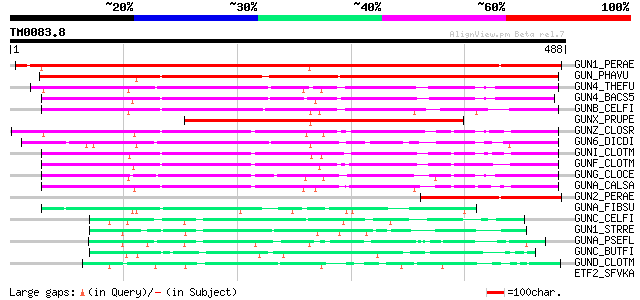
Score E
Sequences producing significant alignments: (bits) Value
GUN1_PERAE (P05522) Endoglucanase 1 precursor (EC 3.2.1.4) (Endo... 565 e-160
GUN_PHAVU (P22503) Endoglucanase precursor (EC 3.2.1.4) (Endo-1,... 491 e-138
GUN4_THEFU (P26221) Endoglucanase E-4 precursor (EC 3.2.1.4) (En... 296 8e-80
GUN4_BACS5 (P28622) Endoglucanase 4 precursor (EC 3.2.1.4) (Endo... 285 2e-76
GUNB_CELFI (P26225) Endoglucanase B precursor (EC 3.2.1.4) (Endo... 278 2e-74
GUNX_PRUPE (P38534) Endoglucanase CX (EC 3.2.1.4) (Endo-1,4-beta... 273 7e-73
GUNZ_CLOSR (P23659) Endoglucanase Z precursor (EC 3.2.1.4) (Endo... 271 4e-72
GUN6_DICDI (P22699) Endoglucanase precursor (EC 3.2.1.4) (Endo-1... 258 2e-68
GUNI_CLOTM (Q02934) Endoglucanase I precursor (EC 3.2.1.4) (EGI)... 256 1e-67
GUNF_CLOTM (P26224) Endoglucanase F precursor (EC 3.2.1.4) (EGF)... 254 5e-67
GUNG_CLOCE (P37700) Endoglucanase G precursor (EC 3.2.1.4) (Endo... 248 3e-65
GUNA_CALSA (P22534) Endoglucanase A precursor (EC 3.2.1.4) (Endo... 246 1e-64
GUN2_PERAE (P23666) Endoglucanase 2 (EC 3.2.1.4) (Endo-1,4-beta-... 167 6e-41
GUNA_FIBSU (P23665) Endoglucanase A precursor (EC 3.2.1.4) (Endo... 57 1e-07
GUNC_CELFI (P14090) Endoglucanase C precursor (EC 3.2.1.4) (Endo... 54 9e-07
GUN1_STRRE (Q05156) Cellulase 1 precursor (EC 3.2.1.4) (Endogluc... 52 3e-06
GUNA_PSEFL (P10476) Endoglucanase A precursor (EC 3.2.1.4) (Endo... 49 2e-05
GUNC_BUTFI (P23658) Cellodextrinase (EC 3.2.1.-) 47 1e-04
GUND_CLOTM (P04954) Endoglucanase D precursor (EC 3.2.1.4) (EGD)... 46 2e-04
ETF2_SFVKA (Q9Q8Y2) Early transcription factor 82 kDa subunit (V... 31 6.3
>GUN1_PERAE (P05522) Endoglucanase 1 precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Abscission cellulase 1)
Length = 494
Score = 565 bits (1455), Expect = e-160
Identities = 283/488 (57%), Positives = 351/488 (70%), Gaps = 11/488 (2%)
Query: 6 SSFMSLLFLIPMLLFVGNVQC----NPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDS 61
SS +SL L+ L+ V+C + +Y +AL KS+LF +GQRSG+LP +Q+L WR DS
Sbjct: 4 SSPLSLFHLL--LVCTVMVKCCSASDLHYSDALEKSILFFEGQRSGKLPTNQRLTWRGDS 61
Query: 62 GLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRW 121
GL DG +VDL GGYYDAGDN+KF PMAFTTTML+W IE+G M Q++ ARAA+RW
Sbjct: 62 GLSDGSSYHVDLVGGYYDAGDNLKFGLPMAFTTTMLAWGIIEFGCLMPEQVENARAALRW 121
Query: 122 GTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAA 181
TDYLLK +T+T LYV VG+PN DH+CWERPEDMDT R VY VS NPGSDVAAETAA
Sbjct: 122 STDYLLKASTATSNSLYVQVGEPNADHRCWERPEDMDTPRNVYKVSTQNPGSDVAAETAA 181
Query: 182 ALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDEL 241
ALAAAS+VF D +YS L+ TA KV++FA QY+GSYSDSLGS VCPFYCSYSG+ DEL
Sbjct: 182 ALAAASIVFGDSDSSYSTKLLHTAVKVFEFADQYRGSYSDSLGSVVCPFYCSYSGYNDEL 241
Query: 242 LWGAAWLFRATNAVKYYNLIK----TLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKN 297
LWGA+WL RA+ Y I+ TLGADD FSWDDK G VLLS+ L + +
Sbjct: 242 LWGASWLHRASQNASYMTYIQSNGHTLGADDDDYSFSWDDKRVGTKVLLSKGFLQDRIEE 301
Query: 298 FDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAK 357
Y+ +N++C ++P + S QYT GGL++K SNLQYVTS FLL TYA Y+++
Sbjct: 302 LQLYKVHTDNYICSLIPGTSSFQAQYTPGGLLYKGSASNLQYVTSTAFLLLTYANYLNSS 361
Query: 358 KYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLA 417
+CG VT L S+AK+QVDYILG+NP +MSYMVG+G +P+ +HHRGSSLPS+
Sbjct: 362 GGHASCGTTTVTAKNLISLAKKQVDYILGQNPAKMSYMVGFGERYPQHVHHRGSSLPSVQ 421
Query: 418 AHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAV 477
HP +I C+ GFQ + +SS PNPNILVGAI+GGP+ D F DDR++Y SEP TYIN +
Sbjct: 422 VHPNSIPCNAGFQ-YLYSSPPNPNILVGAILGGPDNRDSFSDDRNNYQQSEPATYINAPL 480
Query: 478 VGPLAYFA 485
VG LA+FA
Sbjct: 481 VGALAFFA 488
>GUN_PHAVU (P22503) Endoglucanase precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Abscission cellulase)
Length = 496
Score = 491 bits (1263), Expect = e-138
Identities = 238/460 (51%), Positives = 324/460 (69%), Gaps = 11/460 (2%)
Query: 27 NPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKF 86
N +Y +ALAK++LF +GQRSG+LP Q++KWR DS L DG+L NV+L GGYYDAGDNVKF
Sbjct: 38 NYDYADALAKAILFFEGQRSGKLPSSQRVKWREDSALSDGKLQNVNLMGGYYDAGDNVKF 97
Query: 87 NFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDP 144
+PMAF+T++LSW+A+EY + QL ++AIRWG D++L+ TS P LY VGD
Sbjct: 98 GWPMAFSTSLLSWAAVEYESEISSVNQLGYLQSAIRWGADFMLRAHTS-PTTLYTQVGDG 156
Query: 145 NVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRT 204
N DH CWERPEDMDT RTVY + N+PG++VAAE AAAL+AAS+VFKK+D YS L+
Sbjct: 157 NADHNCWERPEDMDTPRTVYKIDANSPGTEVAAEYAAALSAASIVFKKIDAKYSSTLLSH 216
Query: 205 AQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYN-LIKT 263
++ ++ FA + +GSYS S CPFYCSYSG++DELLW AAWL++A+ KY + +I
Sbjct: 217 SKSLFDFADKNRGSYSGS-----CPFYCSYSGYQDELLWAAAWLYKASGESKYLSYIISN 271
Query: 264 LGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQY 323
G FSWD+K+ GA LL+ G K+ + + +AE+F+C ++P S S +
Sbjct: 272 QGWSQTVSEFSWDNKFVGAQTLLTEE-FYGGKKDLAKIKTDAESFICAVMPGSNSRQIKT 330
Query: 324 TQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYT-FNCGNVLVTPNTLRSVAKRQVD 382
T GGL+F SNLQY TS T +L +++ ++ NCG+ T + +R AK QV+
Sbjct: 331 TPGGLLFTRDSSNLQYTTSSTMVLFIFSRILNRNHINGINCGSSHFTASQIRGFAKTQVE 390
Query: 383 YILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNI 442
YILG+NP++MSYMVG+G +PK++HHRGSS+PS+ HP +GC+ G +++S+NPNPN
Sbjct: 391 YILGKNPMKMSYMVGFGSKYPKQLHHRGSSIPSIKVHPAKVGCNAGLSDYYNSANPNPNT 450
Query: 443 LVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
VGAIVGGP+ +D F D RSDYSH+EPTTYIN A V ++
Sbjct: 451 HVGAIVGGPDSNDRFNDARSDYSHAEPTTYINAAFVASIS 490
>GUN4_THEFU (P26221) Endoglucanase E-4 precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase E-4) (Cellulase E-4) (Cellulase
E4)
Length = 880
Score = 296 bits (758), Expect = 8e-80
Identities = 186/482 (38%), Positives = 255/482 (52%), Gaps = 54/482 (11%)
Query: 19 LFVGNVQCNP--NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGG 76
L G P NY EAL KS+ F + QRSG+LP + ++ WR DSGL DG +DL+GG
Sbjct: 39 LATGTAHAEPAFNYAEALQKSMFFYEAQRSGKLPENNRVSWRGDSGLNDGADVGLDLTGG 98
Query: 77 YYDAGDNVKFNFPMAFTTTMLSWSAIE--YGKRMGPQLKEARAAIRWGTDYLLKCATSTP 134
+YDAGD+VKF FPMAFT TML+W AIE G Q+ + +RW DY +K A +P
Sbjct: 99 WYDAGDHVKFGFPMAFTATMLAWGAIESPEGYIRSGQMPYLKDNLRWVNDYFIK-AHPSP 157
Query: 135 GRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVD 194
LYV VGD + DHK W E M R + V P+ PGSDVAAETAAA+AA+S+VF D
Sbjct: 158 NVLYVQVGDGDADHKWWGPAEVMPMERPSFKVDPSCPGSDVAAETAAAMAASSIVFADDD 217
Query: 195 PTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNA 254
P Y+ LV+ A+++Y FA Y+G YSD + + FY S+SG++DEL+WGA WL++AT
Sbjct: 218 PAYAATLVQHAKQLYTFADTYRGVYSDCVPAGA--FYNSWSGYQDELVWGAYWLYKATGD 275
Query: 255 VKY-------YNLIKTLGADDQPDI------FSWDDKYAGAHVLLSRRALLNGDKNFDQY 301
Y Y+ + T + Q D+ +WDDK G +VLL++ + +Y
Sbjct: 276 DSYLAKAEYEYDFLST---EQQTDLRSYRWTIAWDDKSYGTYVLLAK------ETGKQKY 326
Query: 302 RQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTF 361
+A ++ Y+ GG+ L+Y + F+ YAK +
Sbjct: 327 IDDANRWLDYWTVGVNGQRVPYSPGGMAVLDTWGALRYAANTAFVALVYAKVIDDP---- 382
Query: 362 NCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQ 421
V A RQ++Y LG+NP SY+VG+G N P+ HHR AH
Sbjct: 383 ------VRKQRYHDFAVRQINYALGDNPRNSSYVVGFGNNPPRNPHHR-------TAH-- 427
Query: 422 TIGCDGGFQPFFHSSNPNPNILVGAIVGGP-NQSDGFPDDRSDYSHSEPTTYINGAVVGP 480
G + S N ++L GA+VGGP + +D + DDR DY +E T N
Sbjct: 428 -----GSWTDSIASPAENRHVLYGALVGGPGSPNDAYTDDRQDYVANEVATDYNAGFSSA 482
Query: 481 LA 482
LA
Sbjct: 483 LA 484
>GUN4_BACS5 (P28622) Endoglucanase 4 precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase 4) (Cellulase 4) (EG-IV)
Length = 636
Score = 285 bits (729), Expect = 2e-76
Identities = 173/461 (37%), Positives = 247/461 (53%), Gaps = 41/461 (8%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NY E L KS+LF + QRSG+LP +L WR DSGL DG+ DL+GG+YDAGD+VKF
Sbjct: 29 NYVEVLQKSILFYEAQRSGKLPESNRLNWRGDSGLEDGKDVGHDLTGGWYDAGDHVKFGL 88
Query: 89 PMAFTTTMLSWSAIEYGK--RMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNV 146
PMA++ +L+W+ EY + L + I+W TDY LK T P + VGD N
Sbjct: 89 PMAYSAAVLAWTVYEYREAYEEAELLDDMLDQIKWATDYFLKAHTG-PNEFWAQVGDGNA 147
Query: 147 DHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQ 206
DH W E M R + + + PG++VAA+TAAALAA S++FK+ D Y+ L+ A+
Sbjct: 148 DHGWWGPAEVMPMNRPAFKIDEHCPGTEVAAQTAAALAAGSIIFKETDAPYAAKLLTHAK 207
Query: 207 KVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKTLGA 266
++Y FA QY+G Y+D + +A PFY S+SG+ DEL+WG WL+ ATN Y N K L A
Sbjct: 208 QLYAFADQYRGEYTDCVTNAQ-PFYNSWSGYIDELIWGGIWLYLATNDQTYLN--KALKA 264
Query: 267 D-------DQPDIFSWDDKYAGAHVLLSRRALLNGDKNF-DQYRQEAENFMCKILPNSPS 318
D SWD+ + + +LL+R + +K F + + + + + N
Sbjct: 265 VEEWPKDWDYTFTMSWDNTFFLSQILLAR---ITKEKRFIESTERNLDYWSTGFVQNGKV 321
Query: 319 SSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAK 378
YT GGL + +L+Y + FL YA ++S ++ N ++ A
Sbjct: 322 ERITYTPGGLAWLDQWGSLRYTANAAFLAFVYADWVSDQE----------KKNRYQTFAI 371
Query: 379 RQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNP 438
RQ Y+LG+NP SY+VG+G N P HHR AH G + + +
Sbjct: 372 RQTHYMLGDNPQNRSYVVGFGKNPPMHPHHR-------TAH-------GSWSNQLTTPSS 417
Query: 439 NPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVG 479
+ + L G +VGGPN+ D + DD SDY +E T N A G
Sbjct: 418 HRHTLYGPLVGGPNRQDQYTDDISDYVSNEVATDYNAAFTG 458
>GUNB_CELFI (P26225) Endoglucanase B precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase B) (Cellulase B)
Length = 1045
Score = 278 bits (712), Expect = 2e-74
Identities = 178/472 (37%), Positives = 250/472 (52%), Gaps = 55/472 (11%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NY EAL KS+ F Q QRSG LP D + WR DSGL DG DL+GG+YDAGD+VKF F
Sbjct: 38 NYAEALQKSMFFYQAQRSGDLPADFPVSWRGDSGLTDGADVGKDLTGGWYDAGDHVKFGF 97
Query: 89 PMAFTTTMLSWSAIE--YGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNV 146
PMAF+ TML+W AIE G L E + +R+ +DY +K T+ P LYV VGD
Sbjct: 98 PMAFSATMLAWGAIESPTGYSKAGSLDELKDNLRFVSDYFVKAHTA-PNELYVQVGDGEA 156
Query: 147 DHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQ 206
DHK W E M AR + +S + PGSDVAAETAAALA++++V K DP Y+ LV A+
Sbjct: 157 DHKWWGPAEVMTMARPSHKISASCPGSDVAAETAAALASSAIVLKGDDPAYAATLVSHAK 216
Query: 207 KVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT--- 263
++Y FA Y+G+YSD + +A +Y S+SG++DEL+WGA WL++AT Y +
Sbjct: 217 QLYTFADTYRGAYSDCV-TAASAYYKSWSGYQDELVWGAYWLYKATGDATYLAKAEAEYD 275
Query: 264 -LGADDQPD------IFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNS 316
LG ++Q +WD+K G + LL+ + +Y +A ++
Sbjct: 276 KLGTENQSTTRSYKWTIAWDNKQFGTYALLAM------ETGKQKYVDDANRWLDYWTVGV 329
Query: 317 PSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYM--SAKKYTFNCGNVLVTPNTLR 374
Y+ GG L+Y + +F+ Y+ +M + +K ++ V
Sbjct: 330 NGQKVPYSPGGQAVLDSWGALRYAANTSFVALVYSDWMTDATRKARYHDFGV-------- 381
Query: 375 SVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHR---GSSLPSLAAHPQTIGCDGGFQP 431
RQ++Y LG+NP SY+VG+G N P HHR GS L S+ Q
Sbjct: 382 ----RQINYALGDNPRSSSYVVGFGANPPTAPHHRTAHGSWLDSITTPAQ---------- 427
Query: 432 FFHSSNPNPNILVGAIVGGP-NQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
+ ++L GA+VGGP + +D + D R DY +E T N LA
Sbjct: 428 -------SRHVLYGALVGGPGSPNDAYTDSRQDYVANEVATDYNAGFTSALA 472
>GUNX_PRUPE (P38534) Endoglucanase CX (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (CX-cellulase) (Fragment)
Length = 251
Score = 273 bits (698), Expect = 7e-73
Identities = 142/251 (56%), Positives = 172/251 (67%), Gaps = 5/251 (1%)
Query: 154 PEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFAL 213
PEDMDT R VY VS NPGSDVAAETAAALAAAS+VFK DP+YS L+ TA KV+ FA
Sbjct: 1 PEDMDTPRNVYKVSTQNPGSDVAAETAAALAAASIVFKDSDPSYSGKLLHTAMKVFDFAD 60
Query: 214 QYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----LGADDQ 269
+Y+GSYSDS+GS VCPFYCSYSG DELLWGA+W+ RA+ Y IK+ LG DD
Sbjct: 61 RYRGSYSDSIGSVVCPFYCSYSGHHDELLWGASWIHRASQNRSYLVYIKSNGHILGEDDD 120
Query: 270 PDIFSWDDKYAGAHVLLSRRALLNGD-KNFDQYRQEAENFMCKILPNSPSSSTQYTQGGL 328
SWDDK G VLL ++ L + F Y+ + N++C +LP + + QYT GGL
Sbjct: 121 GFSASWDDKETGTKVLLVSKSFLERHVEEFQLYKAHSGNYICSLLPGTSNFQGQYTPGGL 180
Query: 329 MFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGEN 388
++K +SNLQYVTS T LL TYAKY+ CG+ VT TL S AK+QVDYILG N
Sbjct: 181 LYKASESNLQYVTSTTLLLLTYAKYLRTNGGVATCGSSKVTAETLISEAKKQVDYILGNN 240
Query: 389 PLQMSYMVGYG 399
P ++SYMVG+G
Sbjct: 241 PAKISYMVGFG 251
>GUNZ_CLOSR (P23659) Endoglucanase Z precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Thermoactive cellulase)
(Avicelase I)
Length = 986
Score = 271 bits (692), Expect = 4e-72
Identities = 182/496 (36%), Positives = 261/496 (51%), Gaps = 50/496 (10%)
Query: 2 MRTSSSFMSLLFLIPMLLFVGNVQCNP---NYREALAKSLLFLQGQRSGRLPPDQQLKWR 58
MR SF ++ L+ LF+ + NY EAL K+++F + QRSG+LP +++ WR
Sbjct: 1 MRKFYSFAIIISLLVTGLFIHTPKAEAAGYNYGEALQKAIMFYEFQRSGKLPENKRDNWR 60
Query: 59 SDSGLFDGRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRM--GPQLKEAR 116
DSGL DG +DL+GG+YDAGD+VKFN PMA++ TML+W+A E + + Q+
Sbjct: 61 GDSGLNDGADVGLDLTGGWYDAGDHVKFNLPMAYSQTMLAWAAYEAEEALERSGQMGYLL 120
Query: 117 AAIRWGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVA 176
AI+W +DYL+KC S P Y VGD ++DH W E M R Y V NPGS V
Sbjct: 121 DAIKWVSDYLIKCHPS-PNVFYYQVGDGHLDHSWWGPAEVMQMDRPAYKVDLANPGSTVV 179
Query: 177 AETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSG 236
AE AAALA+A++VF DP Y+ ++ A+++Y FA + + SDS +A FY S+SG
Sbjct: 180 AEAAAALASAAVVFADRDPAYAATCIQHAKELYNFA---EITKSDSGYTAASGFYDSHSG 236
Query: 237 FKDELLWGAAWLFRATNAVKYYN----LIKTLGADDQPDIFS------WDDKYAGAHVLL 286
F DEL W WL+ AT Y N + G + Q +I S WDD + GA +LL
Sbjct: 237 FYDELSWAGVWLYLATGDETYLNKAEQYVAYWGTEPQTNIISYKWAHCWDDVHYGACLLL 296
Query: 287 SRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFL 346
++ + G + Y++ E + YT GL + +L+Y T+ FL
Sbjct: 297 AK---ITGKQ---IYKEAIERHLDYWSVGYNGERVHYTPKGLAWLDSWGSLRYATTTAFL 350
Query: 347 LTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRI 406
+ YA + + N AK+Q+DY LG + SY+VG+G N PKR
Sbjct: 351 ASVYADWEGCSREKAAIYN---------DFAKQQIDYALGSS--GRSYVVGFGVNPPKRP 399
Query: 407 HHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSH 466
HHR AH + D P +H ++L+GA+VGGP + D + DD ++Y +
Sbjct: 400 HHR-------TAH--SSWADSMSVPDYHR-----HVLIGALVGGPGKDDSYTDDINNYIN 445
Query: 467 SEPTTYINGAVVGPLA 482
+E N VG LA
Sbjct: 446 NEVACDYNAGFVGALA 461
>GUN6_DICDI (P22699) Endoglucanase precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Spore germination protein
270-6) (Cellulase)
Length = 705
Score = 258 bits (659), Expect = 2e-68
Identities = 176/490 (35%), Positives = 247/490 (49%), Gaps = 64/490 (13%)
Query: 11 LLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDG---- 66
+L +I LL + + +Y L +L+F + R+GRLP D + WR +S L D
Sbjct: 8 ILLIIFGLLSTQLINADTDYCSLLENALMFYKMNRAGRLP-DNDIPWRGNSALNDASPNS 66
Query: 67 -RLANVD--LSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRW 121
+ AN D LSGGY+DAGD VKF PMA++ TML WS IEY + I++
Sbjct: 67 AKDANGDGNLSGGYFDAGDGVKFGLPMAYSMTMLGWSFIEYESNIAQCGLTSLYLDTIKY 126
Query: 122 GTDYLLKCATSTPGRLYVG-VGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETA 180
GTD+L+ A T + G VGD NVDH W PEDM AR Y ++ PG+++A E A
Sbjct: 127 GTDWLI--AAHTADNEFAGQVGDGNVDHSWWGPPEDMTMARPTYMLTTEAPGTEIAMEAA 184
Query: 181 AALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDE 240
+ALAAAS+ FK +PTY+ + A+ ++ F Y+G YSDS+ +A FY S+SG+KD+
Sbjct: 185 SALAAASIAFKSSNPTYAATCLAHAKTLHNFGYTYRGVYSDSITNAQA-FYNSWSGYKDD 243
Query: 241 LLWGAAWLFRATNAVKYYNLIKT------LGADDQPDIFSWDDKYAGAHVLLSRRALLNG 294
L+WG+ WL++AT Y +G Q + WD+K G +LLS+
Sbjct: 244 LVWGSIWLYKATQDSDYLTKAVADYASGGVGGMAQGNSHDWDNKAPGCCLLLSKLV---- 299
Query: 295 DKNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYM 354
Y+ + E ++ L P YT GGL + +Y + FL
Sbjct: 300 -PTTSTYKTDFEGWLNYWL---PGGGVTYTPGGLAWIRQWGPARYAATAAFL----GSLA 351
Query: 355 SAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLP 414
+K T ++QVDY++G NP Q S++VG GPN+P HHR
Sbjct: 352 GTEKGT--------------DFTQKQVDYLIGNNPNQQSFVVGMGPNYPINPHHR----- 392
Query: 415 SLAAHPQTIGCDGGFQPFFHSSNP--NPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTY 472
AAH T +NP N +L GA+VGGP +D + DDR+DY +E T
Sbjct: 393 --AAHHSTTN---------DINNPVNNLYLLKGALVGGPGSNDEYTDDRTDYISNEVATD 441
Query: 473 INGAVVGPLA 482
N VG LA
Sbjct: 442 YNAGFVGALA 451
>GUNI_CLOTM (Q02934) Endoglucanase I precursor (EC 3.2.1.4) (EGI)
(Endo-1,4-beta-glucanase) (Cellulase I)
Length = 879
Score = 256 bits (653), Expect = 1e-67
Identities = 172/467 (36%), Positives = 239/467 (50%), Gaps = 47/467 (10%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQ-QLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFN 87
NY EAL K++ F + QRSG+L P +L WR DSGL DG+ A +DL+GG+YDAGD+VKFN
Sbjct: 77 NYGEALQKAIFFYECQRSGKLDPSTLRLNWRGDSGLDDGKDAGIDLTGGWYDAGDHVKFN 136
Query: 88 FPMAFTTTMLSWSAIEY--GKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPN 145
PM+++ ML W+ EY + Q I+W DY +KC Y VGD +
Sbjct: 137 LPMSYSAAMLGWAVYEYEDAFKQSGQYNHILNNIKWACDYFIKCHPE-KDVYYYQVGDGH 195
Query: 146 VDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTA 205
DH W E M R Y V ++PGS V AET+AALA AS++FKKVD YSK ++ A
Sbjct: 196 ADHAWWGPAEVMPMERPSYKVDRSSPGSTVVAETSAALAIASIIFKKVDGEYSKECLKHA 255
Query: 206 QKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKTL- 264
+++++FA + SD +A FY S+SGF DEL W A WL+ ATN Y + ++
Sbjct: 256 KELFEFA---DTTKSDDGYTAANGFYNSWSGFYDELSWAAVWLYLATNDSSYLDKAESYS 312
Query: 265 ---GADDQPDI------FSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPN 315
G + Q +I WDD G ++LL+R NG +Y++ E +
Sbjct: 313 DKWGYEPQTNIPKYKWAQCWDDVTYGTYLLLARIKNDNG-----KYKEAIERHLDWWTTG 367
Query: 316 SPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRS 375
YT GL + +L+Y T+ FL Y+ + + K T
Sbjct: 368 YNGERITYTPKGLAWLDQWGSLRYATTTAFLACVYSDWENGDK---------EKAKTYLE 418
Query: 376 VAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHS 435
A+ Q DY LG S++VG+G N PKR HHR AH D +P H
Sbjct: 419 FARSQADYALGST--GRSFVVGFGENPPKRPHHR-------TAHGS--WADSQMEPPEHR 467
Query: 436 SNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
++L GA+VGGP+ +D + DD S+Y+ +E N VG LA
Sbjct: 468 -----HVLYGALVGGPDSTDNYTDDISNYTCNEVACDYNAGFVGLLA 509
>GUNF_CLOTM (P26224) Endoglucanase F precursor (EC 3.2.1.4) (EGF)
(Endo-1,4-beta-glucanase) (Cellulase F)
Length = 739
Score = 254 bits (648), Expect = 5e-67
Identities = 169/466 (36%), Positives = 234/466 (49%), Gaps = 47/466 (10%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NY EAL K+++F + QRSG+LP +++ WR DS L DG +DL+GG+YDAGD+VKFN
Sbjct: 31 NYGEALQKAIMFYEFQRSGKLPENKRNNWRGDSALNDGADNGLDLTGGWYDAGDHVKFNL 90
Query: 89 PMAFTTTMLSWSAIEYGKR--MGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNV 146
PMA+ TML+WS E QL I+W TDY +KC S P Y VGD +
Sbjct: 91 PMAYAVTMLAWSVYESRDAYVQSGQLPYILDNIKWATDYFIKCHPS-PNVYYYQVGDGAL 149
Query: 147 DHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQ 206
DH W E M R + V NPGS V AETAAA+AA+S+VFK DP Y+ L+R A+
Sbjct: 150 DHSWWGPAEVMQMPRPSFKVDLTNPGSTVVAETAAAMAASSIVFKPTDPEYAATLLRHAK 209
Query: 207 KVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKTLGA 266
+++ FA + SD+ A +Y S+SGF DEL W + WL+ AT Y + ++
Sbjct: 210 ELFTFA---DTTRSDAGYRAAEGYYSSHSGFYDELTWASIWLYLATGDQSYLDKAESYEP 266
Query: 267 DDQPD----------IFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNS 316
+ + WD+K G+ +LL++ + G Y+Q EN +
Sbjct: 267 HWERERGTTLISYSWAHCWDNKLYGSLLLLAK---ITGK---SYYKQCIENHLDYWTVGF 320
Query: 317 PSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSV 376
S QYT GL + +L+Y T+ FL + YA + +
Sbjct: 321 NGSRVQYTPKGLAYLDRWGSLRYATTQAFLASVYADWSGCDP---------AKAAVYKEF 371
Query: 377 AKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSS 436
AK+QVDY LG S++VG+G N P+ HHR + A + C
Sbjct: 372 AKKQVDYALGST--GRSFVVGFGKNPPRNPHHRTAHSSWSALMTEPAEC----------- 418
Query: 437 NPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
+ILVGA+VGGP+ SD + D DY +E N VG LA
Sbjct: 419 ---RHILVGALVGGPDGSDSYVDRLDDYQCNEVANDYNAGFVGALA 461
>GUNG_CLOCE (P37700) Endoglucanase G precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase G) (Cellulase G) (EGCCG)
Length = 725
Score = 248 bits (632), Expect = 3e-65
Identities = 166/471 (35%), Positives = 237/471 (50%), Gaps = 58/471 (12%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NY EAL KS++F + QRSG LP D++ WR DSG+ DG VDL+GG+YDAGD+VKFN
Sbjct: 40 NYGEALQKSIMFYEFQRSGDLPADKRDNWRDDSGMKDGSDVGVDLTGGWYDAGDHVKFNL 99
Query: 89 PMAFTTTMLSWSAIE----YGKRMGPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDP 144
PM++T+ ML+WS E Y K Q K I+W DY +KC TPG Y VGD
Sbjct: 100 PMSYTSAMLAWSLYEDKDAYDK--SGQTKYIMDGIKWANDYFIKC-NPTPGVYYYQVGDG 156
Query: 145 NVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRT 204
DH W E M R + V + PGS V A TAA+LA+A++VFK DPTY++ +
Sbjct: 157 GKDHSWWGPAEVMQMERPSFKVDASKPGSAVCASTAASLASAAVVFKSSDPTYAEKCISH 216
Query: 205 AQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYY----NL 260
A+ ++ A + SD+ +A +Y S S F D+L W A WL+ ATN Y +
Sbjct: 217 AKNLFDMA---DKAKSDAGYTAASGYYSS-SSFYDDLSWAAVWLYLATNDSTYLDKAESY 272
Query: 261 IKTLGADDQPDIFS------WDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILP 314
+ G + Q DI + WDD + GA +LL++ N Y+ E +
Sbjct: 273 VPNWGKEQQTDIIAYKWGQCWDDVHYGAELLLAKLT------NKQLYKDSIEMNLDFWTT 326
Query: 315 NSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTL- 373
+ YT GL + +L++ T+ FL YA++ TP+ +
Sbjct: 327 GVNGTRVSYTPKGLAWLFQWGSLRHATTQAFLAGVYAEWEGC------------TPSKVS 374
Query: 374 --RSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQP 431
+ K Q+DY LG S++VGYG N P+ HHR AH G +
Sbjct: 375 VYKDFLKSQIDYALGST--GRSFVVGYGVNPPQHPHHR-------TAH-------GSWTD 418
Query: 432 FFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
S + + + GA+VGGP+ +DG+ D+ ++Y ++E N G LA
Sbjct: 419 QMTSPTYHRHTIYGALVGGPDNADGYTDEINNYVNNEIACDYNAGFTGALA 469
>GUNA_CALSA (P22534) Endoglucanase A precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase A) (Cellulase A)
Length = 1742
Score = 246 bits (627), Expect = 1e-64
Identities = 175/469 (37%), Positives = 243/469 (51%), Gaps = 53/469 (11%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNF 88
NY EAL K+++F + Q SG+LP + WR DS L DG+ +DL+GG++DAGD+VKFN
Sbjct: 27 NYGEALQKAIMFYEFQMSGKLPNWVRNNWRGDSALKDGQDNGLDLTGGWFDAGDHVKFNL 86
Query: 89 PMAFTTTMLSWSAIEYGKRM--GPQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNV 146
PM++T TMLSW+A EY QL+ I W DY +KC S Y VGD
Sbjct: 87 PMSYTGTMLSWAAYEYKDAFVKSGQLEHILNQIEWVNDYFVKCHPS-KYVYYYQVGDGGK 145
Query: 147 DHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQ 206
DH W E M R + V+ ++PGS V AETAA+LAAAS+V K +PT + ++ A+
Sbjct: 146 DHAWWGPAEVMQMERPSFKVTQSSPGSAVVAETAASLAAASIVLKDRNPTKAATYLQHAK 205
Query: 207 KVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKY--------Y 258
+Y+FA + + SDS +A +Y S+SGF DEL W A WL+ ATN Y
Sbjct: 206 DLYEFA---EVTKSDSGYTAANGYYNSWSGFYDELSWAAVWLYLATNDSTYLTKAESYVQ 262
Query: 259 NLIKTLGAD--DQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNS 316
N K G++ D WDD + GA +LL++ DK D Y+Q E+ +
Sbjct: 263 NWPKISGSNIIDYKWAHCWDDVHNGAALLLAKIT----DK--DTYKQIIESHLDYWTTGY 316
Query: 317 PSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYM---SAKKYTFNCGNVLVTPNTL 373
+YT GL + +L+Y T+ FL Y+ + + KK T+
Sbjct: 317 NGERIKYTPKGLAWLDQWGSLRYATTTAFLAFVYSDWSGCPTGKKETY------------ 364
Query: 374 RSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFF 433
R + Q+DY LG S++VG+G N PKR HHR + S A Q+I P +
Sbjct: 365 RKFGESQIDYALGST--GRSFVVGFGTNPPKRPHHR--TAHSSWADSQSI-------PSY 413
Query: 434 HSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLA 482
H + L GA+VGGP D + DD S+Y ++E N VG LA
Sbjct: 414 HR-----HTLYGALVGGPGSDDSYTDDISNYVNNEVACDYNAGFVGALA 457
>GUN2_PERAE (P23666) Endoglucanase 2 (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Abscission cellulase 2)
(Fragment)
Length = 130
Score = 167 bits (423), Expect = 6e-41
Identities = 77/124 (62%), Positives = 97/124 (78%), Gaps = 1/124 (0%)
Query: 362 NCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQ 421
+CG+ VT L S+AK+QVDYILGENP +MSYMVG+G +P+ +HHRGSSLPS+ AHP
Sbjct: 2 SCGSTTVTAKNLISLAKKQVDYILGENPAKMSYMVGFGERYPQHVHHRGSSLPSVHAHPN 61
Query: 422 TIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPL 481
I C+ GFQ + +SS+PNPNILVGAI+GGP+ D F DDR++Y SEP TYIN +VG L
Sbjct: 62 PIPCNAGFQ-YLYSSSPNPNILVGAILGGPDSRDSFSDDRNNYQQSEPATYINAPLVGAL 120
Query: 482 AYFA 485
A+FA
Sbjct: 121 AFFA 124
>GUNA_FIBSU (P23665) Endoglucanase A precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Cellulase)
Length = 453
Score = 57.0 bits (136), Expect = 1e-07
Identities = 96/432 (22%), Positives = 161/432 (37%), Gaps = 90/432 (20%)
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLAN-VDLSGGYYDAGDNVKFN 87
+Y EA + F QRSG+ P+ L S+ F N D+SGG++D GD+V +
Sbjct: 29 DYVEAAWMTTRFFGAQRSGQ-GPNWILDGTSNPTSFTKDSYNGKDVSGGWFDCGDHVMYG 87
Query: 88 FPMAFTTTMLSWSAIEYGK---------------------RMGP--QLKEARAAIRWGTD 124
+ + +L+ + E+ + + G ++++ +R+ D
Sbjct: 88 QSQGYASYVLALAYAEFTEVSTTFILVTTPTTRKPTTTPMKSGKPNKVRDLLEELRYEAD 147
Query: 125 YLLKCATSTPGRLYVGV-GDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAAL 183
+ +K A G +V V GD N DH+ W M + P + + L
Sbjct: 148 FWVKAAID--GNNFVTVKGDGNADHQKWVTAGAMSKLGSGEGGEPRCITGNANDGFTSGL 205
Query: 184 AAASL-VFKKVDPTYSKLL--VRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDE 240
AAA L V +VDP + ++ A+ Y +A ++G + GF +
Sbjct: 206 AAAMLAVMARVDPDTANQAKYLKAAKTAYSYAKSHKGVTNSQ-------------GFYES 252
Query: 241 LLWGAAW----------LFRATNAVKYYNLIKTLGADDQPDI-FSWDDKYAGAHVLLSRR 289
W W L+R T Y KT D ++ FS + G H + S
Sbjct: 253 SWWDGRWEDGPFLAELELYRTTGENSY----KTAAIDRYDNLKFSLGE---GTHFMYSNV 305
Query: 290 ALLNG---DKNFDQ----YRQEAENFMCKILPNSPSSSTQYTQGGL-MFKLPDSNLQYVT 341
L+ + F++ R+EA + I G+ K P
Sbjct: 306 VPLSAVMAEAVFEETPHGMRKEAIGVLDLIYEEKAKDKIFQNPNGMGSGKFP-------- 357
Query: 342 SITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGY--- 398
+ S + + + N + ++ V Y+LG+N + SY+VG+
Sbjct: 358 ---------VRVPSGGAFLYALSDKFNNTNEHMEMIEKNVSYLLGDNGSKKSYVVGFSKN 408
Query: 399 GPNFPKRIHHRG 410
G N P R HHRG
Sbjct: 409 GANAPSRPHHRG 420
>GUNC_CELFI (P14090) Endoglucanase C precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase C) (Cellulase C)
Length = 1101
Score = 53.9 bits (128), Expect = 9e-07
Identities = 89/416 (21%), Positives = 150/416 (35%), Gaps = 88/416 (21%)
Query: 71 VDLSGGYYDAGDNVKF--NFPMAFTTTMLSWSAI-----------------EYGKRMGPQ 111
+D+SGG+YDAGD+ K+ N +A + ++ E+G +
Sbjct: 495 LDVSGGWYDAGDHGKYVVNGGIAVGQLLQTYERALHAGTADALADGTLDVPEHGNDVPDV 554
Query: 112 LKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDHKCWE----RPEDMDTARTVYWVS 167
L EAR + W ++ P Y G+ V + W P D AR+++
Sbjct: 555 LDEARWELEWMLSMIV------PEGEYAGMVHHKVHDEGWTGLPLLPADDPQARSLH--- 605
Query: 168 PNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAV 227
P + +A A + + + DP ++ L+ A+ + A ++ Y+ A
Sbjct: 606 --RPSTAATLNLSAVAAQGARLLEPYDPQLAQTLLEAARTTWAAAQEHPALYAPGEAGAD 663
Query: 228 CPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLS 287
+ S DE W AA L+ T + + T D+F+ D G+ L
Sbjct: 664 GGGAYNDSQVADEFYWAAAELYLTTGEDAFATAV-TTSPLHTADVFTADGFGWGSVAALG 722
Query: 288 RRALL---NGDKNFDQYRQEAENFMCKILPNSPSSSTQYTQG-GLMFKLP------DSNL 337
R L N D + + L + Q QG G ++ P S+
Sbjct: 723 RLDLATVPNELPGLDAVQSSVVEGAQEYL------AAQAGQGFGSLYSPPGGEYVWGSSS 776
Query: 338 QYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVG 397
Q ++ + T Y L R+ +DY+ G N L SY+ G
Sbjct: 777 QVANNLVVVATAYD---------------LTGDERFRAATLEGLDYLFGRNALNQSYVTG 821
Query: 398 YGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPN-PNILVGAIVGGPN 452
+G +A+H Q + F H +P+ P+ G++ GGPN
Sbjct: 822 WG---------------EVASHQQ------HSRWFAHQLDPSLPSPPPGSLAGGPN 856
>GUN1_STRRE (Q05156) Cellulase 1 precursor (EC 3.2.1.4)
(Endoglucanase) (Endo-1,4-beta-glucanase) (Avicelase)
Length = 746
Score = 52.0 bits (123), Expect = 3e-06
Identities = 91/422 (21%), Positives = 150/422 (34%), Gaps = 94/422 (22%)
Query: 71 VDLSGGYYDAGDNVKFNFPMAFTTTML---------------------SWSAIEYGKRMG 109
+D++GG+YDAGD+ K+ T L + + E G ++
Sbjct: 331 LDVTGGWYDAGDHGKYVVNGGIATWELLSTYERSLTARTGHPAALGDGTLALPESGNKVP 390
Query: 110 PQLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDPNVDHKCWER----PEDMDTARTVYW 165
L EAR W ++LLK G+ G+ + + W P+ R ++
Sbjct: 391 DVLDEAR----WELEFLLKMQVPA-GQPLAGMAHHKLHDEQWTGLPLLPDQDPQKRELH- 444
Query: 166 VSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGS 225
P + AA A A+ +++ D ++ + A+ +Q AL + +D
Sbjct: 445 ----PPTTAATLNLAATAAQAARLYRPFDKAFAARALTAARTAWQAALAHPDLLADPNDG 500
Query: 226 AVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKTLGADDQ----PDIFSWDDKYAG 281
Y + DE W AA L+ T ++ + + P F W A
Sbjct: 501 TGGGAY-NDDDVTDEFYWAAAELYLTTGERQFADHVLDSPVHTADIFGPTGFDWGHTAAA 559
Query: 282 AHVLLSR--RALLNGDKNFDQYRQEAENFMCKI------LPNSPSSSTQYTQGGLMFKLP 333
+ L+ L D+ + A+ ++ + +P +P+ + +Y G L
Sbjct: 560 GRLDLALVPSRLPGRDQVRRSVIKAADTYLATLTAHPYGMPYAPAGN-RYDWGSSHQVL- 617
Query: 334 DSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMS 393
N V + + LT AKY R A + +DY+LG N L MS
Sbjct: 618 --NNGVVLASAYDLTGAAKY--------------------RDGALQGMDYVLGRNALNMS 655
Query: 394 YMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPN-PNILVGAIVGGPN 452
Y+ GYG H R + H +P PN G + GGPN
Sbjct: 656 YVTGYGEVSSHNQHSRW---------------------YAHQLDPTLPNPPSGTLAGGPN 694
Query: 453 QS 454
S
Sbjct: 695 SS 696
>GUNA_PSEFL (P10476) Endoglucanase A precursor (EC 3.2.1.4)
(Endo-1,4-beta-glucanase) (Cellulase) (EGA)
Length = 962
Score = 49.3 bits (116), Expect = 2e-05
Identities = 100/432 (23%), Positives = 158/432 (36%), Gaps = 83/432 (19%)
Query: 70 NVDLSGGYYDAGDNVKFNFPMAFTTTML------------SWSAIEYGKRMGPQLKEARA 117
+++++ G+YDAGD+ K+ + L + +A+ G P+ A
Sbjct: 191 SLNVTKGWYDAGDHGKYVVNGGISVWTLLNLYERAQHITGNLAAVADGSMNIPESGNGVA 250
Query: 118 AI----RWGTDYLLKCATSTP-GRLYVGVGDPNVDHKCWE----RPEDMDTARTVYWVSP 168
I RW +++L A P G+ G+ + W P + R +
Sbjct: 251 DILDEARWQMEFML--AMQVPQGQAKAGMAHHKIHDVGWTGLPLAPHEDPQQRALV---- 304
Query: 169 NNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQ-----YQGSYSDSL 223
P + AA A A+ ++K +D ++ L + A++ + A Y G+Y +
Sbjct: 305 -PPSTAATLNLAATAAQAARIWKDIDAGFAALCLTAAERAWNAAQANPNDIYSGNYDNGG 363
Query: 224 GSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIK--TLGADDQPDIFSWDDKYAG 281
G F DE W AA L+ T +Y I TL D F W D
Sbjct: 364 GGYGDRFVA------DEFYWAAAELYITTGDSRYLPTINNYTLERTD----FGWPD---- 409
Query: 282 AHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVT 341
+L + + + R A N + I +S+ TQ + P S+L+Y
Sbjct: 410 TELLGVMSLAVVPATHTNSLRIAARNHIQTI-----ASTHLTTQSASGYPAPLSSLEYYW 464
Query: 342 SITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPN 401
++ M Y F+ GN N V+K ++Y+ G N L S++ G G N
Sbjct: 465 GSNSVIANKLVLMGLA-YDFS-GN----QNFALGVSKG-INYLFGINVLPTSFITGLGTN 517
Query: 402 FPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQ--SDGFPD 459
+ HHR F +SN P GA+ GGPN D F
Sbjct: 518 TVAQPHHR-------------------FWAGALNSN-YPWAPPGALSGGPNAGLEDSFSA 557
Query: 460 DRSDYSHSEPTT 471
R S P T
Sbjct: 558 SRLSGCTSRPAT 569
>GUNC_BUTFI (P23658) Cellodextrinase (EC 3.2.1.-)
Length = 547
Score = 47.0 bits (110), Expect = 1e-04
Identities = 91/414 (21%), Positives = 158/414 (37%), Gaps = 68/414 (16%)
Query: 71 VDLSGGYYDAGDNVKFNFPMAFTTTMLSWSA----------IEYGKRMGPQ--LKEARAA 118
VD++GG++DAGD +++ A L + + K G + L E A
Sbjct: 137 VDVTGGWHDAGDYGRYSTAGAVAVAHLLYGVRFFKGLLDVHYDIPKVAGDKGNLPEILAE 196
Query: 119 IRWGTDYLLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAE 178
++ D+L+K G ++ V +H + PED ++ VS S A+
Sbjct: 197 VKVELDFLMKMQREN-GSVWHKV--TTFNHAPFLMPEDDREELFLFSVS-----SLATAD 248
Query: 179 TAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGS--YSDSLGSAVCPFYCSYSG 236
AA A A V+K+ D Y+ L++ + Y++ L + + GS Y
Sbjct: 249 IAAVFALAYTVYKEYDAEYADKLMQKSLLAYKWLLDNPDELLFFNPDGSNT----GQYDE 304
Query: 237 FKD--ELLWGAAWLFRATNAVKYYNLIKTL-GADDQPDIFSWDDKYAGAHVLLSRRALLN 293
+D W A L+ AT+ KYY+ + L ++ D + Y G A +
Sbjct: 305 AEDISNRFWAACALYEATSDGKYYSDAQELKNRLEEFDKNAQKKGYQGNVFTCLGWAEVA 364
Query: 294 GDKNFDQYRQEAENFMCKILPNSPSSSTQ-----YTQGGLMFKLPDSNLQYVTSITFLLT 348
G + + EN +C + NS + + G + +++ + +++ L
Sbjct: 365 GLGSLSLLLKREENALCSLARNSFVAEADRLVKVSKENGFGLCMGENDFIWGSNMELL-- 422
Query: 349 TYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHH 408
KYM N + + +DYILG N + +SY+ G G K H
Sbjct: 423 ---KYMMVLSTAIRIDN----KPEYKLALEAGLDYILGCNSMDISYVTGNGEKAFKNPHL 475
Query: 409 RGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRS 462
R +++ + P P G + GGPN G D+R+
Sbjct: 476 RPTAVDDI-------------------EEPWP----GLVSGGPN--SGLHDERA 504
>GUND_CLOTM (P04954) Endoglucanase D precursor (EC 3.2.1.4) (EGD)
(Endo-1,4-beta-glucanase) (Cellulase D)
Length = 649
Score = 46.2 bits (108), Expect = 2e-04
Identities = 98/450 (21%), Positives = 153/450 (33%), Gaps = 92/450 (20%)
Query: 65 DGRLANVDLSGGYYDAGDNVKF--NFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWG 122
+G+ D + G++DAGD K+ N + + L+W + ++ P E
Sbjct: 184 NGQHTKKDSTKGWHDAGDYNKYVVNAGITVGSMFLAWE--HFKDQLEPVALEIPEKNNSI 241
Query: 123 TDYL--LKCATSTPGRLYVGVGDPNVDHKCWER-------PEDMDTART-VYWVSPNNPG 172
D+L LK + G V HK R PE+ R V W
Sbjct: 242 PDFLDELKYEIDWILTMQYPDGSGRVAHKVSTRNFGGFIMPENEHDERFFVPW------S 295
Query: 173 SDVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYC 232
S A+ A A A+ +F+ DP Y++ + A+ Y+F + + Y
Sbjct: 296 SAATADFVAMTAMAARIFRPYDPQYAEKCINAAKVSYEFLKNNPANVFANQSGFSTGEYA 355
Query: 233 SYSGFKDELLWGAAWLFRATNAVKYYNLIKTLGADDQPDI---FSWDDKYAGAHVLLSRR 289
+ S D+ LW AA ++ +Y + A I F WD+
Sbjct: 356 TVSD-ADDRLWAAAEMWETLGDEEYLRDFENRAAQFSKKIEADFDWDNV----------- 403
Query: 290 ALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSI--TFLL 347
N + +L P + Q + DS L SI T
Sbjct: 404 ------ANLGMFTY--------LLSERPGKNPALVQ-----SIKDSLLSTADSIVRTSQN 444
Query: 348 TTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKR-------------QVDYILGENPLQMSY 394
Y + + Y + C +V + VA + + ++ G N SY
Sbjct: 445 HGYGRTLGT-TYYWGCNGTVVRQTMILQVANKISPNNDYVNAALDAISHVFGRNYYNRSY 503
Query: 395 MVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQS 454
+ G G N P H R S G DG ++P+ P LVG G P
Sbjct: 504 VTGLGINPPMNPHDRRS------------GADGIWEPW-------PGYLVGG--GWPGPK 542
Query: 455 DGFPDDRSDYSHSEPTTYINGAVVGPLAYF 484
D + D + Y +E N A++ LA F
Sbjct: 543 D-WVDIQDSYQTNEIAINWNAALIYALAGF 571
>ETF2_SFVKA (Q9Q8Y2) Early transcription factor 82 kDa subunit (VETF
large subunit)
Length = 711
Score = 31.2 bits (69), Expect = 6.3
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 3/59 (5%)
Query: 26 CNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNV 84
C PNY+ L K+L F + + + D ++ +D L DG+ NVDLS Y N+
Sbjct: 594 CPPNYK--LVKNL-FPRQTKFATIRSDSGMELTTDGFLVDGKEFNVDLSSNYVAFTKNI 649
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 62,782,918
Number of Sequences: 164201
Number of extensions: 2854050
Number of successful extensions: 5652
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 5559
Number of HSP's gapped (non-prelim): 27
length of query: 488
length of database: 59,974,054
effective HSP length: 114
effective length of query: 374
effective length of database: 41,255,140
effective search space: 15429422360
effective search space used: 15429422360
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 68 (30.8 bits)
Lotus: description of TM0083.8


